Conjugative Plasmid Delivery of CRISPR Systems: A Precision Strategy for Biofilm Control and Combating Antimicrobial Resistance
The escalating global health crisis of antimicrobial resistance (AMR) is profoundly fueled by biofilm-associated infections, which confer immense tolerance to conventional antibiotics.
Conjugative Plasmid Delivery of CRISPR Systems: A Precision Strategy for Biofilm Control and Combating Antimicrobial Resistance
Abstract
The escalating global health crisis of antimicrobial resistance (AMR) is profoundly fueled by biofilm-associated infections, which confer immense tolerance to conventional antibiotics. This article explores the transformative potential of leveraging conjugative plasmids for the targeted delivery of CRISPR-Cas systems as a novel anti-biofilm strategy. We provide a comprehensive analysis spanning the foundational science of bacterial conjugation and biofilm biology to the methodological engineering of CRISPR payloads that disrupt antibiotic resistance genes, quorum sensing, and biofilm integrity. The content details critical optimization strategies for enhancing delivery efficiency, including the use of cis-acting plasmids and nanoparticle hybrids, and rigorously evaluates this approach against traditional therapies and alternative delivery vectors. Aimed at researchers and drug development professionals, this review synthesizes cutting-edge advances and translational challenges, positioning conjugative CRISPR delivery as a promising paradigm for next-generation, precision antimicrobials.
The Biofilm Challenge and the CRISPR-Conjugation Synergy
Biofilm Architecture and Its Role in Antibiotic Treatment Failure
Bacterial biofilms are structured microbial communities encased in a self-produced extracellular polymeric substance (EPS) matrix that demonstrate remarkable resilience to antimicrobial treatments [1] [2]. This architectural complexity constitutes a primary factor in the persistence of chronic infections and represents a significant challenge in clinical management. Biofilm-associated bacteria can exhibit up to 1,000-fold increased resistance to antibiotics compared to their planktonic counterparts, leading to treatment failures across diverse medical contexts [3] [4].
The protective nature of biofilms is particularly problematic in healthcare-associated infections (HAIs), where they contribute substantially to morbidity, mortality, and economic burden. Recent estimates indicate that biofilms are implicated in approximately 65-80% of all human microbial infections, with associated costs reaching billions of dollars annually in healthcare expenditures alone [4] [5]. Understanding the structural and functional basis of biofilm-mediated resistance is crucial for developing effective countermeasures, including innovative approaches like conjugative plasmid delivery of CRISPR systems for targeted biofilm control.
Structural Architecture of Biofilms
Composition and Organization
The biofilm matrix is a complex, dynamic assemblage of biopolymers that forms a protective barrier around microbial populations. This extracellular polymeric substance (EPS) comprises primarily polysaccharides, proteins, extracellular DNA (eDNA), and lipids that together create a three-dimensional architecture with heterogeneous structural properties [1] [2] [4]. The matrix can constitute over 90% of the biofilm's dry mass, creating a formidable physical and functional barrier against antimicrobial penetration [2].
The structural organization of biofilms is characterized by microcolonies interspersed with water channels that facilitate nutrient distribution and waste removal. This arrangement creates diverse microenvironments with varying metabolic gradients, pH, oxygen availability, and chemical signaling profiles [6]. The heterogeneous nature of this architecture means that bacteria in different regions of the biofilm experience distinct selective pressures and develop specialized physiological adaptations.
Developmental Lifecycle
Biofilm formation follows a programmed developmental sequence that transforms free-living planktonic bacteria into structured, surface-associated communities:
Initial Reversible Attachment: Planktonic cells adhere to conditioned surfaces through weak interactions (van der Waals forces, electrostatic interactions) [1] [7]. Surface characteristics including roughness, hydrophobicity, and chemistry significantly influence this initial attachment phase.
Irreversible Attachment: Bacterial surface structures (pili, fimbriae, adhesins) strengthen attachment, transitioning to permanent association [1] [2]. Production of early EPS matrix components anchors cells firmly to the substrate.
Microcolony Formation: Attached cells proliferate and form clustered communities, initiating three-dimensional development. Quorum sensing signaling becomes activated, coordinating collective behavior through chemical communication [2] [8].
Maturation: Development of complex architectural features including tower/mushroom structures and fluid channels. The EPS matrix matures with full compositional complexity, and metabolic heterogeneity develops within subpopulations [1] [2].
Dispersion: Active release of cells from the biofilm to colonize new surfaces. This can occur through seeding dispersal, erosion, or sloughing mechanisms, completing the lifecycle and enabling biofilm propagation [2].
Table 1: Biofilm Developmental Stages and Key Characteristics
| Developmental Stage | Key Processes | Regulatory Mechanisms | Structural Features |
|---|---|---|---|
| Initial Attachment | Reversible adhesion, surface conditioning | Van der Waals forces, electrostatic interactions | Single-layer cells, weak binding |
| Irreversible Attachment | Production of adhesins, early EPS secretion | Surface protein expression, signaling initiation | Firmly anchored cells, monolayer |
| Microcolony Formation | Cellular proliferation, cluster formation | Quorum sensing activation, c-di-GMP signaling | Multilayered cell aggregates |
| Maturation | EPS matrix production, structural organization | Full quorum sensing response, metabolic differentiation | Mushroom/tower structures, water channels |
| Dispersion | Active cellular release, matrix degradation | Environmental stress response, nutrient sensing | Detaching cells, hollow cavities |
Mechanisms of Antibiotic Treatment Failure
The architectural and physiological complexity of biofilms confers resistance through multiple concurrent mechanisms that operate at different levels of biofilm organization. These can be broadly categorized into physical/chemical barriers, physiological adaptations, and genetic evolutionary processes.
Physical and Chemical Barrier Mechanisms
The EPS matrix functions as a formidable physical barrier that restricts antibiotic penetration through several mechanisms:
Diffusion Limitation: The dense, anionic matrix structure creates a sieving effect that physically impedes antibiotic penetration, particularly for larger molecules [2] [3]. The EPS meshwork pore size can exclude certain antimicrobial agents based on molecular dimensions.
Binding and Inactivation: Matrix components can directly bind and neutralize antimicrobial compounds. Positively charged aminoglycosides are particularly susceptible to binding with anionic eDNA in the matrix, effectively reducing bioavailable concentrations [2] [3].
Enzyme-Mediated Inactivation: Biofilms harbor extracellular enzymes such as β-lactamases that can degrade antibiotics before they reach their cellular targets. The localized high density of these enzymes in the matrix creates an effective inactivation zone [2] [6].
Altered Microenvironment: Metabolic activity within biofilms creates chemical gradients that reduce antibiotic efficacy. Oxygen depletion in deeper layers diminishes the activity of oxygen-dependent antibiotics, while acidic pH zones can neutralize pH-sensitive drugs [3] [4].
Table 2: Biofilm-Mediated Antibiotic Resistance Mechanisms
| Resistance Mechanism | Functional Basis | Antibiotics Affected | Resistance Factor |
|---|---|---|---|
| Restricted Penetration | Physical barrier by EPS matrix limiting diffusion | Aminoglycosides, β-lactams | Delayed/incomplete penetration |
| Binding/Neutralization | Chemical interaction with matrix components | Aminoglycosides, vancomycin | Reduced effective concentration |
| Enzymatic Inactivation | Extracellular enzyme production | β-lactams, chloramphenicol | Direct degradation |
| Altered Microenvironment | Chemical gradient formation (O₂, pH) | Aminoglycosides, fluoroquinolones | Reduced antibacterial activity |
| Metabolic Heterogeneity | Slow growth/non-growing persister cells | β-lactams, glycopeptides | Target inactivity |
| Enhanced Efflux Pumps | Upregulated efflux systems | Tetracyclines, macrolides | Active antibiotic export |
| Horizontal Gene Transfer | Conjugative plasmid exchange | Multiple classes | Genetic resistance acquisition |
Physiological Adaptations
Biofilm inhabitants undergo significant physiological reprogramming that enhances their tolerance to antimicrobial agents:
Metabolic Heterogeneity: The graded environments within biofilms create subpopulations with diverse metabolic states, including dormant or slow-growing persister cells that are intrinsically tolerant to growth-dependent antibiotics [2] [4] [6].
Stress Response Activation: The biofilm lifestyle induces general stress responses that enhance cellular protection mechanisms, including DNA repair systems, chaperone proteins, and oxidative stress defenses [4] [7].
Persister Cell Formation: A subpopulation of dormant bacterial cells exhibits exceptional tolerance to antimicrobials without genetic mutation. These persister cells can reseed biofilms after antibiotic treatment is discontinued, leading to chronic recurrence [3] [4].
Application Notes: Conjugative Plasmid Delivery of CRISPR Systems
Conceptual Framework and Rationale
The development of conjugative plasmid systems for delivering CRISPR-based antimicrobials represents a promising strategic approach to overcome biofilm-mediated treatment failure. This methodology leverages bacterial mating mechanisms to introduce targeted genetic interventions directly into biofilm communities [9] [6]. The approach offers several distinct advantages:
Precision Targeting: CRISPR-Cas systems can be programmed to selectively disrupt antibiotic resistance genes, virulence factors, or essential biofilm maintenance genes without affecting non-target species [9] [10].
Bypassing Physical Barriers: Conjugative transfer actively delivers CRISPR components through the EPS matrix using natural bacterial mating mechanisms, effectively circumventing diffusion limitation issues that plague conventional antibiotics [9] [6].
Programmable Resistance Reversal: By specifically targeting and eliminating resistance genes (e.g., β-lactamases, efflux pump components), CRISPR delivery can resensitize biofilm populations to conventional antibiotics [6].
Synergistic Combinatorial Therapy: Conjugative CRISPR delivery can be combined with traditional antibiotics in sequential treatment regimens, where biofilm architectural disruption enhances subsequent antibiotic penetration and efficacy [6].
Experimental Protocol: Conjugative Plasmid Delivery for Biofilm Control
Materials and Reagents
Table 3: Research Reagent Solutions for Conjugative CRISPR Delivery
| Reagent/Category | Specific Examples | Function/Application | Considerations |
|---|---|---|---|
| CRISPR-Cas System | Cas9 nuclease, guide RNA (gRNA) constructs | Targeted gene disruption in biofilm cells | Species-specific gRNA design required |
| Delivery Vector | Conjugative plasmids (RP4, R6K derivatives), phagemids | Intercellular transfer of CRISPR components | Broad host range vs. narrow specificity |
| Donor Strains | E. coli S17-1, WM3064 (diaminopimelic acid auxotroph) | Conjugation machinery source | Selection marker compatibility |
| Biofilm Models | Flow cell systems, MBEC assay plates, catheter segments | Experimental biofilm cultivation | In vitro vs. in vivo model relevance |
| Selection Agents | Antibiotics, nutritional supplements | Donor/transconjugant selection | Resistance marker expression |
| Detection Tools | Fluorescence reporters, PCR validation primers | Transfer efficiency assessment | Quantitative vs. qualitative measurement |
Protocol Workflow
Phase 1: Conjugative Plasmid Design and Donor Strain Preparation
CRISPR Payload Construction: Clone CRISPR-Cas9 components with species-specific guide RNAs targeting biofilm-associated genes (e.g., pelA, pslA, algD for polysaccharide synthesis; lasI, rhlI for quorum sensing; antibiotic resistance genes) into a broad-host-range conjugative plasmid backbone [9] [6].
Donor Strain Transformation: Introduce the constructed plasmid into appropriate donor strains (e.g., E. coli S17-1) via electroporation or chemical transformation. Validate CRISPR function in donor strains prior to conjugation experiments.
Biofilm Cultivation: Grow mature biofilms (48-72 hours) of target pathogens (e.g., P. aeruginosa, S. aureus) in relevant model systems (flow cells, catheter segments, or 96-well peg lids). Standardized biofilm assays like the MBEC (Minimum Biofilm Eradication Concentration) system provide reproducible platforms [2] [4].
Phase 2: Conjugation and Plasmid Transfer
Donor-Biofilm Co-incubation: Introduce donor strains to established biofilms at optimized donor:recipient ratios (typically 1:1 to 10:1) in appropriate conjugation media. Incubation periods typically range from 4-24 hours depending on model system and bacterial species [9].
Selection and Counter-Selection: Apply appropriate selection agents to eliminate donor strains while selecting for transconjugants (biofilm cells that have received the conjugative plasmid). Nutritional counter-selection or differential antibiotic resistance profiles are commonly employed.
Conjugation Efficiency Assessment: Quantify transfer efficiency through CFU enumeration, fluorescence activation, or PCR-based detection of plasmid markers in biofilm residents. Optimal systems typically achieve conjugation efficiencies of 10⁻² to 10⁻⁴ transconjugants per recipient [9] [6].
Phase 3: Functional Assessment and Combinatorial Treatment
Biofilm Disruption Analysis: Quantify changes in biofilm architecture using confocal laser scanning microscopy (CLSM) with specific staining (SYTO9/propidium iodide for viability, concanavalin A for matrix polysaccharides). Measure reductions in biofilm biomass (50-90% reduction) and structural integrity [6].
Gene Editing Validation: Confirm targeted gene disruption in transconjugants through PCR amplification, sequencing, or functional assays for lost gene function (e.g., reduced matrix production, abolished antibiotic resistance).
Combinatorial Antibiotic Sensitivity Testing: Assess resensitization to conventional antibiotics by determining MBEC reductions following CRISPR delivery. Successful interventions typically demonstrate 10-1000 fold reductions in biofilm eradication concentrations [6].
Dispersal and Viability Impact: Quantify changes in cellular dispersal from biofilms and overall reductions in viable counts. Effective treatments typically reduce viable counts by 3-5 log units compared to untreated controls [6].
Technical Considerations and Optimization Parameters
Successful implementation of conjugative CRISPR delivery for biofilm control requires careful optimization of several technical parameters:
Donor-Recipient Compatibility: Match conjugative systems with appropriate origin of transfer (oriT) sequences and mating pair formation mechanisms compatible with target biofilm species [9].
Temporal Control: Coordinate CRISPR expression with conjugation timing using inducible promoter systems to prevent premature Cas9/gRNA expression in donor strains.
Delivery Efficiency Enhancement: Utilize nanoparticle co-delivery systems to improve conjugation efficiency in dense biofilm matrices. Lipid-based nanoparticles have demonstrated 3.5-fold increases in delivery efficiency [6].
Multiplexed Targeting: Design gRNA arrays to simultaneously disrupt multiple redundant biofilm pathways (e.g., matrix production, quorum sensing, resistance genes) to prevent compensatory adaptations [9] [10].
Escape Mutation Monitoring: Include multiple experimental replicates and control for potential CRISPR immune escape through target site mutation or spacer acquisition.
The structural and functional complexity of biofilm architecture presents a multi-faceted barrier to effective antibiotic therapy, contributing significantly to treatment failure in chronic infections. The integrated physical, physiological, and genetic resistance mechanisms employed by biofilm communities necessitate innovative approaches that specifically target biofilm vulnerabilities.
Conjugative plasmid delivery of CRISPR systems represents a promising strategic platform for precision biofilm control that directly addresses several key resistance mechanisms. This approach enables targeted genetic interventions against resistance determinants and biofilm maintenance genes while leveraging natural bacterial mating mechanisms to bypass physical diffusion barriers. When combined with conventional antibiotics in sequential treatment regimens, this technology demonstrates potential for significant biofilm eradication and resensitization of persistent infections to standard therapies.
Future development in this field will likely focus on improved delivery efficiency through engineered conjugative systems, multiplexed targeting strategies against redundant biofilm pathways, and integration with nanoparticle technologies for enhanced penetration. As these approaches mature, conjugative CRISPR delivery systems may provide powerful tools for addressing the persistent challenge of biofilm-mediated antibiotic treatment failure in clinical settings.
Biofilm-mediated resistance represents a significant challenge in treating chronic bacterial infections and combating antimicrobial resistance (AMR). Biofilms are structured communities of microbial cells enclosed in a self-produced extracellular polymeric substance (EPS) matrix that can be attached to a biotic or abiotic surface [1]. This mode of growth provides inherent protection against antibiotics and host immune responses, leading to persistent infections that are difficult to eradicate [2] [11]. The resistance mechanisms employed by biofilm-associated bacteria are multifaceted, encompassing both physical barrier functions and physiological adaptations that distinguish them from their planktonic counterparts [2] [12]. Understanding these mechanisms is crucial for developing novel therapeutic strategies, including emerging approaches that utilize conjugative plasmid delivery of CRISPR systems for precise biofilm control [13] [14].
Biofilm Architecture and Developmental Lifecycle
Structural Composition of Biofilms
The biofilm matrix is a complex, dynamic environment composed primarily of extracellular polymeric substances (EPS) that can constitute up to 90% of the biofilm's biomass [12]. This matrix forms a protective barrier that encases microbial cells in a three-dimensional architecture, creating heterogeneous microenvironments with gradients of nutrients, oxygen, and metabolic activity [15] [1]. The EPS comprises a variety of biopolymers including polysaccharides, proteins, lipids, and extracellular DNA (eDNA) [2]. This composition varies significantly depending on the microbial species present, nutrient availability, and environmental conditions [2].
The structural integrity of biofilms is maintained by this matrix, which not only provides physical protection but also facilitates social interactions and horizontal gene transfer between bacterial cells [12]. The matrix architecture often features water channels that allow for nutrient distribution and waste removal, while the heterogeneous organization results in subpopulations of bacteria with distinct metabolic states and functions [13] [1].
Developmental Stages of Biofilm Formation
Biofilm formation follows a regulated developmental cycle comprising distinct stages:
Initial Reversible Attachment: Planktonic cells adhere to surfaces through weak interactions such as van der Waals forces and electrostatic interactions [1]. Surface characteristics including roughness significantly influence this attachment phase [1].
Irreversible Attachment: Production of adhesive extracellular polymeric substances enables firm attachment of cells to the substrate and to each other [2] [1]. This stage is often regulated by intracellular signaling molecules such as cyclic diguanylate monophosphate (c-di-GMP) [2].
Microcolony Formation and Maturation: Attached cells proliferate and develop into structured microcolonies with characteristic architectural features [2]. The EPS matrix is extensively produced during this phase, creating the protective three-dimensional structure [2].
Dispersion: Active or passive release of cells from the biofilm enables colonization of new niches [2]. This can occur through seeding (central hollowing), erosion, or sloughing in response to environmental cues such as nutrient limitation [2].
Table 1: Key Stages in Biofilm Development
| Developmental Stage | Key Processes | Regulatory Factors |
|---|---|---|
| Initial Attachment | Reversible adhesion to surfaces | Surface roughness, preconditioning, weak physical forces |
| Irreversible Attachment | EPS production, firm adhesion | c-di-GMP signaling, adhesin production |
| Maturation | Microcolony formation, structural development | Quorum sensing, continued EPS production |
| Dispersion | Active or passive cell release | Nutrient limitation, environmental stress signals |
Figure 1: Biofilm Developmental Lifecycle. The process begins with initial attachment of planktonic cells, progresses through irreversible attachment and maturation, and culminates in dispersion that releases cells to colonize new surfaces.
Fundamental Mechanisms of Biofilm-Mediated Resistance
Physical Barrier Function of the Extracellular Matrix
The EPS matrix serves as a formidable physical barrier that significantly limits antibiotic penetration to cells embedded within the biofilm structure [2] [13]. This matrix can bind to antimicrobial agents through various mechanisms, effectively reducing the concentration that reaches bacterial cells in the deeper layers of the biofilm [2]. Positively charged antibiotics such as aminoglycosides particularly interact with negatively charged components of the matrix like eDNA, leading to sequestration and reduced efficacy [2]. In some cases, enzymes within the biofilm matrix can directly degrade or modify antibiotics before they reach their cellular targets [2].
The barrier function is further enhanced in clinical settings where host components integrate into the biofilm structure. For instance, in cystic fibrosis lung infections, eDNA produced by Pseudomonas aeruginosa combines with host eDNA to form a protective shield that limits tobramycin penetration [2]. Similarly, neutrophil extracellular traps (NETs) released by host immune cells can surround biofilms, creating an additional physical barrier that hinders antibiotic access while simultaneously containing bacterial dissemination [2].
Physiological Heterogeneity and Metabolic Dormancy
Biofilms exhibit significant physiological heterogeneity due to the creation of microenvironments with varying nutrient and oxygen availability [12] [15]. This heterogeneity leads to distinct subpopulations of cells with different metabolic states and growth rates [12]. Cells in the inner layers of biofilms often experience nutrient limitation and develop into metabolically dormant states [12] [15]. Since most conventional antibiotics target actively growing cells, these dormant populations exhibit dramatically increased tolerance to antimicrobial treatments [12].
The metabolic dormancy in biofilms is not uniform but represents a continuum of metabolic states, with the most dormant cells often localized in the deepest regions of the biofilm where nutrient diffusion is most limited [12] [15]. This gradient of metabolic activity contributes to the recalcitrance of biofilm infections, as dormant cells can survive antibiotic treatment and repopulate the biofilm once antimicrobial pressure is removed [12].
Persister Cell Formation and Their Role in Chronic Infections
Persister cells represent a subpopulation of phenotypic variants that exhibit extreme tolerance to antibiotics without undergoing genetic change [12]. These cells are characterized by reduced metabolic activity and a transient, non-growing state that allows them to survive concentrations of antibiotics that kill their genetically identical counterparts [12]. The formation of persister cells is controlled by bacterial growth phases and environmental stress factors, with their proportion increasing significantly during stationary phase and in mature biofilms [12].
Persister cells play a crucial role in chronic and relapsing infections. Their ability to survive antimicrobial treatment and resume growth once antibiotics are removed contributes significantly to treatment failures [12]. In diseases such as cystic fibrosis, where P. aeruginosa establishes chronic lung infections, high-persister (hip) mutants have been identified in patients undergoing repeated antibiotic therapies, demonstrating the clinical relevance of this tolerance mechanism [12]. Similarly, in cancer patients with oral Candida albicans infections, long-term carriers developed hip mutants that contributed to persistent biofilm-associated infections [12].
Table 2: Key Mechanisms of Biofilm-Mediated Antimicrobial Resistance
| Resistance Mechanism | Functional Basis | Impact on Antibiotic Efficacy |
|---|---|---|
| Physical Barrier | EPS matrix limits antibiotic penetration | Reduced drug concentration at target sites |
| Metabolic Dormancy | Heterogeneous metabolic states | Tolerance to growth-dependent antibiotics |
| Persister Cells | Dormant phenotypic variants | Survival after high-dose antibiotic treatment |
| Enhanced HGT | Efficient intercellular gene transfer | Dissemination of genetic resistance determinants |
Figure 2: Mechanisms of Biofilm-Mediated Antibiotic Resistance. Antibiotics face multiple barriers including limited penetration through the EPS matrix, reduced efficacy against dormant cells, and complete tolerance by persister cells.
Conjugative Plasmid Delivery of CRISPR Systems for Biofilm Control
Principles of CRISPR-Based Antimicrobial Strategy
The CRISPR/Cas9 system has emerged as a powerful tool for precision targeting of bacterial pathogens through specific genetic interventions [13] [15]. This system consists of two key components: the Cas9 nuclease, which introduces double-strand breaks in DNA, and a guide RNA (gRNA) that directs Cas9 to specific genomic sequences [13]. By designing gRNAs to target essential genes, antibiotic resistance genes, or biofilm regulatory elements, researchers can selectively eliminate bacterial pathogens or resensitize them to conventional antibiotics [13] [15].
The specificity of CRISPR-based antimicrobials offers significant advantages over broad-spectrum antibiotics, particularly for targeting pathogens within complex microbial communities without disrupting commensal populations [13] [14]. This precision approach is especially valuable for tackling biofilm-associated infections where traditional antibiotics often fail due to the mechanisms described previously [13].
Conjugative Plasmid Delivery Systems
Conjugative plasmid delivery represents an efficient method for introducing CRISPR systems into target bacterial populations [14]. These plasmid-based systems exploit natural bacterial mating mechanisms to transfer genetic material between cells, offering several advantages including broad host range, resistance to restriction-modification systems, and the capacity to deliver large genetic payloads [14].
Research has demonstrated that cis-acting conjugative plasmids, which encode both the conjugation machinery and CRISPR nuclease on a single vector, achieve significantly higher conjugation frequencies compared to trans systems where these elements are separated [14]. In co-culture experiments with Escherichia coli donors and Salmonella enterica recipients, cis-conjugative plasmids demonstrated conjugation frequencies approaching 100% under conditions that enhanced cell-to-cell contact, such as inclusion of glass beads to promote biofilm-like environments [14]. This high efficiency stems from the fact that bacteria receiving the cis-conjugative plasmid become donors themselves, leading to exponentially increasing numbers of conjugative donors in the population [14].
Experimental Protocol: Conjugative Transfer of CRISPR Systems
Protocol Title: Assessment of Conjugative Plasmid Delivery of CRISPR/Cas9 for Targeted Bacterial Killing
Principle: This protocol describes a method for evaluating the efficacy of cis-conjugative plasmids in delivering CRISPR/Cas9 systems to target bacterial biofilms. The approach leverages the innate efficiency of conjugative transfer under conditions that promote cell-to-cell contact, mimicking biofilm environments [14].
Materials:
- Donor strain: E. coli carrying pNuc-cis plasmid (or similar cis-conjugative CRISPR plasmid)
- Recipient strain: Target biofilm-forming bacteria (e.g., S. enterica, P. aeruginosa)
- Low-salt LB media (LSLB: 0.25% NaCl w/v)
- Antibiotics for selection (appropriate for plasmid markers and recipient strain)
- Arabinose for induction of Cas9 expression
- 0.5 mm glass beads (for enhanced cell contact)
- Filter membranes (0.22 μm) for filter mating assays
Procedure:
Donor and Recipient Culture Preparation
- Grow donor E. coli overnight in LSLB with appropriate antibiotic selection at 37°C with shaking.
- Grow recipient biofilm cultures for 24-48 hours in suitable media to establish mature biofilms.
Liquid Conjugation Assay
- Mix donor and recipient cells at optimal ratios (typically 10:1 donor:recipient) in LSLB media.
- Add 0.5 mm glass beads to enhance cell-to-cell contact.
- Incubate at 37°C with mild agitation (60 RPM) for 24-72 hours.
Filter Mating Assay (Alternative Method)
- Mix donor and recipient cultures and concentrate by centrifugation.
- Resuspend in fresh media and apply to filter membranes placed on agar plates.
- Incubate for 4-24 hours to allow conjugation.
Selection and Analysis
- Plate conjugation mixtures on selective media containing antibiotics that distinguish transconjugants from donors and recipients.
- Induce Cas9 expression with arabinose to assess targeted killing efficiency.
- Quantify conjugation frequency (transconjugants/total recipients) and killing efficiency.
Biofilm Disruption Assessment
- Establish biofilms of target strains in appropriate flow cells or microtiter plates.
- Introduce donor strains with CRISPR-conjugative plasmids.
- Assess biofilm disruption through biomass quantification, microscopy, or viability staining.
Troubleshooting Notes:
- Optimize donor:recipient ratios for specific target species (test 1:1 to 50:1).
- Adjust NaCl concentration in media (lower concentrations may enhance conjugation).
- Modulate agitation speed—lower speeds may enhance contact but reduce mixing.
- For difficult-to-transform species, consider extended conjugation times or addition of conjugation-promoting factors.
Research Toolkit: Essential Reagents and Materials
Table 3: Research Reagent Solutions for Conjugative CRISPR-Biofilm Studies
| Reagent/Material | Function/Application | Specific Examples |
|---|---|---|
| Cis-Conjugative Plasmid System | Delivery of CRISPR components | pNuc-cis based on IncP RK2 system [14] |
| CRISPR Nuclease Variants | Targeted DNA cleavage | TevSpCas9, Cas9, Cas12a [13] [14] |
| Guide RNA Design | Target specificity | sgRNAs targeting essential genes, antibiotic resistance genes, or biofilm regulators [13] [14] |
| Specialized Growth Media | Enhanced conjugation efficiency | Low-salt LB (0.25% NaCl) [14] |
| Biofilm Promotion Surfaces | Mimic in vivo conditions | Glass beads, flow cells, preconditioned surfaces [2] [14] |
| Selection Antibiotics | Transconjugant identification | Chloramphenicol, tetracycline (plasmid-dependent) [14] |
| Induction Compounds | Controlled Cas9 expression | Arabinose for pBAD promoter systems [14] |
Quantitative Assessment of Conjugative CRISPR Efficacy
Table 4: Efficacy Metrics for Conjugative CRISPR Delivery Against Biofilms
| Parameter | Performance Metric | Experimental Conditions |
|---|---|---|
| Conjugation Frequency | Up to 100% with cis-system in liquid culture with beads [14] | E. coli to S. enterica, 72h, LSLB media, 0.5mm beads |
| Biofilm Reduction | >90% reduction in P. aeruginosa biofilm biomass with liposomal Cas9 [13] | In vitro biofilm models, lipid nanoparticle delivery |
| Gene Editing Efficiency | 3.5-fold increase with gold nanoparticle carriers [13] | Comparison to non-carrier delivery systems |
| Killing Efficiency | High efficiency with single or multiplexed sgRNAs targeting non-essential genes [14] | S. enterica, induced TevSpCas9 expression |
The multifaceted mechanisms of biofilm-mediated resistance, encompassing physical barrier protection, physiological heterogeneity, and persister cell formation, present significant challenges for conventional antibiotic therapies. The integration of conjugative plasmid delivery with CRISPR-based precision targeting represents a promising approach to overcome these challenges. By exploiting natural bacterial mating mechanisms to deliver targeted genetic interventions, this strategy offers the potential for species-specific pathogen control without disrupting commensal microbiota. Further optimization of delivery efficiency, host range, and safety profiles will be essential for translating these innovative approaches into clinically viable therapies for biofilm-associated infections.
The global crisis of antimicrobial resistance (AMR) demands innovative solutions that move beyond conventional antibiotics. Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) systems, originally identified as the adaptive immune system in prokaryotes, have emerged as transformative tools for sequence-specific antimicrobial therapy [16]. These systems offer a paradigm shift from broad-spectrum antimicrobials to programmable, precision strategies capable of targeting antibiotic-resistant pathogens while preserving beneficial microbiota [17]. This application note focuses specifically on conjugative plasmid delivery of CRISPR systems for biofilm control, a critical application within the broader antimicrobial field. Biofilms, which are structured microbial communities embedded in an extracellular polymeric substance (EPS), are responsible for approximately 65% of all microbial infections and exhibit dramatically enhanced tolerance to antimicrobial agents [6]. The integration of CRISPR technologies with advanced delivery platforms represents a promising approach for overcoming biofilm-associated treatment failures.
Molecular Mechanisms: From Bacterial Immunity to Programmable Tools
Natural Function and Classification
CRISPR-Cas systems provide adaptive immunity in bacteria and archaea through a three-stage process: adaptation, expression, and interference. During adaptation, Cas proteins capture fragments of invading nucleic acids and integrate them as "spacers" within the CRISPR array. In expression, these spacers are transcribed into CRISPR RNAs (crRNAs). Finally, during interference, crRNAs guide Cas effector complexes to recognize and cleave complementary foreign nucleic acids upon re-exposure [16]. The evolving diversity of these systems is reflected in the updated classification, which now includes 2 classes, 7 types, and 46 subtypes [18]. Class 1 systems (Types I, III, IV, and VII) utilize multi-protein effector complexes, while Class 2 systems (Types II, V, and VI) employ single-protein effectors such as Cas9, Cas12, and Cas13, making them particularly suitable for biotechnological applications [18] [17].
Reprogramming for Antimicrobial Applications
The reprogrammability of CRISPR-Cas systems enables two primary antimicrobial strategies: (1) targeted killing of resistant bacterial strains through cleavage of essential genes or chromosomal DNA, and (2) genetic inactivation of antimicrobial resistance genes to resensitize bacteria to conventional antibiotics [17]. The specificity of these interventions is determined by guide RNAs (gRNAs) designed to target unique genetic sequences associated with resistance determinants or essential bacterial functions, thereby avoiding collateral damage to commensal microorganisms [17].
Application Note: Conjugative Plasmid Delivery for Biofilm Control
Scientific Rationale and Strategic Advantage
Biofilms in food processing and healthcare environments present a formidable challenge due to their structural complexity, multispecies composition, and enhanced tolerance to disinfectants [9]. Conjugative plasmids offer an efficient delivery mechanism for CRISPR-Cas payloads to target bacteria within these complex communities. This approach leverages the natural bacterial mating processes for horizontal gene transfer, enabling widespread dissemination of antimicrobial constructs throughout biofilm architectures that often limit the penetration of conventional therapeutic agents [17]. Research demonstrates that conjugative plasmid-based delivery of CRISPR-Cas9 can selectively eradicate resistant Escherichia coli populations in mixed microbial communities, effectively reducing resistance gene prevalence without disrupting the surrounding microbiota [17].
Quantitative Efficacy Data
Table 1: Efficacy Metrics for CRISPR-Based Biofilm Control Strategies
| Intervention Strategy | Target System | Efficacy Outcome | Reference |
|---|---|---|---|
| Conjugative plasmid-delivered Cas9 | β-lactamase genes in E. coli | Selective elimination of resistant strains; preservation of commensals | [17] |
| Liposomal Cas9 formulations | Pseudomonas aeruginosa biofilm | >90% reduction in biofilm biomass in vitro | [6] |
| CRISPR-gold nanoparticle hybrids | Bacterial biofilms | 3.5-fold increase in gene-editing efficiency | [6] |
| CRISPRi targeting quorum sensing | E. coli urinary catheter biofilms | Significant reduction in biofilm formation | [9] |
| Phage-delivered CRISPR-Cas | Plasmids in Acinetobacter baumannii | Clearance of target pathogens in animal models | [17] |
Regulatory Pathways in Biofilm Formation
CRISPR-Cas systems not only serve as intervention tools but also participate in the natural regulation of biofilm formation. Recent research has revealed that Cas3 protein of the Type I-F CRISPR-Cas system in Acinetobacter baumannii functions as a repressor of virulence traits, with deletion leading to significantly enhanced biofilm formation, increased extracellular matrix production, and elevated epithelial colonization capacity [19]. This regulatory function is mediated through a hierarchical axis where transcriptional regulators H-NS and BaeR suppress Cas3 expression, thereby modulating biofilm phenotypes [19]. Conversely, in the Type I-Fa system, Cas3 deletion significantly reduces biofilm formation, virulence, and pathogenicity in mice, demonstrating subtype-specific regulatory roles [20]. These findings highlight the dual functionality of CRISPR-Cas systems as both genetic regulators and programmable antimicrobial tools.
Diagram 1: CRISPR-Cas Regulatory Network in Biofilm Formation. This diagram illustrates the hierarchical regulatory axis where BaeR positively regulates H-NS, which suppresses Cas3 expression. Cas3 subsequently inhibits biofilm formation and virulence factor production, including PNAG and pilus expression, ultimately modulating host adhesion capacity [19] [20].
Experimental Protocols
Protocol 1: Conjugative Plasmid Assembly for CRISPR-Cas Delivery
Objective: To construct a conjugative plasmid system for delivering CRISPR-Cas components to target biofilm-forming bacteria.
Materials:
- Donor strain: E. coli WM3064 or similar diaminopimelic acid (DAP) auxotroph
- Recipient strain: Target biofilm-forming bacteria (e.g., A. baumannii, P. aeruginosa)
- Conjugative plasmid backbone (e.g., pRPF185 or similar IncP-type plasmid)
- CRISPR expression cassette with appropriate promoters
- LB broth and agar plates with/without DAP (300 μM)
- Appropriate antibiotics for selection
Procedure:
- Clone the CRISPR-Cas system (Cas9 and sgRNA expression cassettes) into the conjugative plasmid backbone using Gibson assembly or traditional restriction-ligation.
- Transform the constructed plasmid into the donor E. coli strain via electroporation.
- Culture donor and recipient strains separately to mid-log phase (OD600 ≈ 0.5-0.6).
- Mix donor and recipient cells at a 1:2 ratio on a sterile filter placed on LB agar containing DAP.
- Incubate the mating filter for 8-12 hours at 30°C or 37°C depending on recipient requirements.
- Resuspend the mating mixture and plate on selective media without DAP but with antibiotics selective for the plasmid and counter-selective against the donor.
- Verify transconjugants by colony PCR and sequencing of the targeted genomic locus.
Technical Notes: Efficiency can be improved by using broad-host-range replicons and optimizing incubation times. For difficult strains, consider triparental mating with a helper plasmid providing mobilization functions [17].
Protocol 2: Assessment of Biofilm Disruption Efficacy
Objective: To quantitatively evaluate the impact of CRISPR-Cas delivery on pre-formed biofilms.
Materials:
- 96-well polystyrene microtiter plates
- Crystal violet solution (0.1%)
- Acetic acid (30%)
- Confocal laser scanning microscope (CLSM)
- SYTO9 and propidium iodide stains
- PCR reagents for resistance gene detection
Procedure:
- Grow biofilms of the target strain for 24-48 hours under appropriate conditions.
- Introduce the CRISPR-Cas conjugative system to mature biofilms via conjugation.
- Incubate for an additional 24 hours to allow gene editing.
- For biomass quantification: Stain with crystal violet, solubilize with acetic acid, and measure OD590.
- For viability assessment: Stain with SYTO9/propidium iodide and visualize via CLSM.
- For genetic verification: Harvest biofilm cells, extract genomic DNA, and perform PCR to confirm editing of target genes.
- Compare results to non-treated biofilms and biofilms treated with control plasmids.
Technical Notes: Combine with transcriptomic analysis (qRT-PCR) to verify downregulation of biofilm-related genes (e.g., ompA, csuA/B, pilA) [19] [20].
Diagram 2: Conjugative Plasmid Workflow for Biofilm Control. This experimental workflow outlines the key steps for delivering CRISPR-Cas systems via conjugation to target bacteria within biofilms, followed by selection and comprehensive analysis of biofilm disruption efficacy.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Conjugative CRISPR-Cas Biofilm Research
| Reagent/Category | Specific Examples | Function/Application | Technical Notes |
|---|---|---|---|
| Conjugative Plasmids | pRPF185, pKM101, R6K derivatives | Self-transmissible vectors for CRISPR delivery | Choose based on host range; include appropriate origin of transfer (oriT) |
| Cas Effector Variants | Cas9, dCas9, Cas12a, Cas13a | DNA/RNA targeting and editing or regulation | dCas9 for CRISPRi without cleavage; Cas12a for staggered cuts |
| Delivery Strains | E. coli WM3064 (DAP auxotroph) | Conjugation donor with plasmid maintenance | DAP auxotrophy enables counter-selection post-conjugation |
| Selection Markers | Kanamycin, Carbenicillin, Tetracycline resistance genes | Selection of transconjugants | Match antibiotic to recipient strain sensitivity profile |
| Biofilm Assay Kits | Crystal violet, SYTO9/propidium iodide, resazurin | Quantification of biofilm biomass and viability | Combine multiple methods for comprehensive assessment |
| gRNA Design Tools | CHOPCHOP, CRISPRdirect, Cas-OFFinder | In silico design and off-target prediction | Optimize for PAM compatibility and minimal off-target effects |
| Analysis Software | CRISPResso, ImageJ for biofilm analysis | Verification of editing efficiency and biofilm architecture | Use COMSTAT with ImageJ for biofilm structural metrics |
Conjugative plasmid delivery of CRISPR-Cas systems represents a promising strategy for precision biofilm control with potential applications in clinical, industrial, and environmental settings. The programmable nature of these systems enables selective targeting of resistance genes or virulence factors without disrupting beneficial microbiota—a significant advantage over broad-spectrum antimicrobials [9] [17]. Future developments will likely focus on optimizing delivery efficiency through engineered nanoparticles [6], enhancing specificity through improved gRNA design algorithms [21], and integrating artificial intelligence for predictive modeling of optimal gene targets [9]. As regulatory frameworks evolve and delivery challenges are addressed, CRISPR-based antimicrobials delivered via conjugative plasmids may transform our approach to managing persistent biofilm-associated infections and combating the global AMR crisis.
Bacterial conjugation, a process first discovered in 1946 by Joshua Lederberg and Edward Tatum, is a fundamental horizontal gene transfer mechanism that enables the direct cell-to-cell transfer of genetic material [22] [23] [24]. This natural delivery system involves the unidirectional transfer of DNA from a donor to a recipient bacterium through a specialized conjugative pilus, resulting in the recipient cell acquiring new genetic traits [22] [23]. Within bacterial populations, conjugation drives rapid evolution and adaptation by propagating advantageous properties including virulence factors, metabolic capabilities, and—most critically—antibiotic resistance genes [22].
The inherent efficiency and broad host range of conjugative systems present a powerful opportunity for biomedical applications. Recent innovative approaches have harnessed this natural DNA delivery mechanism to transport CRISPR-based antimicrobial systems for precise targeting of antibiotic-resistant pathogens and biofilms [25] [14] [15]. This application note explores the molecular mechanisms of bacterial conjugation and provides detailed protocols for leveraging conjugative plasmid systems to deliver CRISPR nucleases for targeted bacterial killing and biofilm control.
Molecular Mechanism of Bacterial Conjugation
The conjugation process is mediated by conjugative plasmids—extrachromosomal genetic elements that encode all necessary functions for their own transfer. The F (fertility) plasmid of Escherichia coli, approximately 100 kilobases in size, represents one of the best-characterized conjugative systems and serves as a paradigm for understanding conjugation mechanisms [22] [23] [26].
Key Steps in Conjugative DNA Transfer
The following diagram illustrates the core mechanism of plasmid transfer via bacterial conjugation:
Conjugation Mechanism Diagram Title: Conjugation DNA Transfer Process
The molecular process of conjugation proceeds through these key stages:
Conjugative Pilus Assembly and Cell Contact: Donor cells express transfer (tra) genes that encode the proteins necessary for forming a conjugative pilus—a hair-like appendage that projects from the donor cell surface [22] [23] [24]. The pilus recognizes and attaches to recipient cells, then retracts to establish direct membrane-to-membrane contact between donor and recipient cells [24].
Relaxosome Assembly: A protein complex called the relaxosome assembles at the origin of transfer (oriT) on the plasmid [22] [24]. In the F plasmid system, the relaxosome consists of the relaxase enzyme TraI, along with accessory proteins TraY, TraM, and the integrated host factor (IHF) [22] [23].
DNA Processing and Transfer: The relaxase enzyme creates a single-strand nick at the nic site within oriT [23] [24]. The nicked strand (T-strand) is unwound from its complementary strand and transferred in the 5' to 3' direction into the recipient cell through a channel formed by the type IV secretion system (T4SS) [22] [24]. A type IV coupling protein (T4CP) coordinates this process by coupling the relaxosome to the T4SS [22].
DNA Replication in Recipient Cell: Once the single-stranded DNA enters the recipient cell, the relaxase rejoins the DNA ends to regenerate a circular single-stranded plasmid molecule [24]. Both the donor and recipient cells then synthesize complementary strands to form double-stranded DNA through rolling circle replication, converting the recipient into a new donor cell capable of future conjugation events [23] [26].
Regulation of Transfer Gene Expression
The expression of tra genes is tightly regulated to minimize the fitness cost associated with conjugation machinery production [22]. In F-like plasmids, the primary activator TraJ promotes transcription of the main tra operon [22]. This activation is controlled by the FinOP fertility inhibition system, where FinP (an antisense RNA) and FinO (an RNA chaperone) repress TraJ translation at the post-transcriptional level [22]. Additionally, host-encoded factors like the histone-like nucleoid structuring protein (H-NS) silence tra promoters, making conjugation efficiency growth phase-dependent [22].
Conjugative Delivery of CRISPR Systems
The broad host range and high efficiency of conjugative plasmids make them ideal vectors for delivering CRISPR-based antimicrobial systems to target bacterial pathogens [25] [14]. The following diagram illustrates the strategic approach to designing conjugative CRISPR delivery systems for targeted bacterial killing:
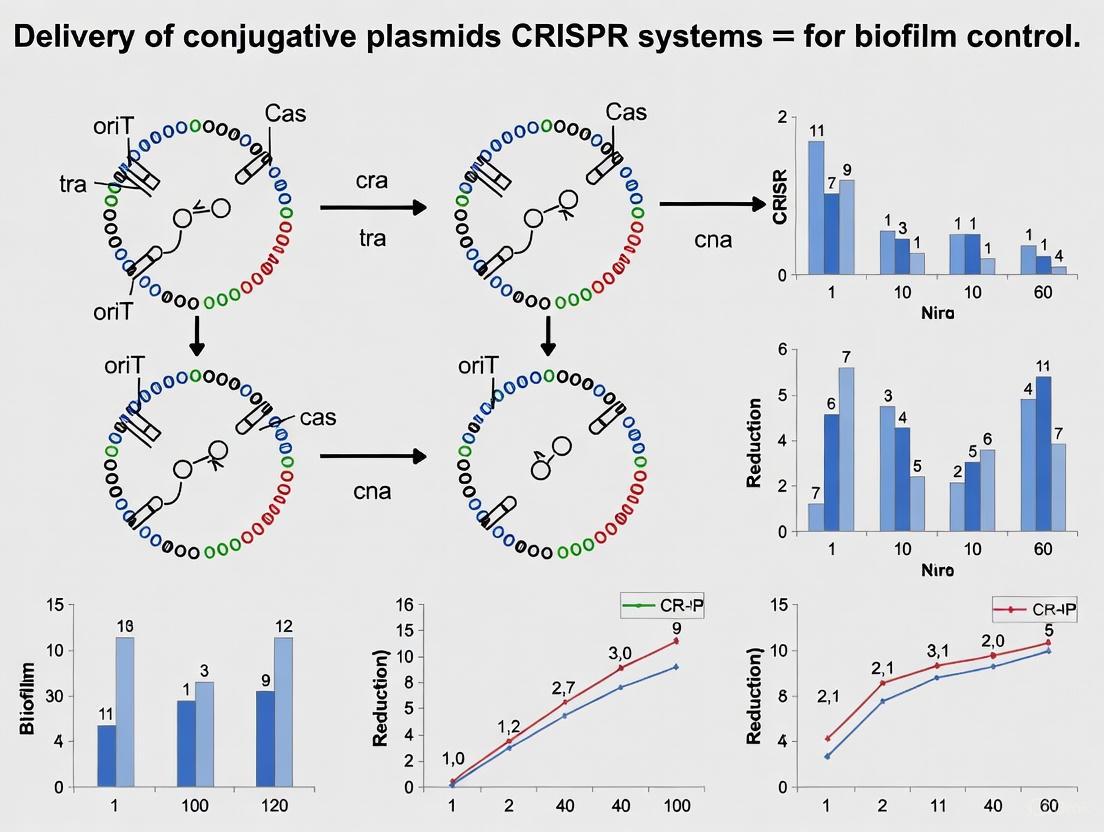
Conjugation CRISPR Strategy Diagram Title: CRISPR Delivery Design Strategy
Conjugative CRISPR Delivery Platforms
Two primary configurations have been developed for delivering CRISPR systems via conjugation:
Cis-Acting Conjugative Systems Cis-acting plasmids encode both the CRISPR nuclease machinery and the complete conjugation apparatus on a single plasmid backbone [14]. The IncP RK2-based pNuc-cis plasmid exemplifies this approach, containing the TevSpCas9 nuclease, guide RNA expression cassette, and all necessary tra genes for self-transfer [14]. This configuration enables extremely high conjugation frequencies—approaching 100% under optimized conditions—because each transconjugant becomes a potential donor for subsequent conjugation rounds, leading to exponential propagation throughout the bacterial population [14].
Trans-Acting Conjugative Systems Trans-acting systems separate the CRISPR payload and conjugation machinery onto different plasmids [14]. The CRISPR components are encoded on a mobilizable plasmid containing oriT, while the conjugation proteins are supplied in trans by a helper plasmid [14]. This configuration results in significantly lower conjugation frequencies (typically 10⁻⁵ to 10⁻³) compared to cis-acting systems because transconjugants cannot serve as donors for further conjugation [14].
Table 1: Comparison of Conjugative CRISPR Delivery Systems
| Feature | Cis-Acting System | Trans-Acting System |
|---|---|---|
| Configuration | Single plasmid encoding both CRISPR and conjugation machinery | Two plasmids: mobilizable CRISPR plasmid + helper conjugation plasmid |
| Conjugation Frequency | Up to 100% under optimal conditions [14] | Typically 10⁻⁵ to 10⁻³ [14] |
| Propagation in Population | Exponential (transconjugants become donors) | Limited to primary transconjugants |
| Delivery Efficiency | Liposomal Cas9 formulations reduce biofilm biomass by >90% [13] | Significantly lower efficiency |
| Key Advantage | Self-propagating through bacterial population | Reduced potential for unintended spread |
Quantitative Efficacy of Conjugative CRISPR Antimicrobials
Conjugative delivery of CRISPR nucleases has demonstrated remarkable efficacy in selectively eliminating targeted bacterial populations. The table below summarizes key performance metrics from recent studies:
Table 2: Efficacy Metrics of Conjugative CRISPR Antimicrobials
| Application | Target | Delivery System | Efficiency | Reference |
|---|---|---|---|---|
| Enterococcus faecalis killing | ermB and tetM resistance genes | Pheromone-responsive plasmid pPD1 | Significant reduction of antibiotic-resistant populations in murine intestine [25] | [25] |
| Salmonella enterica killing | Non-essential genes | IncP RK2 cis-conjugative system | High killing efficiency with single or multiplexed sgRNAs [14] | [14] |
| Biofilm disruption | P. aeruginosa biofilm genes | Liposomal Cas9 formulations | >90% reduction in biofilm biomass [13] | [13] |
| Gene editing enhancement | Various bacterial targets | Gold nanoparticle-CRISPR hybrids | 3.5-fold increase in editing efficiency [13] | [13] |
Experimental Protocols
Protocol: Conjugative Transfer of CRISPR Plasmids in Liquid Culture
This protocol describes a method for conjugative transfer of cis-acting CRISPR plasmids from E. coli to Salmonella enterica, adapted from [14] with modifications to enhance conjugation efficiency in biofilm-promoting conditions.
Materials Required
- Donor strain: E. coli harboring pNuc-cis or similar cis-conjugative CRISPR plasmid
- Recipient strain: Target pathogen (e.g., S. enterica, E. faecalis)
- Low-salt LB (LSLB) media: 1% tryptone, 0.5% yeast extract, 0.25% NaCl
- Antibiotics for selection
- 0.5 mm glass beads (sterile)
- 12-well cell culture plates
Procedure
- Culture Preparation
- Inoculate donor and recipient strains separately in 5 mL LSLB media with appropriate antibiotics.
- Incubate overnight at 37°C with shaking at 180 rpm.
Conjugation Setup
- Pellet 1 mL of each culture by centrifugation at 5,000 × g for 5 minutes.
- Resuspend cells in 1 mL fresh LSLB media without antibiotics.
- Mix donor and recipient cells at a 10:1 ratio in a final volume of 1 mL.
- Add 0.1 g of sterile 0.5 mm glass beads to each well of a 12-well plate.
- Add the cell mixture to the wells containing beads.
- Incubate statically at 37°C for 72 hours.
Transconjugant Selection
- Resuspend the conjugation mixture by pipetting.
- Prepare serial dilutions in phosphate-buffered saline (PBS).
- Plate appropriate dilutions on selective media containing antibiotics that select for transconjugants.
- Incubate plates at 37°C for 24-48 hours.
- Count transconjugant colonies and calculate conjugation frequency as transconjugants per recipient.
Notes
- The inclusion of glass beads provides surfaces for cell-to-cell contact, enhancing conjugation frequency up to 500-1000-fold compared to solution-based assays [14].
- Lower NaCl concentration (0.25%) in media further enhances conjugation efficiency [14].
- Static incubation or mild agitation (60 RPM) maximizes conjugation frequency compared to vigorous shaking [14].
Protocol: Assessment of CRISPR-Mediated Bacterial Killing
This protocol describes methods to quantify the efficacy of conjugatively delivered CRISPR nucleases in killing target bacteria.
Materials Required
- Transconjugants from Protocol 4.1
- Arabinose for induction of Cas9 expression
- Appropriate antibiotics
- Crystal violet solution (0.1%) for biofilm quantification
- Microtiter plates (96-well)
Procedure
- Killing Efficiency Assay
- Inoculate transconjugants in media with and without arabinose (0.2% final concentration) to induce Cas9 expression.
- Incubate at 37°C with shaking for 24 hours.
- Prepare serial dilutions and spot on non-selective agar plates.
- Compare colony forming units (CFUs) between induced and non-induced conditions.
- Calculate killing efficiency as: (1 - CFUinduced/CFUuninduced) × 100%.
- Biofilm Disruption Assay
- Inoculate transconjugants in 96-well plates with appropriate media.
- Induce Cas9 expression with arabinose.
- Incubate statically at 37°C for 48 hours.
- Remove planktonic cells and gently wash adhered cells with PBS.
- Fix biofilms with methanol for 15 minutes.
- Stain with 0.1% crystal violet for 20 minutes.
- Wash excess stain and solubilize bound stain with 30% acetic acid.
- Measure absorbance at 595 nm to quantify remaining biofilm biomass.
Notes
- Include controls with non-targeting sgRNAs to assess sequence-specificity of killing.
- For microscopy analysis, include fluorescent markers to visualize biofilm architecture disruption.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Conjugative CRISPR Delivery Studies
| Reagent/Cell Line | Key Features | Application/Function | Examples |
|---|---|---|---|
| Conjugative Plasmids | |||
| pNuc-cis plasmid | IncP RK2 backbone, encodes TevSpCas9 nuclease and conjugation machinery | Cis-acting conjugative CRISPR delivery [14] | [14] |
| pPD1 derivatives | Pheromone-responsive, narrow host range (E. faecalis) | Conjugative delivery in Enterococci [25] | [25] |
| Model Organisms | |||
| Escherichia coli | Common donor strain, well-characterized genetics | Conjugative plasmid propagation and delivery [14] | MG1655, S17-1 |
| Salmonella enterica | Gram-negative pathogen, model for infection studies | Recipient for conjugative CRISPR delivery [14] | SL1344, LT2 |
| Enterococcus faecalis | Gram-positive pathogen, hospital-acquired infections | Target for narrow host-range conjugation [25] | OG1RF, V583 |
| Specialized Media | |||
| Low-salt LB media | 0.25% NaCl concentration | Enhances conjugation frequency [14] | [14] |
| Brain Heart Infusion | Nutrient-rich medium for Gram-positives | Supports Enterococcus growth and conjugation [25] | [25] |
Bacterial conjugation represents a highly efficient natural mechanism for horizontal gene transfer that can be repurposed as a powerful delivery system for CRISPR-based antimicrobials. The protocols and applications detailed in this document provide researchers with practical methodologies to exploit conjugative systems for precise targeting of antibiotic-resistant pathogens and biofilms. Cis-acting conjugative plasmids that encode both CRISPR machinery and transfer functions demonstrate particularly high efficiency, enabling near-complete population penetration under optimized conditions that enhance cell-to-cell contact. As antibiotic resistance continues to pose severe threats to global health, conjugative delivery of CRISPR nucleases offers a promising precision antimicrobial strategy that can selectively eliminate targeted bacterial populations while preserving beneficial microbiota. Future directions in this field will likely focus on refining the specificity and control of these systems to enable translational applications in managing complex microbial communities.
The Rationale for Combining Conjugative Plasmids and CRISPR for Biofilm Eradication
Biofilms represent a significant challenge in both clinical and industrial settings, forming structured communities of microorganisms embedded in a protective extracellular polymeric substance (EPS) that confers high tolerance to antimicrobial agents and environmental stresses [9] [13]. The persistent nature of biofilms on medical devices and food processing surfaces leads to chronic infections and product contamination, with biofilm-related losses in the global agrifood sector alone estimated at approximately $324 billion annually [9]. Traditional broad-spectrum antimicrobials often fail to penetrate the biofilm matrix and disrupt these communities effectively, frequently selecting for tolerant strains and leaving persister cells intact [9].
The CRISPR-Cas system, originally identified as a bacterial adaptive immune mechanism, has emerged as a powerful platform for precision antimicrobial development [9] [27]. These systems can be programmed to target specific genetic sequences, enabling selective killing of pathogens, disruption of virulence genes, or elimination of antibiotic resistance determinants without affecting beneficial microbes [28] [29]. However, the efficacy of CRISPR-based antimicrobials depends critically on delivery systems that can efficiently transport the CRISPR components through protective biofilm matrices and into target bacterial cells [13] [28].
Conjugative plasmids offer a promising delivery vector for CRISPR antimicrobials due to their natural ability to transfer genetic material between bacterial cells through direct cell-to-cell contact [30] [14]. This review explores the scientific rationale, experimental evidence, and practical methodologies for harnessing conjugative plasmids to deliver CRISPR systems for precision biofilm control, providing researchers with a framework for developing next-generation antimicrobial strategies.
The Biofilm Challenge and Conventional Limitations
Structural and Functional Complexity of Biofilms
Biofilms exhibit a highly organized architecture characterized by microcolonies interspersed with water channels that facilitate nutrient distribution and waste removal [13]. The extracellular matrix, composed primarily of polysaccharides, proteins, and extracellular DNA (eDNA), forms a protective barrier that limits the penetration of antibiotics and plays a pivotal role in maintaining biofilm integrity and resilience [13]. This heterogeneous structure creates microenvironments where bacterial cells experience varying levels of nutrient availability, pH, oxygen, and waste products, contributing to survival under challenging conditions [13].
Bacteria within biofilms can exhibit up to 1000-fold greater tolerance to antibiotics compared to their planktonic counterparts [13]. This resistance arises through multiple mechanisms: reduced metabolic activity of persister cells, quorum sensing-regulated efflux systems, and the physical barrier posed by the EPS matrix that limits antibiotic penetration [13]. Additionally, the proximity of cells within biofilms enhances horizontal gene transfer, facilitating the spread of antibiotic resistance genes among community members [30].
Drawbacks of Conventional Antimicrobial Approaches
Traditional biofilm control strategies rely heavily on broad-spectrum disinfectants and antibiotics that provide non-specific antimicrobial activity but often fail to prevent recurrence [9]. These conventional approaches present significant limitations:
- Selection for tolerant strains through non-lethal exposure to antimicrobials
- Disruption of beneficial microbes and overall microbial ecology
- Inability to eliminate persister cells that remain dormant within the biofilm
- Acceleration of resistance development through horizontal gene transfer [9]
The inefficacy of these conventional methods has created an urgent need for precision antimicrobial strategies capable of selectively dismantling biofilms without compromising the overall microbial ecology [9].
CRISPR-Cas Systems: Precision Antimicrobial Tools
Mechanisms of CRISPR-Based Antimicrobial Activity
CRISPR-Cas systems employ Cas proteins and programmable RNA molecules to guide the recognition and cleavage of specific DNA or RNA sequences, permitting accurate genome editing and gene regulation [27]. These systems can be repurposed as antimicrobials through two primary mechanisms:
- Chromosomal Gene Inactivation: Programmed double-strand breaks in essential bacterial genes lead to replication fork collapse and cell death [14] [28].
- Plasmid Curing: Targeted elimination of plasmids encoding antibiotic resistance or virulence factors resensitizes bacteria to conventional antibiotics [28].
The specificity of the CRISPR-Cas approach relies upon the interaction between the Cas protein and a guide RNA (gRNA) sequence designed for targeting unique DNA sequences within the pathogenic target strain [28]. This precision enables selective killing of a particular bacterial member within a large population, which allows CRISPR-Cas antimicrobials to precisely modulate the composition of a complex bacterial population—a significant advantage over conventional antibiotics which tend to be broad spectrum without killing specificity [29].
CRISPR System Diversity and Selection
CRISPR-Cas systems are categorized into two classes based on their effector modules. Class 1 systems (types I, III, and IV) utilize multi-subunit effector complexes, while Class 2 systems (types II, V, and VI) employ a single multi-domain effector protein [27] [29]. The table below summarizes the primary CRISPR systems used for antimicrobial development:
Table 1: CRISPR-Cas Systems for Antimicrobial Applications
| System Type | Signature Protein | Mechanism of Action | Advantages for Antimicrobial Use |
|---|---|---|---|
| Type I | Cas3 | Creates long DNA fragments via nickase activity and recursive degradation [29] | Predominant in prokaryotes; highly efficient chromosomal degradation |
| Type II | Cas9 | Introduces double-strand breaks at specific DNA sites [28] | Well-characterized; versatile targeting; multiple delivery platforms |
| Type VI | Cas13 | Targets and cleaves RNA sequences [28] | Effective against RNA pathogens; collateral activity for detection |
Conjugative Plasmids: Ideal Delivery Vectors for Biofilm Environments
Natural Advantages for Biofilm Penetration
Conjugative plasmids possess inherent properties that make them particularly suitable for delivering CRISPR systems into biofilm communities:
- Broad Host Range: Many conjugative plasmids can transfer across taxonomic boundaries, enabling targeting of diverse bacterial species within multispecies biofilms [14] [28].
- Self-Replicating Capacity: Unlike phage vectors, plasmids maintain themselves episomally without immediate integration requirements [14].
- Resistance to Restriction: Plasmid-encoded modifications help evade bacterial restriction systems that often degrade foreign DNA [14].
- Biofilm Enhancement: Conjugative plasmids naturally promote biofilm formation to enhance cell-to-cell contact and transfer efficiency [14].
The conjugative process is particularly efficient in biofilm environments due to the dense, structured nature of these communities that facilitates cell-to-cell contact [14]. Studies have demonstrated that conditions enhancing cell-to-cell contact through biofilm formation can increase conjugation rates to nearly 100% [14].
Plasmid Engineering Strategies
The engineering of conjugative plasmids for CRISPR delivery involves two primary configurations:
- cis-Acting Systems: The plasmid encodes both the conjugative machinery and CRISPR system, enabling bacteria that receive the plasmid to become donors for subsequent conjugation rounds [14].
- trans-Acting Systems: The conjugation machinery and CRISPR components are encoded on separate DNA molecules, limiting transfer to a single conjugation event [14].
Research has demonstrated that cis-acting plasmids achieve significantly higher conjugation frequencies (up to 1000-fold greater than trans systems) because bacteria that receive the plasmid become potential donors for subsequent rounds of conjugation, leading to exponentially increasing numbers of conjugative donor bacteria in the population [14].
Quantitative Evidence: Efficacy of Conjugative CRISPR Delivery
Antimicrobial Performance Metrics
The combination of conjugative plasmids and CRISPR systems has demonstrated significant efficacy in biofilm eradication across multiple experimental models. The table below summarizes key quantitative findings from recent studies:
Table 2: Efficacy Metrics for Conjugative Plasmid-Delivered CRISPR Systems
| Target Organism | Delivery System | CRISPR System | Key Efficacy Findings | Reference |
|---|---|---|---|---|
| Salmonella enterica | IncP RK2 cis-conjugative plasmid | TevSpCas9 | Conjugation frequency up to 100% in biofilm conditions; high killing efficiency with targeted sgRNAs | [14] |
| Escherichia coli | CGV-EcCas conjugative vector | Type I-E CRISPR-Cas | Average reduction of 3.5 log10 CFU/mL; reduction of 1-6 log10 across E. coli panel | [31] |
| Escherichia coli | Engineered phage with plasmid elements | Type I-E CRISPR-Cas | Significant killing in biofilm conditions with PbolA promoter | [31] |
| Pseudomonas aeruginosa | Liposomal Cas9 formulations | Cas9 | >90% reduction in biofilm biomass in vitro | [13] |
| Multiple species | Gold nanoparticle carriers | Cas9 | 3.5-fold enhancement in editing efficiency compared to non-carrier systems | [13] |
Advantages Over Alternative Delivery Systems
While bacteriophages and nanoparticles represent alternative delivery mechanisms for CRISPR antimicrobials, conjugative plasmids offer distinct advantages:
- Superior Biofilm Penetration: Conjugative transfer efficiency increases in high-cell-density environments like biofilms, whereas phage diffusion may be limited by matrix barriers [14] [31].
- Broad Host Range: Many conjugative plasmids transfer across species boundaries more effectively than phages with narrow host specificity [14] [28].
- Continuous Propagation: cis-Acting conjugative systems create self-amplifying delivery through sequential rounds of conjugation [14].
- Large Cargo Capacity: Plasmids can accommodate multiple CRISPR components and regulatory elements in a single vector [14].
Experimental Protocols and Workflows
Protocol 1: Construction of cis-Acting Conjugative CRISPR Plasmids
This protocol outlines the methodology for creating efficient conjugative plasmids for CRISPR delivery, based on established systems such as the IncP RK2 plasmid [14].
Materials:
- Backbone vector with conjugative machinery (e.g., pTA-Mob)
- Cas nuclease gene with inducible promoter (e.g., pBAD-TevSpCas9)
- sgRNA expression cassette with constitutive promoter (e.g., pTet)
- Origin of transfer (oriT) sequence
- Restriction enzymes and ligation reagents
- E. coli donor strains and recipient biofilm-forming strains
Methodology:
- Vector Preparation: Digest the conjugative backbone vector with appropriate restriction enzymes to create compatible ends for insertion of CRISPR components.
- CRISPR Cassette Assembly: Clone the Cas nuclease gene under control of an inducible promoter (e.g., arabinose-inducible pBAD) into the prepared vector.
- sgRNA Integration: Insert the sgRNA expression cassette driven by a constitutive promoter (e.g., tetracycline resistance promoter, pTet) targeting specific biofilm-related genes.
- OriT Inclusion: Ensure the origin of transfer sequence is present and functional for conjugation machinery recognition.
- Transformation and Validation: Transform the constructed plasmid into donor E. coli strains and validate functionality through:
- Restriction mapping and sequence verification
- Cas protein expression analysis under inducing conditions
- sgRNA expression confirmation
- Conjugation efficiency testing
Critical Steps:
- Maintain the cis-configuration where conjugation machinery and CRISPR components reside on the same plasmid
- Select inducible promoters for Cas expression to prevent toxicity in donor strains
- Include appropriate selective markers for both donor and recipient selection
- Validate sgRNA specificity to minimize off-target effects in complex biofilm communities
Protocol 2: Biofilm Conjugation and Eradication Assay
This protocol describes the evaluation of conjugative CRISPR plasmid efficacy against established biofilms, adapted from established conjugation methodologies [14] [31].
Materials:
- Biofilm growth chambers (e.g., peg lids for 96-well plates, flow cells)
- Donor strains carrying conjugative CRISPR plasmids
- Biofilm-forming recipient strains
- Culture media with appropriate selective agents
- Confocal laser scanning microscopy (CLSM) equipment
- Viability staining reagents (e.g., LIVE/DEAD BacLight kit)
Methodology:
- Biofilm Establishment: Grow recipient biofilms for 48-72 hours under optimal conditions for the target species, with medium refreshment every 24 hours.
- Donor Preparation: Grow donor strains carrying conjugative CRISPR plasmids to mid-log phase (OD600 ≈ 0.5-0.6) under selective conditions.
- Conjugation Conditions:
- For liquid conjugation: Mix donors and biofilm recipients at optimal ratios (e.g., 10:1) in low-salt LB media with 0.5mm glass beads to enhance cell-to-cell contact
- Incubate at 37°C with mild agitation (60 RPM) for 24-72 hours
- CRISPR Induction: Add inducing agent (e.g., arabinose for pBAD promoter) at appropriate concentration to activate Cas expression.
- Efficacy Assessment:
- Viability counts: Perform serial dilution and plating on selective media to quantify transconjugants and surviving biofilm cells
- Biomass quantification: Use crystal violet staining or protein assays to measure total biofilm biomass
- Metabolic activity: Assess via ATP assays or tetrazolium reduction tests
- Visualization: Employ CLSM with viability staining to visualize spatial distribution of killing within biofilm architecture
Optimization Considerations:
- Test varying donor:recipient ratios (1:1 to 50:1) to determine optimal conjugation efficiency
- Evaluate different promoter systems (e.g., PbolA for biofilm conditions) to enhance expression in biofilm environments [31]
- Assess timing of induction relative to conjugation for maximal killing effect
- Include controls with empty vectors and non-targeting sgRNAs to distinguish specific CRISPR effects
Research Reagent Solutions
Table 3: Essential Research Reagents for Conjugative CRISPR-Biofilm Studies
| Reagent Category | Specific Examples | Function and Application | Key Considerations |
|---|---|---|---|
| Conjugative Backbones | IncP RK2, pTA-Mob, TP114, pPD1 | Provide conjugation machinery and broad host range | Select based on target species and cargo capacity requirements |
| CRISPR Effectors | TevSpCas9, Cas3, Cascade complex | Execute targeted DNA cleavage and degradation | Balance between size constraints and killing efficiency |
| Promoter Systems | pBAD (inducible), PbolA (biofilm-activated), pTet (constitutive) | Regulate temporal and spatial expression of Cas proteins | PbolA shows superior performance in biofilm conditions [31] |
| Biofilm Growth Systems | Peg lids, flow cells, microtiter plates | Provide standardized surfaces for biofilm development | Select based on analytical requirements and scalability needs |
| Analytical Tools | CLSM, viability stains, qPCR for transconjugants | Quantify conjugation efficiency and biofilm eradication | Combine multiple methods for comprehensive assessment |
Visualizing Workflows and Mechanisms
Conceptual Workflow for Conjugative CRISPR Biofilm Eradication
Experimental Protocol Visualization
The strategic combination of conjugative plasmids and CRISPR-Cas systems represents a paradigm shift in precision antimicrobial development for biofilm control. This approach directly addresses the fundamental limitations of conventional broad-spectrum antimicrobials by leveraging nature's own mechanisms for genetic transfer and bacterial immunity. The exceptional specificity of CRISPR targeting, combined with the efficient biofilm penetration enabled by conjugative transfer, creates a powerful synergy that enables selective eradication of target pathogens within complex microbial communities.
The experimental evidence demonstrates that properly engineered systems can achieve near-complete conjugation in biofilm environments and significant reductions (3-6 log10) in bacterial load through targeted genetic disruption. The self-amplifying nature of cis-acting conjugative plasmids further enhances their efficacy by creating continuously expanding populations of donor cells within the biofilm architecture.
As research in this field advances, future developments will likely focus on optimizing delivery efficiency, expanding host range, and implementing sophisticated regulatory controls to ensure precise spatial and temporal activation of CRISPR systems. With continued refinement, conjugative CRISPR delivery platforms hold exceptional promise for addressing some of the most challenging biofilm-associated infections in clinical, industrial, and environmental settings.
Engineering and Deploying Conjugative CRISPR Systems
Within the strategic framework of using conjugative plasmids to deliver CRISPR systems for biofilm control, the design of guide RNAs (gRNAs) is a foundational step that determines the success and specificity of the antimicrobial intervention. The programmable nature of CRISPR-Cas systems allows for precise targeting of bacterial genomic elements, enabling the disruption of biofilm integrity, the elimination of antibiotic resistance, and the inactivation of virulence mechanisms. This protocol details the key principles and methodologies for designing and validating effective gRNAs against three critical target categories: essential genes for bacterial lethality, antibiotic resistance determinants for resensitization, and virulence factors for pathogenicity attenuation. The application of these guidelines is particularly critical for conjugative delivery systems, where transfer efficiency and payload size must be balanced with therapeutic efficacy [13] [14].
The biological rationale for targeting these specific genetic elements stems from their distinct roles in bacterial persistence and pathogenesis. Essential genes encode functions indispensable for bacterial survival; their disruption leads to rapid cell death and effectively reduces biofilm biomass [13]. Antibiotic resistance genes, often carried on mobile genetic elements, can be selectively inactivated to restore antibiotic susceptibility in target pathogens [32]. Virulence factors contribute to pathogenicity and biofilm formation; their targeting can attenuate infection without directly causing bacterial death, potentially reducing selective pressure for resistance [13] [27]. When deployed via conjugative plasmids, CRISPR systems with specifically designed gRNAs can transform recipient bacteria into donors of these antimicrobial constructs, creating a propagating therapeutic effect within biofilm communities [14].
gRNA Design Principles and Strategic Considerations
Fundamental Design Parameters
The efficacy of CRISPR-based antimicrobial activity depends on careful consideration of several gRNA design parameters that influence binding efficiency, cleavage accuracy, and functional impact. The following factors must be addressed during the design phase:
Protospacer Adjacent Motif (PAM) Requirements: The PAM sequence is essential for Cas nuclease recognition and binding to target DNA. For the commonly used Streptococcus pyogenes Cas9 (SpCas9), the PAM sequence is 5'-NGG-3' located immediately downstream (3') of the target sequence [27] [32]. The PAM must be present in the target DNA but is not included in the gRNA sequence itself.
gRNA Length and Sequence Composition: The target-specific portion of the gRNA typically comprises 17-24 nucleotides with a GC content of 40-80% [27]. Higher GC content generally strengthens binding through enhanced complementary base pairing, while extremely high GC content may reduce specificity.
Target Strand Selection: The targeting efficiency can vary depending on whether the coding or template strand is selected, with some evidence suggesting preference for the non-template strand in certain systems [14].
Specificity Considerations: The gRNA sequence should be unique to the intended target to minimize off-target effects. This is particularly important when designing gRNAs for antimicrobial applications where non-specific cleavage could disrupt beneficial microbiota or lead to unintended genetic consequences [33].
Target Selection Strategy
Different target categories require distinct strategic approaches to maximize the desired phenotypic outcome:
Essential Genes: Select gRNAs targeting early exons of essential genes to maximize the probability of generating loss-of-function mutations through frameshifts or early stop codons [33]. Targeting multiple sites within the same essential gene can enhance lethality and reduce the likelihood of escape through compensatory mutations [14].
Antibiotic Resistance Determinants: Design gRNAs against conserved regions of resistance genes to enable broad targeting across bacterial strains and species [32]. For plasmid-borne resistance, target replication or maintenance genes in addition to the resistance cassette itself to facilitate plasmid clearance.
Virulence Factors: Identify functional domains critical for virulence factor activity and design gRNAs to disrupt these specific regions. For biofilm-specific virulence, target genes involved in quorum sensing, adhesion, or extracellular matrix production [13] [10].
Table 1: gRNA Design Specifications for Different Target Categories
| Target Category | Optimal gRNA Length | GC Content Range | Preferred Target Region | Special Considerations |
|---|---|---|---|---|
| Essential Genes | 20-22 nt | 45-65% | Early exons, catalytic domains | Multiplexing recommended to prevent escape mutants |
| Resistance Genes | 20 nt | 40-70% | Conserved catalytic sites, promoter regions | Target multiple alleles for broad-spectrum efficacy |
| Virulence Factors | 19-21 nt | 40-60% | Functional domains, secretion signals | Consider temporal expression patterns during infection |
Protocol: gRNA Design and Validation Workflow
In Silico Design and Specificity Analysis
Materials and Reagents:
- Reference genome sequences for target bacteria (NCBI, PATRIC)
- gRNA design software (Benchling, CHOPCHOP, CRISPy)
- Off-target prediction tools (Cas-OFFinder, BLAST)
Procedure:
Sequence Acquisition and Alignment:
- Obtain complete genome sequences for target bacterial strains from public databases (NCBI Genome, PATRIC).
- For resistance genes and virulence factors, compile representative sequences from multiple strains to identify conserved regions.
- Perform multiple sequence alignment using ClustalOmega or MUSCLE to identify conserved regions for broad-spectrum targeting.
gRNA Candidate Identification:
- Scan target genes for PAM sequences (5'-NGG-3' for SpCas9) using genome browser tools.
- Select 20-nucleotide sequences immediately 5' to each PAM as potential gRNA targets.
- Filter candidates based on GC content (40-70% optimal) and absence of secondary structure formation.
Specificity Validation:
- Perform genome-wide similarity searches using BLASTN against the host bacterium genome to identify potential off-target sites.
- Use specialized off-target prediction tools (Cas-OFFinder) that allow up to 3 nucleotide mismatches.
- Exclude gRNA candidates with significant homology to non-target genes, especially in conserved functional domains.
- For conjugative systems, verify absence of targets in potential recipient commensal bacteria.
Efficiency Prediction:
- Utilize machine learning-based prediction tools (DeepCRISPR, CRISPRscan) to rank gRNAs by predicted cleavage efficiency.
- Select 3-5 top candidates per target for empirical testing.
Table 2: Experimentally Validated gRNA Targets from Literature
| Target Category | Gene Example | gRNA Target Sequence (5'-3') | PAM | Efficiency | Application |
|---|---|---|---|---|---|
| Essential Gene | gyrA (S. enterica) | CTGATCATCGGTCGTACGAC | CGG | >99% killing | Bacterial elimination [14] |
| Resistance Gene | blaTEM (E. coli) | Not specified in results | GG | 2-3 log protection | AMR gene blockade [32] |
| Biofilm Gene | Quorum sensing (P. aeruginosa) | Not specified in results | GG | >90% biomass reduction | Biofilm disruption [13] |
Experimental Validation of gRNA Efficacy
Materials and Reagents:
- Conjugative plasmid backbone (e.g., IncP RK2-based system)
- Cas9 expression cassette (constitutive or inducible)
- gRNA cloning vector with appropriate promoters
- Donor and recipient bacterial strains
- Selective antibiotics for transconjugant selection
- Sanger sequencing capabilities
- ICE (Inference of CRISPR Edits) analysis tool [34] [35]
Procedure:
gRNA Cloning and Plasmid Construction:
- Clone selected gRNA sequences into the conjugative CRISPR plasmid under appropriate promoter control (e.g., constitutive pTet promoter [14]).
- For multiplexed targeting, clone multiple gRNA expression cassettes in tandem or utilize a single array with processing elements.
- Transform constructed plasmids into donor Escherichia coli strain.
Conjugative Transfer and Editing Assessment:
- Co-culture donor E. coli (carrying CRISPR conjugative plasmid) with recipient target bacteria at donor:recipient ratios between 1:1 to 50:1 [14].
- Use optimal conjugation conditions: low-salt LB media (0.25% NaCl), 37°C, mild agitation (60 RPM) or inclusion of solid surfaces (0.5mm glass beads) to enhance cell-to-cell contact.
- Allow conjugation for 24-72 hours to achieve high transfer frequency (approaching 100% with cis-conjugative systems).
- Select transconjugants using appropriate antibiotics.
Editing Efficiency Analysis:
- Isolate genomic DNA from transconjugants and amplify target regions using PCR.
- Perform Sanger sequencing of amplified products.
- Upload sequencing data to ICE (Inference of CRISPR Edits) tool along with gRNA sequence and nuclease information [34] [35].
- Calculate editing efficiency (Indel Percentage), functional knockout probability (Knockout Score), and model fit (R² score) from ICE output.
Phenotypic Validation:
- For essential gene targets: quantify bacterial killing through colony forming unit (CFU) counts before and after CRISPR induction.
- For resistance gene targets: perform antibiotic susceptibility testing to confirm resensitization.
- For virulence factor targets: conduct functional assays (adhesion, invasion, toxin production) relevant to the targeted virulence mechanism.
- For biofilm-related targets: quantify biofilm biomass using crystal violet staining or confocal microscopy after CRISPR treatment [13].
Research Reagent Solutions
Table 3: Essential Materials for gRNA Design and Validation
| Reagent/Resource | Function/Purpose | Examples/Specifications |
|---|---|---|
| Conjugative Plasmid Backbone | Delivery of CRISPR components to target bacteria | IncP RK2-based system; cis-configuration for high transfer efficiency [14] |
| Cas9 Nuclease Variants | DNA cleavage at target sites | TevSpCas9 dual nuclease; SpCas9 wildtype; high-fidelity variants [14] |
| gRNA Cloning Vector | gRNA expression and maintenance | Contains constitutive (pTet) or inducible promoters; multiple cloning sites for gRNA insertion [14] |
| Bioinformatics Tools | gRNA design and off-target prediction | CHOPCHOP, Benchling, Cas-OFFinder, BLASTN [33] |
| ICE Analysis Software | Quantification of editing efficiency from Sanger data | Synthego ICE; analyzes indels, calculates knockout scores, model fit R² [34] [35] |
| Delivery Optimization Reagents | Enhance conjugative transfer | Low-salt LB media; glass beads for surface attachment; varying agitation conditions [14] |
Workflow Visualization
gRNA Design and Validation Workflow
Technical Notes and Troubleshooting
Low Editing Efficiency: If ICE analysis reveals low indel percentages (<20%), consider optimizing gRNA length (increase to 22nt), verify PAM accessibility through chromatin analysis, or test alternative gRNA targets within the same gene [34] [35].
Poor Conjugative Transfer: For suboptimal plasmid transfer frequencies (<10^-3), implement conditions that enhance cell-to-cell contact including solid surface addition (glass beads), reduced agitation (0-60 RPM), and extended co-culture periods (up to 72 hours) [14].
Unexpected Off-target Effects: When observed, utilize more stringent off-target prediction parameters, consider high-fidelity Cas9 variants, or employ dual nuclease systems requiring two gRNAs for cleavage activity [33].
Incomplete Phenotypic Effect: For partial killing or resensitization, implement multiplexed gRNA strategies targeting multiple sites within the same gene or across redundant pathways to enhance efficacy and reduce escape frequency [14].
The protocols and design principles outlined herein provide a comprehensive framework for developing effective gRNA-based antimicrobials for delivery via conjugative plasmids. By adhering to these guidelines and utilizing the accompanying validation methodologies, researchers can create targeted interventions against biofilm-associated infections with enhanced precision and efficacy.
Conjugative plasmids are extrachromosomal DNA molecules that can transfer themselves between bacterial cells through a process called conjugation, a major horizontal gene transfer mechanism [22]. In the context of delivering CRISPR systems for biofilm control, understanding the design of these plasmid vectors—specifically whether their conjugation machinery is configured in cis (on the same plasmid) or trans (on separate elements)—is fundamental to predicting their transfer efficiency, host range, and stability within target populations [36]. Biofilms, which are structured microbial communities embedded in extracellular polymeric substances, pose a significant challenge in clinical and industrial settings due to their enhanced tolerance to antimicrobials [9] [13]. Conjugative delivery of CRISPR-based systems offers a promising approach to precisely target and disrupt biofilm-forming or antibiotic-resistant genes [13].
The F plasmid, first discovered in Escherichia coli, serves as the paradigm for conjugation studies in Gram-negative bacteria [22] [37]. The core components of a conjugative system include the Origin of Transfer (oriT), the Relaxosome complex that processes DNA at the oriT, and the Mating Pair Formation (Mpf) apparatus, which includes the type IV secretion system (T4SS) and the conjugative pilus [22]. This article delineates the design principles, applications, and experimental protocols for cis and trans configurations of these systems, with a specific focus on their use for delivering CRISPR countermeasures against bacterial biofilms.
Molecular Mechanisms of Conjugation
Key Steps in Plasmid Transfer
The conjugation process, as exemplified by the F plasmid, involves a series of coordinated steps [22]:
- Regulation of tra Gene Expression: The expression of transfer (tra) genes is tightly regulated. The key activator, TraJ, is itself regulated by the FinOP fertility inhibition system, comprising the antisense RNA FinP and the RNA chaperone FinO. Mutations, such as an IS3 insertion into finO in the F plasmid, create "superspreader" variants with constitutively high transfer rates [22] [37].
- Relaxosome Assembly: A complex of proteins (e.g., TraI, TraY, TraM in F) binds the oriT and nicks one strand of the plasmid DNA.
- Mating Pair Formation: The Mpf system, a Type IV Secretion System (T4SS), assembles a conjugative pilus that contacts a recipient cell. The pilus retracts to establish close cell-to-cell contact [22] [37].
- DNA Transfer and Replication: The nicked DNA strand is unwound and translocated into the recipient cell through the T4SS. Complementary strands are synthesized in both the donor and recipient cells, resulting in a functional plasmid in the transconjugant [22].
Defining Cis and Trans Configurations
The organization of the genetic determinants for conjugation defines the two main configurations:
- Cis Configuration: All necessary genes for conjugation—including the oriT, relaxosome components, and the entire Mpf system—are located on a single plasmid. This is the natural state of most conjugative plasmids, such as the F plasmid [22].
- Trans Configuration: The conjugation machinery is split across multiple genetic elements. A common setup involves a "helper" system, where the Mpf apparatus is provided in trans by a conjugative plasmid residing in the same donor cell, enabling the mobilization of a second, otherwise non-conjugative plasmid carrying the oriT [36].
Comparative Analysis: Cis vs. Trans Systems
The choice between a cis and a trans configuration has profound implications for the application of conjugative plasmids in biofilm research. The table below summarizes the core characteristics and trade-offs.
Table 1: Comparative analysis of cis and trans conjugative system configurations
| Feature | Cis Configuration (Single Plasmid) | Trans Configuration (Helper System) |
|---|---|---|
| System Complexity | All-in-one, self-sufficient system [22] | Multi-component system requiring coordination |
| Payload Capacity | Limited by plasmid size; large payloads may reduce transfer efficiency [36] | Large payloads possible on the mobilizable plasmid |
| Transfer Efficiency | High; optimized, coordinated expression of tra genes [22] | Variable; can be lower, depends on helper plasmid compatibility and copy number [36] |
| Host Range | Determined by the Mpf system of the plasmid (e.g., F-like plasmids in Enterobacteriaceae) [37] | Can be broadened by choosing a helper plasmid with a broad host range |
| Genetic Stability | High for the trait, as transfer genes and payload are linked | Risk of losing the helper plasmid or the mobilizable plasmid over generations |
| Flexibility & Design | Low; the system is fixed once engineered | High; modular, allows for "shuttle" vectors and specialized helper strains |
| Ideal Application | Efficient delivery of smaller CRISPR cassettes to a specific bacterial group | Delivery of large or complex genetic payloads, or when a broad host range is needed |
Application in CRISPR Delivery for Biofilm Control
Conjugative plasmids are ideal vectors for delivering CRISPR-based systems to bacterial populations within a biofilm, as they leverage a natural bacterial process for gene transfer. The protective extracellular matrix of biofilms can limit the penetration of conventional antibiotics and antimicrobials, but conjugative transfer can occur efficiently within these structured communities [22] [13].
Target Selection for Biofilm Disruption
CRISPR systems can be programmed to target specific genetic elements essential for biofilm integrity and persistence:
- Antibiotic Resistance Genes: Target and cleave plasmid-borne genes like bla (β-lactamase) or mcr-1 (colistin resistance) to resensitize bacteria to antibiotics [13].
- Quorum Sensing (QS) Pathways: Disrupt genes involved in the production or detection of autoinducers (e.g., luxS), impairing cell-cell communication and biofilm maturation [9].
- Biofilm Structural Genes: Target genes responsible for the production of adhesins, curli fibers, and extracellular polysaccharides (e.g., algD in P. aeruginosa) to dismantle the biofilm matrix [9].
- Conjugative Elements Themselves: A self-targeting strategy can use CRISPR to disrupt the conjugative plasmid's tra genes or oriT, potentially halting the spread of the delivery vector after its therapeutic function is complete [13].
Configuration Selection for Biofilm Applications
- Cis Systems are advantageous for creating a robust, "fire-and-forget" delivery vehicle. A single, well-engineered conjugative plasmid carrying a CRISPR-Cas9 system targeting a biofilm-specific gene can efficiently spread through a population from an initial, small number of donors.
- Trans Systems offer superior flexibility. A mobilizable plasmid can be designed to carry a large CRISPR array or multiple effector genes (e.g., Cas9 and an antimicrobial peptide). This mobilizable payload can then be delivered using different, specialized helper strains, allowing researchers to test conjugation across a wide range of conditions and recipient species without re-engineering the payload plasmid.
Experimental Protocols
Protocol 1: Testing Conjugation Efficiency in Biofilm vs. Planktonic Cultures
This protocol measures the transfer frequency of a conjugative plasmid in a controlled setting.
Research Reagent Solutions:
- Donor Strain: E. coli harboring the conjugative plasmid (e.g., a cis-configured F-plasmid derivative or a mobilizable plasmid with a helper system).
- Recipient Strain: A plasmid-free, antibiotic-sensitive strain, preferably a biofilm-forming clinical isolate, marked with a different selectable antibiotic resistance (e.g., rifampicin or kanamycin).
- Growth Media: Lysogeny Broth (LB) and minimal M9 media [38].
- Antibiotics: As required for selection of donors, recipients, and transconjugants.
- PBS (Phosphate Buffered Saline): For washing and diluting cells.
Procedure:
- Overnight Cultures: Grow donor and recipient strains separately in LB with appropriate antibiotics overnight at 37°C with shaking.
- Biofilm Setup:
- Mix donor and recipient cultures at a 1:1 ratio in a fresh, antibiotic-free medium.
- Dispense 2 mL aliquots into sterile 12-well polystyrene plates or flow cells.
- Incubate statically for 24-48 hours at 37°C to allow biofilm formation.
- Planktonic Mating:
- Mix donor and recipient cultures at a 1:1 ratio in a tube with fresh, antibiotic-free medium.
- Incubate with shaking for the same duration as the biofilm mating.
- Harvesting:
- Biofilm: Gently wash the biofilms twice with PBS to remove non-adherent cells. Add 1 mL PBS and scrape the biofilm from the well surface using a sterile pipette tip or cell scraper. Vortex vigorously to disaggregate the biofilm.
- Planktonic: Take 1 mL directly from the mating mixture.
- Enumeration and Calculation:
- Perform serial dilutions of the harvested cells in PBS.
- Plate appropriate dilutions on selective agar plates to count the number of donor cells, recipient cells, and transconjugants.
- Calculate the conjugation frequency as: Transconjugants / (Donors + Recipients) or as Transconjugants / Recipients.
Protocol 2: Assessing Biofilm Disruption Post-Conjugation
This protocol evaluates the functional outcome of delivering a CRISPR system targeting a biofilm-related gene.
Research Reagent Solutions:
- Transconjugant Strain: Recipient cells that have acquired the CRISPR-conjugative plasmid from Protocol 1.
- Control Strain: Recipient cells with a non-targeting control plasmid.
- Crystal Violet Solution (0.1%): For biofilm biomass staining.
- Acetic Acid (33%): To solubilize crystal violet.
Procedure:
- Strain Preparation: Isolate pure transconjugant and control strains using selective agar.
- Biofilm Growth: Grow the transconjugant and control strains in 96-well plates for 24-48 hours under conditions that promote biofilm formation.
- Crystal Violet Staining:
- Carefully remove the planktonic culture from the wells.
- Wash the wells gently with PBS to remove loosely attached cells.
- Add 125 µL of 0.1% crystal violet to each well and incubate for 15 minutes.
- Wash the wells thoroughly with water to remove excess stain.
- Add 125 µL of 33% acetic acid to solubilize the stain bound to the biofilm.
- Quantification: Transfer 100 µL of the solubilized crystal violet to a new 96-well plate and measure the absorbance at 550-600 nm using a plate reader. Higher absorbance correlates with greater biofilm biomass.
The Scientist's Toolkit
Table 2: Essential research reagents for conjugative plasmid and biofilm studies
| Research Reagent | Function & Application |
|---|---|
| F-like Plasmid (e.g., F, R1, R100) | Serves as a model cis-configured system or a helper plasmid; provides the Mpf machinery and oriT [22] [37]. |
| Mobilizable Plasmid with oriT | Core of a trans system; carries the CRISPR payload and can be mobilized by a helper plasmid providing transfer functions in trans [36]. |
| Broad-Host-Range Helper Plasmid | Expands the recipient range of a trans system beyond Enterobacteriaceae (e.g., using an IncP-type helper) [36]. |
| PhlF Repressor / PP7 System | A single-cell method using PhlF-RFP to label and count plasmid copies, and PP7-CFP to label RNA transcripts in living cells [38]. |
| Crystal Violet | A standard dye used to quantify total biofilm biomass adhered to abiotic surfaces [13]. |
| Liposomal CRISPR-Cas9 Formulations | Nanoparticle carriers that can enhance the delivery and efficacy of CRISPR components, potentially used in conjunction with conjugation for a combinatorial approach [13]. |
System Diagrams and Workflows
Cis vs. Trans System Architecture
Conjugative CRISPR Delivery Workflow
Regulation of tra Genes in F-like Plasmids
Broad-Host-Range IncP Plasmids and Other Delivery Vectors
Broad-host-range plasmids, particularly those of the IncP group, are indispensable tools in microbial biotechnology and environmental microbiology. Their capacity to transfer and replicate across a wide spectrum of bacterial species makes them exceptionally valuable for genetic engineering, especially in complex microbial communities. Within the specific research context of conjugative plasmid delivery of CRISPR systems for biofilm control, these vectors enable the precise introduction of antimicrobial agents into target organisms. This application note details the key plasmid systems, quantitative performance data, and standardized protocols for employing these vectors in experimental settings, providing researchers with a framework for investigating and developing novel biofilm control strategies.
Key Plasmid Delivery Systems and Their Performance
Broad-host-range vectors facilitate the introduction of genetic cargo, such as CRISPR-Cas systems, into diverse bacterial targets. The table below summarizes the characteristics and documented performance of major delivery systems.
Table 1: Key Broad-Host-Range Delivery Vectors for CRISPR System Delivery
| Vector Name | Backbone / Type | Key Features | Documented Experimental Performance |
|---|---|---|---|
| pNuc-cis [14] | IncP RK2 (cis-conjugative) | Encodes both conjugation machinery and CRISPR nuclease; enables secondary transmission. | Conjugation frequency from E. coli to S. enterica approached 100% in liquid culture with glass beads to enhance cell contact [14]. |
| pKJK5::csg [39] | IncP-1ε (conjugative) | Engineered to encode Cas9 and target-specific sgRNA; broad host-range. | Reduced transformation efficiency of a targeted AMR plasmid by ~4 orders of magnitude; conjugatively removed resident plasmids from E. coli, reducing plasmid-carrying recipients by ~2-fold [39]. |
| pTA-Mob 2.0 Derivatives [40] | Conjugative Plasmid | Superior variants with mutations (e.g., in traJ promoter) for enhanced transfer. | Demonstrated effective conjugation from bacteria to 7 diverse yeast species, including Candida auris [40]. |
| Three-Plasmid CRISPR-Cas9 [41] | Thermosensitive Shuttle Vectors | Utilizes pET194ts origin; constitutive P43 promoter for Cas9; for Bacillus velezensis. | Achieved 96% single-gene and 61% dual-gene editing efficiency in B. velezensis [41]. |
| CRISPR-Nanoparticle Hybrids [13] | Nanoparticle Carriers | e.g., Liposomal or gold nanoparticles; protect and enhance cellular uptake of CRISPR components. | Liposomal Cas9 reduced P. aeruginosa biofilm biomass by >90% in vitro; gold NPs increased gene-editing efficiency by 3.5-fold [13]. |
Experimental Protocols
Protocol: Conjugative Transfer of a Cis-Acting CRISPR Plasmid (e.g., pNuc-cis)
This protocol describes a filter mating assay for transferring a cis-acting conjugative plasmid from a donor to a recipient strain, adapted from published methodologies [14].
Materials:
- Donor strain (e.g., E. coli harboring pNuc-cis)
- Recipient strain (e.g., Salmonella enterica)
- Luria-Bertani (LB) broth and LB agar plates
- Appropriate antibiotics for selection
- Sterile nitrocellulose filters (0.45 µm pore size)
- Microfuge tubes
- Sterile forceps
Procedure:
- Culture Preparation: Grow donor and recipient strains separately in LB broth overnight with appropriate aeration and conditions.
- Cell Harvesting: Harvest bacterial cells by centrifuging 1 mL of each culture at 5,000-8,000 × g for 2 minutes. Gently resuspend the cell pellets in 1 mL of fresh LB broth to remove residual antibiotics.
- Mating Mixture: Combine donor and recipient cells at the desired ratio (e.g., 1:1 to 10:1 donor:recipient) in a microfuge tube. A total volume of 100-200 µL is typical.
- Filter Mating:
- Place a sterile nitrocellulose filter on an LB agar plate without antibiotics.
- Pipette the mating mixture onto the center of the filter and allow the liquid to be absorbed.
- Incubate the plate, right-side up, at the appropriate temperature (e.g., 37°C) for a defined period (e.g., 4-24 hours).
- Harvesting Transconjugants:
- After incubation, use sterile forceps to transfer the filter to a tube containing a known volume of sterile saline or LB broth.
- Vortex thoroughly to resuspend the cells from the filter.
- Selection and Enumeration:
- Plate serial dilutions of the resuspended cells onto LB agar plates containing antibiotics that selectively count the recipient strain (to determine total recipients) and the transconjugants (recipients that have acquired the plasmid).
- Incubate plates and count colonies to calculate conjugation frequency as (Number of Transconjugants / Total Number of Recipients).
Notes: For liquid conjugation, adding 0.5 mm glass beads to the medium can dramatically enhance cell-to-cell contact and conjugation frequency [14].
Protocol: Assessing CRISPR-Mediated Plasmid Removal or Blocking
This protocol uses a broad-host-range CRISPR delivery tool (e.g., pKJK5::csg) to either prevent the acquisition of an antimicrobial resistance (AMR) plasmid or to remove a resident AMR plasmid from a target strain [39].
Materials:
- Donor strain with mobilizable CRISPR plasmid (e.g., E. coli with pKJK5::csg[aacC1])
- Recipient strain with or without a target AMR plasmid (e.g., E. coli K12 with pHERD30T)
- LB broth and agar plates with appropriate antibiotics (e.g., Gentamicin, Trimethoprim)
- Sterile culture tubes and plates
Procedure: Part A: Blocking Plasmid Uptake
- Preparation: Introduce the targeting CRISPR plasmid (pKJK5::csg[aacC1]) and a non-targeting control (pKJK5::csg[nt]) into separate batches of the recipient strain (which lacks the target AMR plasmid) via conjugation or transformation.
- Transformation: Attempt to transform the targeted AMR plasmid (pHERD30T) into these prepared recipient strains using a standard transformation protocol.
- Analysis: Plate the transformation mixture on media selecting for the AMR plasmid (e.g., Gentamicin). A significant reduction (several orders of magnitude) in transformation efficiency for the strain with the targeting CRISPR plasmid compared to the control indicates successful blocking of plasmid uptake [39].
Part B: Removing a Resident Plasmid
- Liquid Mating:
- Grow donor (with pKJK5::csg[aacC1] or pKJK5::csg[nt]) and recipient (with resident AMR plasmid) strains separately overnight.
- Mix donor and recipient cultures in a defined ratio (e.g., 1:1) in LB broth and incubate for several hours to overnight with mild agitation.
- Selection and Analysis:
- Plate serial dilutions of the mating mixture onto selective media to enumerate:
- Total recipients (antibiotics for the recipient's chromosomal marker).
- Recipients that have acquired the CRISPR plasmid (antibiotics for the CRISPR plasmid).
- Recipients that have retained the target AMR plasmid (antibiotics for the AMR plasmid).
- Recipients that carry both plasmids (double selection).
- Plate serial dilutions of the mating mixture onto selective media to enumerate:
- Calculation: The proportion of recipients that have lost the AMR plasmid is significantly higher in matings with the targeting CRISPR donor compared to the non-targeting control [39].
Visualization of Experimental Workflows
The following diagrams outline the core logical relationships and experimental workflows for the protocols described above.
Conjugative Plasmid Workflow
CRISPR Plasmid Curing Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Conjugative Delivery of CRISPR Systems
| Reagent / Material | Function / Application | Specific Examples |
|---|---|---|
| IncP Plasmid Backbones | Broad-host-range conjugative delivery of genetic cargo. | pTA-Mob (for cis-conjugation) [14], pKJK5 (IncP-1ε) [39]. |
| CRISPR-Cas9 Components | Provides sequence-specific nuclease activity for gene editing or antimicrobial targeting. | Cas9 gene, sgRNA expression cassette [39]. |
| Fluorescent Reporter Genes | Visual tracking of plasmid transfer and gene expression in donor, recipient, and transconjugant cells. | Genes encoding GFP, mRuby2, or sfGFP [42] [39]. |
| Selective Antibiotics | Selection and maintenance of plasmids in bacterial cultures; enumeration of specific cell types post-experiment. | Trimethoprim, Gentamicin, Ampicillin, Chloramphenicol [41] [39]. |
| Microfluidic Chambers | Real-time, high-resolution imaging of conjugation dynamics and biofilm architecture. | Used for quasi-2D cell monolayers to study plasmid transfer [42]. |
| Glass Beads (0.5 mm) | Enhancement of cell-to-cell contact in liquid conjugation assays by providing a solid surface. | Increased conjugation frequency of pNuc-cis to nearly 100% [14]. |
Model Organisms and Co-culture Systems for Testing Efficacy
The rise of antimicrobial resistance (AMR) represents one of the most urgent threats to global health, with biofilm-associated infections being particularly challenging to treat due to their inherent resistance to conventional antibiotics [43] [44]. Biofilms are structured communities of microorganisms embedded in a self-produced extracellular polymeric substance (EPS) that can exhibit up to 1000-fold greater tolerance to antibiotics compared to their planktonic counterparts [6]. Within the broader thesis on conjugative plasmid delivery of CRISPR systems for biofilm control, this application note provides detailed protocols for utilizing model organisms and co-culture systems to test the efficacy of these innovative antimicrobial strategies. Conjugative plasmids have emerged as promising delivery vectors for CRISPR nucleases due to their broad-host range, resistance to restriction-modification systems, and capacity for large coding sequences [14]. This document outlines standardized methodologies for evaluating these systems in both simple and complex microbial communities, enabling researchers to assess bacterial killing efficiency, conjugation frequency, and biofilm disruption with quantitative precision.
Key Model Organisms for CRISPR Conjugative Delivery Studies
Selection Criteria and Representative Strains
The selection of appropriate model organisms is critical for evaluating conjugative CRISPR delivery systems. Ideal species are genetically tractable, relevant to human health, and capable of biofilm formation. The table below summarizes the primary model organisms used in this field.
Table 1: Key Model Organisms for CRISPR Conjugative Delivery Studies
| Organism | Relevance to AMR | Biofilm Formation Capability | Genetic Features | Key Applications |
|---|---|---|---|---|
| Escherichia coli | Common host for delivery plasmid propagation [14] | Forms biofilms on abiotic surfaces and in host tissues [43] | Well-characterized genetics; IncP RK2 conjugative system compatibility [14] | Donor strain for conjugation studies; Model for Gram-negative biofilm control |
| Salmonella enterica | Leading foodborne pathogen with emerging drug-resistant strains [14] | Robust biofilm former on epithelial surfaces and medical devices [14] | Susceptible to CRISPR targeting with designed sgRNAs [14] | Primary target for CRISPR-killing in co-culture systems; Model for host-pathogen interactions |
| Enterococcus faecalis | Leading cause of hospital-acquired infections (HAIs); multidrug-resistant strains prevalent [25] | Forms tenacious biofilms on catheters and surgical sites [43] [25] | Compatible with pheromone-responsive plasmids (PRPs); lacks native Cas9 in clinical strains [25] | Model for narrow-host-range conjugation; Target for ermB and tetM resistance gene removal |
| Staphylococcus aureus | Methicillin-resistant (MRSA) strains are major healthcare threat [44] | Strong biofilm producer associated with medical implant infections [45] | Susceptible to CRISPR targeting of resistance genes (e.g., mecA) [44] | Model for Gram-positive biofilm disruption; Assessment of combinatorial therapies |
These model organisms represent diverse bacterial families, resistance mechanisms, and biofilm-forming capabilities, allowing comprehensive evaluation of CRISPR conjugative systems across different microbial contexts.
Co-culture Systems for Efficacy Testing
Escherichia coli and Salmonella enterica Co-culture Model
The E. coli-S. enterica co-culture system represents a well-established model for studying inter-species conjugative transfer and targeted bacterial killing. This system allows researchers to simulate complex microbial environments where a CRISPR-carrying donor strain (E. coli) targets a pathogen recipient (S. enterica).
Quantitative Parameters and Experimental Outcomes
Table 2: Efficacy Parameters in E. coli-S. enterica Co-culture Models
| Parameter | Measurement Method | Reported Efficiency | Factors Influencing Outcome |
|---|---|---|---|
| Conjugation Frequency | Transconjugants per total recipients [14] | cis-plasmid: Up to 100% with enhanced cell contact; trans-plasmid: 10⁻⁸ to 10⁻⁴ [14] | Donor:recipient ratio; Culture conditions; Plasmid configuration (cis vs. trans) |
| Bacterial Killing Efficiency | Colony counts under induced vs. non-induced conditions [14] | Varies by sgRNA target: Essential genes >90% reduction; Non-essential genes variable [14] | sgRNA specificity and efficiency; Cas9 expression level; Target gene essentiality |
| Biofilm Disruption | Biomass reduction via microscopy or staining [45] | Liposomal Cas9 formulations: >90% reduction in P. aeruginosa biofilms [6] | Nanoparticle delivery system; Biofilm maturity; EPS composition |
| Resistance Gene Removal | Loss of antibiotic resistance in populations [25] | Several orders of magnitude reduction in resistant E. faecalis in murine intestine [25] | Guide RNA specificity; Conjugation efficiency; Selective pressure |
Enterococcus faecalis Model System
The E. faecalis system utilizes pheromone-responsive plasmids (PRPs) for highly efficient, species-specific conjugation. This model is particularly valuable for studying CRISPR delivery within Gram-positive bacteria and in intestinal environments.
Detailed Experimental Protocols
Protocol 1: Conjugative Transfer in E. coli-S. enterica Co-culture System
This protocol describes the procedure for assessing conjugative transfer of CRISPR-carrying plasmids from E. coli to S. enterica and subsequent targeted killing.
Materials and Reagents
Table 3: Essential Research Reagent Solutions
| Reagent/Equipment | Specification | Function/Application |
|---|---|---|
| Bacterial Strains | E. coli donor with pNuc-cis plasmid; S. enterica recipient with chromosomal target [14] | Conjugative transfer and CRISPR targeting assessment |
| Plasmid System | pNuc-cis with TevSpCas9, sgRNA cassette, and conjugative machinery [14] | All-in-one delivery of CRISPR and conjugation functions |
| Culture Media | Low-salt LB (LSLB) with 0.25% NaCl [14] | Enhanced conjugation efficiency through promoted cell contact |
| Induction Reagent | Arabinose for pBAD promoter induction [14] | Controlled Cas9 nuclease expression |
| Glass Beads | 0.5mm diameter [14] | Solid surface for enhanced cell-to-cell contact and biofilm formation |
| Selection Antibiotics | Chloramphenicol for plasmid maintenance [14] | Selection of transconjugants and donor strains |
| Confocal Microscope | Leica TCS SPE or equivalent [45] | Biofilm architecture and component analysis |
Procedure
Donor and Recipient Preparation
- Inoculate E. coli donor strain (carrying pNuc-cis plasmid with TevSpCas9 and sgRNA expression cassette) in LSLB media with appropriate antibiotic selection. Incubate overnight at 37°C with shaking at 120 RPM.
- In parallel, inoculate S. enterica recipient strain in LSLB media without antibiotics. Incubate overnight at 37°C with shaking at 120 RPM.
Co-culture Setup
- Mix donor and recipient cultures at a 10:1 ratio in fresh LSLB media supplemented with 0.5mm glass beads to enhance cell-to-cell contact.
- Include control conditions: donor alone, recipient alone, and co-culture with non-targeting sgRNA plasmid.
Conjugation Conditions
- Incubate co-culture at 37°C with mild agitation (60 RPM) for 72 hours to allow conjugative transfer.
- Maintain repressive conditions for Cas9 expression (0.2% glucose) during conjugation phase.
CRISPR Induction and Killing Assessment
- After 72 hours, induce TevSpCas9 expression by adding arabinose to a final concentration of 0.2%.
- Continue incubation for additional 24 hours to allow CRISPR-mediated killing.
Analysis and Quantification
- Serially dilute cultures and plate on selective media to quantify:
- Total viable recipients (antibiotic to count only S. enterica)
- Transconjugants (antibiotic that selects for plasmid-containing S. enterica)
- Donor cells (antibiotic that selects for E. coli with different resistance)
- Calculate conjugation frequency as: Transconjugants / Total Recipients
- Calculate killing efficiency as: 1 - (Transconjugants with targeting sgRNA / Transconjugants with non-targeting sgRNA)
- Serially dilute cultures and plate on selective media to quantify:
Protocol 2: Biofilm Disruption Assessment with Confocal Microscopy
This protocol details the quantification of biofilm components following CRISPR conjugative delivery, using multiple fluorescent staining reagents to assess different matrix elements.
Materials and Reagents
- Biofilm Growth Surface: 24-well plates with glass slides pre-coated with 10% poly-L-lysine [45]
- Fixation Solution: 4% formaldehyde with 0.5% Triton-X 100 [45]
- Fluorescent Stains:
- Imaging Equipment: Confocal laser scanning microscope (e.g., Leica TCS SPE) [45]
Procedure
Biofilm Formation
- Prepare bacterial suspension at 10⁸ CFU/mL in Tryptic Soy Broth (TSB).
- Inoculate into 24-multiwell plates containing poly-L-lysine coated glass slides.
- Incubate under orbital shaking (150 RPM) for 24 hours at 37°C to promote biofilm formation.
Treatment Application
- Wash established biofilms three times with PBS to remove non-adherent cells.
- Apply treatment conditions: CRISPR conjugative system vs. control.
- Incubate for additional 24 hours at 37°C.
Biofilm Fixation and Staining
- Wash biofilms three times with PBS and allow to air dry.
- Treat with 0.5% Triton-X-100 and 4% formaldehyde solution to disrupt and fix biofilms.
- Apply fluorescent staining reagents according to manufacturer recommendations:
- Sypro Ruby: 30 minutes incubation
- ConA-Alexa fluor 633: 60 minutes incubation
- GS-II-Alexa fluor 488: 45 minutes incubation
- PI: 15 minutes incubation
- TOTO-1: 30 minutes incubation
Image Acquisition and Analysis
- Examine stained biofilm samples using confocal fluorescence microscope.
- Measure biofilm depth at 4µm intervals along 80µm with 10× objective.
- Capture three fields per sample for statistical robustness.
- Process images using FIJI (Image J) software.
- Calculate biofilm component density as percentage of occupied area for each channel.
Table 4: Expected Biofilm Reduction After Effective CRISPR Treatment
| Biofilm Component | Staining Reagent | Control Occupied Area (%) | Treatment Occupied Area (%) | Expected Reduction (%) |
|---|---|---|---|---|
| Extracellular Proteins | Sypro Ruby | 17.58 ± 1.22 | 0.15 ± 0.01 | ≥99.2 [45] |
| α-Polysaccharides | ConA-Alexa fluor 633 | 16.34 ± 4.71 | 1.69 ± 0.69 | ≥89.7 [45] |
| α-β-N-acetylglucosamine | GS-II-Alexa fluor 488 | 16.77 ± 1.36 | 0.57 ± 0.28 | ≥96.6 [45] |
| Bacterial DNA | Propidium Iodide (PI) | 16.55 ± 13.42 | 1.60 ± 0.81 | ≥90.3 [45] |
| eDNA | TOTO-1 | 12.43 ± 6.23 | 0.07 ± 0.02 | ≥99.4 [45] |
Troubleshooting and Optimization Guidelines
Enhancing Conjugation Efficiency
- Low Conjugation Frequency: Increase donor:recipient ratio to 10:1 or higher; use low-salt LB media (0.25% NaCl); include solid surfaces (0.5mm glass beads) to enhance cell-to-cell contact [14].
- Poor Plasmid Transfer to Clinical Strains: Address restriction-modification barriers by using plasmid systems resistant to restriction enzymes or pre-modifying DNA to avoid recognition [25].
- Variable Killing Efficiency: Optimize sgRNA design by targeting essential genes; test multiple sgRNAs for each target; ensure proper induction of Cas9 expression with appropriate arabinose concentrations [14].
Standardizing Biofilm Assessment
- Inconsistent Biofilm Formation: Pre-coat surfaces with poly-L-lysine; standardize inoculum density (10⁸ CFU/mL); control agitation speed (150 RPM) during formation [45].
- High Variability in Staining: Follow manufacturer incubation times precisely; include appropriate controls for autofluorescence; use fresh staining solutions for each experiment.
- Image Analysis Challenges: Standardize microscope settings across samples; capture multiple fields per sample (minimum 3); use consistent thresholding parameters in FIJI/ImageJ [45].
Data Interpretation and Reporting Standards
When reporting results from these model systems, include the following key parameters:
- Conjugation frequency calculated as transconjugants per total recipients
- Killing efficiency relative to non-targeting controls
- Percentage reduction for each biofilm component compared to untreated controls
- Statistical significance (p-values) using appropriate tests (e.g., Mann-Whitney for non-normal distributions)
- Sample size and number of biological replicates
These standardized protocols for model organisms and co-culture systems provide robust frameworks for evaluating the efficacy of conjugative plasmid delivery of CRISPR systems for biofilm control, enabling direct comparison across studies and accelerating the development of novel antimicrobial strategies.
The escalating crisis of antibiotic-resistant biofilm-associated infections represents a critical challenge to global public health. Biofilms, structured communities of bacteria encased in an extracellular polymeric substance (EPS), demonstrate up to 1000-fold greater tolerance to antibiotics compared to their planktonic counterparts [13]. This resilience is mediated through dual mechanisms: phenotypic resistance, conferred by the physical and physiological barrier of the EPS, and heritable genetic resistance, driven by the acquisition of resistance genes via horizontal gene transfer within the biofilm matrix [13]. Conventional monotherapies often fail to overcome this multifaceted defense system, necessitating the development of innovative combinatorial approaches.
The CRISPR/Cas9 gene-editing system has emerged as a revolutionary tool for precision antimicrobial therapy. It offers the unique capability to directly target and disrupt the genetic foundations of antibiotic resistance and biofilm integrity [13]. However, the clinical translation of CRISPR-based antimicrobials is hampered by significant delivery challenges, particularly the inefficient penetration of biofilm matrices and the instability of genetic material en route to bacterial targets [13] [46].
This protocol details a synergistic strategy that integrates the genetic precision of CRISPR/Cas9 with the disruptive power of conventional antibiotics, enhanced by nanoparticle-mediated co-delivery. Nanoparticles serve a dual purpose: they act as sophisticated carriers that protect CRISPR components and ensure their targeted release within the biofilm, while also exhibiting intrinsic antibacterial and biofilm-penetrating properties [13]. By conjugating this delivery platform with conventional antibiotics, a powerful synergistic effect is achieved, simultaneously dismantling the biofilm's physical defenses and eradicating its underlying genetic resistance mechanisms. The following sections provide a detailed application note and standardized protocols for implementing this co-delivery strategy against robust biofilm-forming bacteria such as Pseudomonas aeruginosa.
Background and Rationale
The Biofilm Challenge and CRISPR/Cas9 Precision
Biofilms are highly organized microbial consortia embedded within a self-produced matrix of extracellular polymeric substances (EPS), including polysaccharides, proteins, and extracellular DNA (eDNA). This complex architecture creates heterogeneous microenvironments that limit antibiotic penetration and foster the emergence of metabolically dormant persister cells, which are highly tolerant to antimicrobials [13]. Furthermore, the dense, structured community of biofilms facilitates horizontal gene transfer, accelerating the dissemination of antibiotic resistance genes (ARGs) such as bla, mecA, and ndm-1 [13].
The CRISPR/Cas9 system can be programmed to introduce double-strand breaks in DNA, enabling the precise disruption of specific genetic elements. In the context of biofilms, key targets include:
- Antibiotic Resistance Genes (ARGs): Disarming bacteria by cleaving genes responsible for antibiotic inactivation, efflux, or target modification.
- Quorum Sensing (QS) Pathways: Impairing cell-to-cell communication, a critical process for biofilm maturation and virulence.
- Biofilm-Specific Regulators: Targeting genes responsible for EPS production, adhesion, and biofilm stability [13] [9].
Nanoparticles as Synergistic Carriers and Agents
Nanoparticles (NPs) provide an innovative solution to the central challenge of delivering CRISPR components into bacterial cells within a biofilm. Their small size and tunable surface chemistry allow for enhanced penetration through the EPS barrier. Different NP types offer distinct advantages [13]:
- Lipid-Based NPs: Facilitate fusion with bacterial membranes, enabling efficient delivery of CRISPR payloads.
- Metallic NPs (e.g., Gold NPs): Offer facile surface functionalization and can be engineered for controlled release; some also exhibit intrinsic antimicrobial properties.
- Polymeric NPs: Provide high stability and the capacity for large payloads.
Recent advances demonstrate the profound efficacy of this approach. For instance, liposomal formulations delivering CRISPR-Cas9 have been shown to reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro [13]. Similarly, CRISPR components complexed with gold nanoparticles demonstrated a 3.5-fold increase in gene-editing efficiency compared to delivery without a carrier system [13]. These platforms also enable the co-delivery of antibiotics or antimicrobial peptides, creating a multi-pronged attack that results in superior biofilm disruption and bacterial killing [13] [46].
Key Reagents and Equipment
Table 1: Essential Research Reagent Solutions for Co-delivery Experiments
| Item Name | Function/Description | Key Characteristics |
|---|---|---|
| dCas9/gRNA Ribonucleoprotein (RNP) | Core editing complex; dCas9 for CRISPRi/a or active Cas9 for cleavage. | Target-specific gRNA; purified complex for minimal off-target effects. |
| Cationic Liposomal Nanoparticles | Primary carrier for RNP and antibiotic co-delivery. | Positive surface charge for nucleic acid complexation; ~100 nm size for biofilm penetration. |
| Gold Nanoparticles (AuNPs) | Alternative carrier; allows conjugation via Au-thiol bonds. | Easy functionalization; intrinsic biofilm-disrupting properties. |
| Tobramycin | Aminoglycoside antibiotic for co-delivery against P. aeruginosa. | Synergizes with CRISPR-targeting of resistance genes (e.g., ampC). |
| Conjugative Plasmid Vector | Enables bacterial cell-to-cell delivery of CRISPR system. | Contains oriT origin of transfer; mobilizable into target pathogens. |
| CRISPRi/a System (dCas9) | For precise gene repression (CRISPRi) or activation (CRISPRa). | Allows reversible gene modulation without double-strand breaks. |
The following tables consolidate key quantitative findings from foundational studies employing CRISPR/nanoparticle co-delivery strategies.
Table 2: Efficacy of Nanoparticle-Mediated CRISPR Delivery Against Biofilms
| Nanoparticle Type | CRISPR Payload | Target Bacteria | Key Outcome Metric | Result | Citation |
|---|---|---|---|---|---|
| Liposomal Nanoparticles | Cas9/gRNA targeting pelA | Pseudomonas aeruginosa | Reduction in Biofilm Biomass | >90% reduction in vitro | [13] |
| Gold Nanoparticles (AuNPs) | Cas9/gRNA targeting ampC | Pseudomonas aeruginosa | Gene-Editing Efficiency | 3.5-fold increase vs. carrier-free system | [13] |
Table 3: Synergistic Effects of CRISPR-Antibiotic Co-Delivery
| Combination Therapy | Delivery Platform | Effect on Biofilm Viability | Synergy Metric | Notes | |
|---|---|---|---|---|---|
| CRISPR (targeting ndm-1) + Meropenem | Lipid Nanoparticles | >4-log reduction | FIC Index: <0.5 | Re-sensitized carbapenem-resistant strains | [46] |
| CRISPR (targeting qsrA) + Tobramycin | Conjugative Plasmids | ~99% cell death | 8-fold lower MIC for Tobramycin | Disrupted QS enhanced antibiotic uptake | [9] |
Detailed Experimental Protocols
Protocol 1: Co-delivery via Liposomal Nanoparticles
This protocol describes the formulation of cationic liposomal nanoparticles for the co-encapsulation of CRISPR-Cas9 ribonucleoprotein (RNP) and tobramycin.
I. Materials
- Cationic lipid (e.g., DOTAP), cholesterol, and PEG-lipid
- CRISPR-Cas9 RNP complex (pre-assembled)
- Tobramycin sulfate
- Saline buffer (10 mM HEPES, pH 7.4)
- Mini-extruder with 100 nm polycarbonate membranes
II. Step-by-Step Procedure
- Lipid Film Formation: Dissolve cationic lipid, cholesterol, and PEG-lipid (50:45:5 molar ratio) in chloroform in a round-bottom flask. Evaporate the solvent under reduced pressure using a rotary evaporator to form a thin, uniform lipid film.
- Hydration and Loading: Hydrate the dried lipid film with 1 mL of HEPES buffer containing the pre-complexed Cas9 RNP (500 nM) and tobramycin (1 mg/mL). Vortex vigorously for 5 minutes to form multilamellar vesicles.
- Size Reduction: Subject the lipid suspension to 10 freeze-thaw cycles (liquid nitrogen/40°C water bath). Subsequently, extrude the suspension 21 times through a polycarbonate membrane with a 100 nm pore size using a mini-extruder.
- Purification: Purify the formed liposomes from unencapsulated drugs and RNPs using a Sephadex G-50 size exclusion column. Elute with HEPES buffer.
- Characterization: Determine the particle size and zeta potential using dynamic light scattering (DLS). Measure the encapsulation efficiency of tobramycin via HPLC and of RNP via a fluorescence-based Ribogreen assay.
III. Application to Biofilm Assay
- Grow P. aeruginosa (e.g., strain PAO1) biofilms in a 96-well peg plate for 24-48 hours.
- Treat the mature biofilms with the formulated liposomes (diluted in fresh media) for another 24 hours.
- Assess biofilm biomass using the crystal violet staining method and quantify viable cells by sonicating the pegs and plating for colony-forming units (CFU).
Protocol 2: Conjugative Plasmid Delivery of CRISPRi System
This protocol outlines the design and deployment of a conjugative plasmid to deliver a CRISPR interference (CRISPRi) system for targeted gene repression in biofilm-forming bacteria.
I. Plasmid Construction
- Clone a dCas9 gene and a guide RNA (gRNA) targeting a specific gene (e.g., a quorum sensing regulator, lasR) into a broad-host-range, mobilizable plasmid vector (e.g., derived from RP4) containing an oriT site.
- The gRNA expression should be driven by a strong, constitutive bacterial promoter.
II. Conjugation Procedure
- Strain Preparation: Grow the donor strain (e.g., E. coli S17-1 λpir containing the conjugative plasmid) and the recipient biofilm-forming pathogen (e.g., P. aeruginosa) to mid-log phase (OD600 ~0.5).
- Mating: Mix donor and recipient cells at a 1:2 ratio on a sterile filter placed on a non-selective agar plate. Incubate for 6-8 hours at 37°C.
- Selection: Resuspend the cells from the filter in fresh medium and plate onto selective agar containing antibiotics that inhibit the donor strain and select for the recipient with the plasmid.
III. Biofilm Analysis
- Induce the CRISPRi system in the transconjugants and allow biofilms to form under selective conditions.
- Quantify gene repression efficacy via RT-qPCR of the target mRNA.
- Evaluate the impact on biofilm formation and antibiotic sensitivity by performing a minimum inhibitory concentration (MIC) assay in a biofilm mode and measuring EPS production.
Visualization of Workflows and Pathways
The following diagrams, generated with Graphviz DOT language, illustrate the core mechanisms and experimental workflows.
Diagram Title: Mechanism of Nanoparticle Co-delivery for Biofilm Eradication
Diagram Title: High-Throughput Screening Workflow for Synergistic Therapy
Overcoming Hurdles in Delivery Efficiency and Specificity
Conjugative plasmid delivery is a promising strategy for introducing CRISPR-based antimicrobial systems into bacterial populations, offering a sequence-specific method to combat antibiotic-resistant biofilms. A critical factor determining the success of this approach is the efficiency of plasmid transfer from donor to recipient cells, a process governed by physical cell-to-cell contact and profoundly influenced by the spatial organization of the bacterial community [47] [42]. This Application Note delineates the quantitative impact of cell density and biofilm maturation on conjugation frequency and provides detailed protocols for optimizing this process within the context of conjugative CRISPR delivery systems for biofilm control. Evidence demonstrates that while high cell density generally promotes conjugation, the complex architecture of mature biofilms can paradoxically hinder plasmid dissemination, creating a key challenge that these protocols aim to address [42].
Quantitative Impact of Cell Density and Biofilm Architecture
The data summarized in the table below reveal that conjugation frequency is maximized under conditions that promote high cell density and direct physical contact, yet is significantly restricted by the structural complexity of mature biofilms.
Table 1: Impact of Community Structure on Conjugative Transfer Efficiency
| Community Structure | Experimental Setup | Key Finding | Conjugation Frequency/ Efficiency |
|---|---|---|---|
| Liquid Culture (Low Cell Density) | LSLB media, 10:1 donor:recipient, 60 RPM [47] | Suboptimal frequency due to limited cell contact | ~1 x 10⁻⁵ |
| Filter Mating (Solid Surface) | 24-hour filter mating assay [47] | High-frequency transfer with a cis-conjugative plasmid | ~1 x 10⁻² |
| Liquid Culture with Beads | LSLB media with 0.5 mm glass beads [47] | Near-total conjugation due to biofilm-like conditions promoting contact | Approaches 100% |
| 2D Monolayer (High Density) | Microfluidic chamber, 1:10 donor:recipient [42] | Direct cell contact triggers efficient transfer; 92% of recipients become transconjugants | 92% ± 2% of contacted recipients |
| 3D Mature Biofilm | Pre-formed biofilm invaded by donors [42] | Limited donor penetration restricts transfer to periphery | Significantly hindered |
The data demonstrates a clear hierarchy of efficiency. Cis-conjugative plasmids, which encode their own conjugation machinery, are superior to trans-conjugative systems, showing a ~1000-fold higher conjugation frequency in filter mating assays because every new transconjugant becomes a potential donor, creating a chain reaction [47]. Furthermore, providing a solid surface for attachment, such as in filter mating or by adding glass beads to liquid culture, dramatically increases conjugation frequency by promoting stable cell-to-cell contacts that are essential for the conjugation machinery to function [47]. Crucially, the architecture of the bacterial community is a decisive factor. While high-density 2D monolayers support rapid and efficient plasmid propagation, the 3D structure of mature biofilms physically impedes donor cells from penetrating deeply, limiting conjugation primarily to the biofilm periphery [42].
Experimental Protocols
Protocol 1: Filter Mating Assay for Conjugation Efficiency
This protocol measures conjugation frequency under conditions that enhance cell-to-cell contact on a solid surface.
- Strain Preparation: Grow overnight cultures of the donor E. coli strain (e.g., carrying the pNuc-cis plasmid [47]) and the recipient Salmonella enterica strain in appropriate selective media.
- Cell Mixing and Filtration:
- Mix donor and recipient cells at a 1:1 ratio in a microcentrifuge tube. A 10:1 donor-to-recipient ratio can also be used for higher frequencies [47].
- Pellet the mixed culture and resuspend in a non-selective, low-salt LB (LSLB) medium (e.g., 0.25% NaCl) to promote contact [47].
- Transfer the cell suspension onto a sterile membrane filter (e.g., 0.22 µm pore size) using a vacuum filtration apparatus.
- Incubation for Conjugation:
- Place the filter on a pre-warmed non-selective LB agar plate.
- Incubate the plate at 37°C for a defined period (e.g., 2-24 hours) to allow conjugation.
- Harvesting and Plating:
- After incubation, transfer the filter to a tube containing a known volume of sterile saline or buffer.
- Vortex thoroughly to resuspend the cells from the filter.
- Perform serial dilutions and plate onto selective agar plates that permit growth only of transconjugants (e.g., containing antibiotics that select for the plasmid and the recipient strain's chromosomal markers).
- Calculation:
- Conjugation frequency is calculated as the number of transconjugants (CFU/mL) divided by the total number of recipient cells (CFU/mL).
Protocol 2: Assessing Conjugation Dynamics in a 3D Biofilm
This protocol uses live-cell microscopy to visualize plasmid invasion in pre-formed biofilms, revealing spatial constraints.
- Biofilm Growth:
- Grow a recipient strain (e.g., E. coli expressing a chromosomal red fluorescent protein, mRuby2 [42]) in a flow cell or microfluidic chamber with a suitable growth medium for 24-48 hours to form a mature 3D biofilm.
- Donor Invasion:
- Introduce the donor strain (e.g., carrying an RP4 plasmid with a constitutively expressed sfGFP [42]) in fresh medium into the flow cell containing the pre-formed biofilm.
- Time-Lapse Imaging:
- Use a confocal laser scanning microscope (CLSM) to acquire Z-stack images of the biofilm at regular intervals (e.g., every 15-30 minutes) over several hours.
- Use appropriate laser lines to excite the donor (GFP, green) and recipient/transconjugant (RFP, red) fluorophores.
- Image and Data Analysis:
- Use software tools like BiofilmQ and StarDistOPP to quantitatively analyze the images [42].
- Quantify the spatial distribution of donors and transconjugants (which appear yellow in overlay images) over time.
- Measure the depth of donor penetration and the percentage of transconjugants formed in the biofilm core versus the periphery.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Conjugation and Biofilm Studies
| Reagent / Material | Function / Description | Example & Rationale |
|---|---|---|
| Cis-Conjugative Plasmid | A plasmid encoding both CRISPR machinery and conjugation functions. | pNuc-cis [47]: Based on IncP RK2 system, enables high-frequency transfer as transconjugants become new donors. |
| Fluorescent Reporter Strains | Genetically modified donor/recipient strains for visualization. | RP4-sfGFP donors and mRuby2 recipients [42]: Allow real-time, single-cell tracking of plasmid transfer in biofilms. |
| Microfluidic Flow Cell | Device for growing biofilms under controlled fluid dynamics for microscopy. | Enables high-resolution imaging of 3D biofilm structure and plasmid invasion dynamics [42]. |
| Biofilm Image Analysis Software | Computational tools for quantifying 3D biofilm architecture and cell locations. | BiofilmQ & StarDistOPP [42]: Extract quantitative data on cell number, spatial distribution, and fluorescence from CLSM images. |
| Low-Salt LB Media | Culture medium with reduced ionic strength to enhance cell aggregation. | LSLB (0.25% NaCl) [47]: Promotes cell-to-cell contact, significantly boosting conjugation frequency in liquid assays. |
Optimizing conjugation frequency requires a nuanced understanding of bacterial community organization. The protocols and data presented here provide a framework for enhancing the delivery of CRISPR systems via conjugation. The key is to leverage conditions that mimic the high cell density of early-stage biofilms or microcolonies, while developing strategies to overcome the physical diffusion barrier presented by the extracellular matrix of mature biofilms. Mastering these parameters is essential for advancing conjugative plasmid delivery as a viable and potent strategy for precision biofilm control.
Strategies to Promote Donor-Recipient Contact in 3D Biofilm Structures
The efficacy of conjugative plasmid delivery systems for biofilm control is fundamentally constrained by the physical architecture of biofilms, which limits direct cell-to-cell contact between donor and recipient bacteria. Conjugative transfer of plasmids, including those carrying CRISPR-Cas systems, requires direct contact between donor and recipient cells [42]. However, the three-dimensional structure of mature biofilms presents a significant barrier, as their emergent community architecture limits the ability of donor cells to enter regions with high cell density, thereby hindering the establishment of productive contacts for plasmid transfer [42]. This application note details practical strategies to overcome these physical constraints, with particular emphasis on enhancing contact for conjugative delivery of CRISPR-based antimicrobials. We present optimized protocols and quantitative frameworks to maximize plasmid dissemination in biofilm environments for research and therapeutic development.
The Biofilm Conjugation Challenge
Architectural Barriers to Plasmid Transfer in Biofilms
Biofilms are surface-attached, spatially structured communities embedded in a self-produced polymeric extracellular matrix that significantly influences conjugation efficiency [42]. While high bacterial density within biofilms was historically presumed to enhance conjugation, recent evidence demonstrates that the 3D architecture of mature biofilms actually imposes significant physical constraints on plasmid transfer. Key challenges include:
- Limited donor penetration: Donor cells experience restricted access to high-density regions in mature biofilms [42]
- Matrix barrier function: Extracellular polymeric substances (EPS) composed of polysaccharides, proteins, and DNA create a protective barrier that limits cellular mobility and contact [48] [6]
- Spatial heterogeneity: The formation of microcolonies interspersed with water channels creates uneven distribution patterns for donor cells [6]
The timing of donor introduction relative to biofilm development critically impacts conjugation success. Studies with the broad host range IncP-1α RP4 plasmid demonstrate that early-stage biofilms and microcolonies with lower surface coverage provide open access points for donor cells to regions that later become high-cell-density areas in mature biofilms, thereby facilitating significantly more efficient plasmid transfer compared to established mature biofilms [42].
Quantitative Impact of Biofilm Architecture on Conjugation
Table 1: Conjugation Efficiency Across Different Biofilm Architectures
| Biofilm Architecture | Relative Conjugation Efficiency | Key Limiting Factors | Optimal Intervention Points |
|---|---|---|---|
| Mature 3D Biofilms | Severely limited | Restricted donor penetration, dense EPS matrix, spatial segregation | Matrix disruption, donor engineering |
| Early-Stage Biofilms/Microcolonies | High | Limited surface coverage, developing architecture | Early donor introduction, surface modification |
| High-Density 2D Monolayers | Optimal (Reference) | Minimal physical barriers | Not applicable - ideal reference condition |
| Mixed-Species Biofilms | Variable | Interspecies interactions, niche competition | Community engineering, broad-host-range vectors |
Strategic Approaches to Enhance Donor-Recipient Contact
Biofilm Growth Stage Manipulation
The developmental stage of a biofilm significantly determines its accessibility to donor cells. Research demonstrates that microcolonies and early-stage biofilms permit substantially greater donor access compared to mature structures [42]. Strategic approaches include:
- Early donor introduction: Introducing donor cells during initial attachment or microcolony formation phases, when biofilm architecture is less developed and physical barriers are minimal
- Developmental synchronization: Coordinating donor-recipient co-culture initiation to ensure simultaneous presence during optimal conjugation windows
- Biofilm maturation monitoring: Using established benchmarks such as the 48-hour maturation model for Staphylococcus epidermidis to time interventions [48]
Matrix-Targeted Disruption Strategies
The extracellular matrix represents a primary physical barrier to donor-recipient contact. Effective disruption strategies include:
- Enzymatic matrix degradation: Using glycoside hydrolases to target polysaccharide components or DNAse I to degrade extracellular DNA (eDNA), both universal biofilm matrix constituents [49]
- Dispersion induction: Triggering native biofilm dispersal mechanisms through compounds like nitric oxide (NO) and cis-2-decenoic acid, which reduce intracellular cyclic di-GMP levels and promote transition to planktonic state [49]
- EPS-targeted nanoparticles: Employing functionalized nanoparticles that penetrate and disrupt the matrix structure while potentially carrying therapeutic payloads [6]
Conjugative Plasmid Engineering
Engineering conjugation systems to enhance transfer efficiency represents a powerful approach:
- Cis-acting conjugative plasmids: Utilizing systems where conjugation machinery and payload (e.g., CRISPR-Cas) are encoded on the same plasmid, enabling recipients to become donors and exponentially increase transfer rates [14]
- Broad-host-range systems: Implementing IncP RK2-based plasmids that transfer efficiently across species boundaries [14]
- Pheromone-responsive plasmids: For species like Enterococcus faecalis, exploiting naturally high-transfer systems that comprehensively infiltrate target populations [25]
Table 2: Engineered Conjugative Plasmid Systems for Enhanced Biofilm Transfer
| Plasmid System | Transfer Mechanism | Advantages for Biofilm Applications | Demonstrated Efficacy |
|---|---|---|---|
| Cis-Acting (pNuc-cis) | Single plasmid encodes both conjugation machinery and payload | Recipients become donors; exponential spread; ~1000x higher frequency than trans systems [14] | Approaching 100% conjugation frequency in optimized conditions [14] |
| Pheromone-Responsive (pPD1) | Responds to recipient pheromones; narrow host range | High specificity; efficient intestinal colonization; bacteriocin-enhanced maintenance [25] | Significant reduction of antibiotic-resistant E. faecalis in murine intestine [25] |
| Trans-Acting Systems | Conjugation machinery and payload on separate plasmids | Modular design; easier engineering | Limited by low transfer frequency; not optimal for biofilms [14] |
Experimental Protocols
Microfluidic Biofilm Conjugation Assay
This protocol enables real-time, single-cell quantification of conjugation dynamics in controlled biofilm environments, based on methodology from live-cell fluorescence microscopy studies of RP4 plasmid transfer [42].
Materials and Reagents
Table 3: Essential Research Reagent Solutions
| Reagent/Cell Line | Function/Application | Key Characteristics |
|---|---|---|
| E. coli RP4-sfGFP Donor | Conjugative donor strain | Constitutively expresses sfGFP; contains broad host range IncP-1α RP4 plasmid with fluorescent reporter [42] |
| E. coli mRuby2 Recipient | Conjugation recipient | Chromosomal mRuby2 expression (red fluorescence); enables transconjugant detection (yellow in overlay) [42] |
| Ssb-Ypet Reporter | Visualizes ssDNA transfer | Membrane-associated conjugative foci indicate single-stranded DNA transfer during conjugation [42] |
| mCherry-ParB/parS System | Detects dsDNA conversion in transconjugants | Binds double-stranded parS DNA sites on RP4 plasmid; confirms successful plasmid establishment [42] |
| Quasi-2D Microfluidic Chambers | Biofilm growth platform | Enables high-density cell monolayer formation; ideal for real-time microscopy of conjugation events [42] |
| Low-Salt LB (LSLB) Media | Enhanced conjugation medium | 0.25% NaCl w/v; significantly increases conjugation frequency in liquid assays [14] |
Procedure
- Strain preparation: Grow donor and recipient strains overnight in appropriate selective media.
- Cell loading: Mix donor and recipient cells at 1:10 ratio in fresh medium and introduce into microfluidic chambers.
- Monolayer formation: Allow cells to attach and form high-density monolayers under continuous medium flow (0.5-1.0 μL/min).
- Time-lapse imaging: Acquire images at 1-minute intervals using fluorescence microscopy with appropriate filter sets.
- Conjugation quantification:
- Detect Ssb-Ypet foci to identify ongoing ssDNA transfer events
- Monitor mCherry-ParB focus formation to confirm dsDNA conversion
- Track sfGFP fluorescence increase in recipient cells to identify transconjugants
Data Analysis
- Calculate plasmid transfer rate based on Ssb-Ypet focus lifespan (approximately 1.8±0.8 minutes for 60kb RP4 plasmid)
- Determine ss-to-dsDNA conversion time (approximately 1.4±0.7 minutes)
- Quantify percentage of donor-recipient contacts resulting in productive conjugation (approximately 92±2% within 20 minutes post-contact)
Enhanced Liquid Conjugation Assay with Surface Contact
This protocol adapts filter mating principles to liquid culture with surface enhancement, achieving near-100% conjugation frequency with cis-acting plasmids [14].
Materials and Reagents
- Donor and recipient strains (e.g., E. coli donor with S. enterica recipient)
- Cis-acting conjugative plasmid (e.g., pNuc-cis with CRISPR payload)
- Low-salt LB media (0.25% NaCl w/v)
- Sterile glass beads (0.5mm diameter)
- Orbital shaker with temperature control
Procedure
- Culture preparation: Grow donor and recipient strains to mid-log phase (OD600 ≈ 0.4-0.6).
- Cell mixing: Combine donor and recipient at 10:1 ratio in LSLB media.
- Surface enhancement: Add sterile glass beads (approximately 20% volume) to provide attachment surfaces.
- Incubation conditions: Incubate at 37°C with mild agitation (60 RPM) for 24-72 hours.
- Transconjugant selection: Plate serial dilutions on selective media to quantify transconjugants, donors, and recipients.
- Conjugation frequency calculation: Express as transconjugants per recipient.
Optimization Notes
- Donor:Recipient ratio: 10:1 optimal for most applications [14]
- Agitation: Mild agitation (60 RPM) superior to static conditions or high agitation (120 RPM)
- Time course: Conjugation frequency increases over 72 hours with cis-acting systems
- Salt concentration: Reduced NaCl (0.25%) significantly enhances conjugation frequency
Biofilm Dispersion-Enhanced Conjugation Protocol
This approach utilizes biofilm dispersion triggers to temporarily increase planktonic cell populations and enhance conjugation opportunities [49].
Materials
- Pre-formed recipient biofilms (24-48 hour maturation)
- Donor cell suspension
- Dispersion induces: Nitric oxide donors (e.g., DETA-NO) or cis-2-decenoic acid
- Appropriate culture vessels
Procedure
- Biofilm establishment: Grow recipient biofilms for 24-48 hours under optimal conditions.
- Dispersion triggering: Add nitric oxide donor (50-100 μM) or cis-2-decenoic acid (1-10 nM) to mature biofilms.
- Donor introduction: Add donor cells during maximum dispersion (typically 2-4 hours post-induction).
- Co-incubation: Allow 4-24 hours for conjugation during dispersion window.
- Assessment: Quantify transconjugants in dispersed and residual biofilm fractions.
Expected Outcomes
- Nitric oxide treatment typically disperses 63% of biofilm biomass [49]
- Combination with antimicrobials (e.g., colistin) near-completely removes biofilms
- Conjugation frequency increases significantly during dispersion phase
Data Analysis and Interpretation
Quantitative Conjugation Metrics
Effective evaluation of contact enhancement strategies requires standardized metrics:
- Conjugation frequency: Transconjugants per recipient under defined conditions
- Spatial distribution analysis: Mapping of transconjugants within biofilm architecture
- Temporal dynamics: Rate of plasmid spread through population over time
- Persistence assessment: Stability of plasmid maintenance post-transfer
Technical Validation
- Single-cell fluorescence: Confirm actual plasmid transfer versus potential probe transfer
- Spatial correlation: Associate conjugation hotspots with specific architectural features
- Control experiments: Include essential controls for spontaneous resistance and donor contamination
Troubleshooting Guide
| Problem | Potential Causes | Solutions |
|---|---|---|
| Low conjugation frequency in mature biofilms | Physical barriers limit donor access | Introduce donors during early biofilm stages; implement matrix disruption strategies |
| Inconsistent results between replicates | Biofilm heterogeneity | Standardize inoculation density; increase replicate number; use flow-based systems for uniformity |
| Poor donor penetration | Dense EPS matrix | Incorporate matrix-degrading enzymes; use motility-enhanced donor strains |
| High donor-to-recipient ratio required | Inefficient contact formation | Switch to cis-acting plasmid systems; optimize surface contact conditions |
Enhancing donor-recipient contact in 3D biofilm structures requires integrated approaches that address both the physical barriers imposed by biofilm architecture and the biological efficiency of conjugation systems. Strategic manipulation of biofilm developmental stage, targeted disruption of matrix components, and implementation of engineered conjugative plasmids collectively enable significantly improved plasmid dissemination. The protocols detailed herein provide validated methodologies for achieving high-efficiency conjugation in biofilm environments, with particular relevance for deploying CRISPR-Cas systems as precision antimicrobials. As biofilm-associated infections continue to challenge conventional antibiotics, these strategies offer promising avenues for developing next-generation interventions that exploit rather than combat bacterial community behaviors.
Addressing Off-Target Effects and Ensuring Strain Specificity
The precision of conjugative plasmid delivery of CRISPR systems is paramount for effective biofilm control research. A primary challenge is the risk of off-target effects, where the CRISPR machinery acts on unintended genomic locations, and a lack of strain specificity, which can lead to the elimination of non-targeted bacteria. This document details protocols and application notes to quantify, minimize, and control for these factors, ensuring the integrity and safety of your research outcomes.
Understanding and Quantifying Off-Target Effects
Off-target effects occur when the Cas nuclease cleaves DNA at sites other than the intended target due to partial complementarity between the guide RNA (gRNA) and non-target genomic sequences [50] [51].
Mechanisms and Contributing Factors
The following table summarizes the key factors influencing off-target activity.
Table 1: Key Factors Influencing CRISPR/Cas9 Off-Target Effects
| Factor | Mechanism of Influence | Impact on Specificity |
|---|---|---|
| Mismatch Tolerance | Cas9 can bind and cleave DNA with up to 3-5 base pair mismatches in the gRNA, especially in the PAM-distal region [51]. | High number of tolerated mismatches directly increases the pool of potential off-target sites. |
| PAM Flexibility | While the canonical PAM for SpCas9 is NGG, it can tolerate suboptimal PAMs like NAG or NGA, expanding the range of possible target sites [51]. | Relaxed PAM requirements significantly increase the number of genomic sequences susceptible to off-target binding. |
| gRNA Secondary Structure | The formation of hairpins or other structures in the gRNA itself can affect its binding affinity and specificity [50]. | Alters the effective concentration and kinetics of correct target recognition. |
| Enzyme Concentration & Exposure Time | High concentrations of Cas9-sgRNA complex and prolonged cellular exposure can exacerbate off-target cleavage [51]. | Increases the likelihood of cleavage at sites with lower binding affinity. |
| Genetic Variations (e.g., SNPs) | Pre-existing single nucleotide polymorphisms (SNPs) in the target genome can create or destroy PAM sites or introduce mismatches that mislead gRNA design [51]. | Can render an on-target site inaccessible or create novel, unpredictable off-target sites. |
Quantitative Models for Predicting Off-Target Dynamics
Advanced models are being developed to quantitatively predict the dynamics of off-target binding. One such model treats R-loop formation (the key step in target recognition) as a random walk process in a one-dimensional free energy landscape [52]. In this model:
- Each base-pair step in R-loop formation is assigned a free energy value.
- Mismatches act as energy barriers, with their impact quantified by a free energy penalty (ΔG_MM).
- The model can predict the non-trivial dependence of R-loop formation on the proximity between multiple mismatches and reveals that the initiation "seed" region is influenced by DNA supercoiling [52].
This mechanistic, quantitative approach provides a more robust foundation for off-target prediction compared to purely heuristic scoring methods.
Experimental Protocols for Off-Target Assessment
A comprehensive off-target assessment is critical. The following workflow and protocols outline a multi-faceted approach.
Diagram 1: Comprehensive workflow for off-target assessment, integrating in silico prediction with experimental validation.
Protocol 1: Gel-Based Mutation Detection (T7E1/CEL-I Assay)
This is a rapid, economical method for initial screening of nuclease activity, including potential off-target cleavage [53].
- Sample Preparation: Harvest cells 48-72 hours post-transfection with the CRISPR-conjugative plasmid. Extract genomic DNA using a standard kit.
- PCR Amplification: Design primers flanking the on-target site and predicted off-target sites. Perform PCR to generate amplicons of 400-800 bp.
- DNA Hybridization: Purify the PCR products. For each site, hybridize the amplicons using a thermal cycler program: 95°C for 5 minutes, ramp down to 85°C at -2°C/seconds, then to 25°C at -0.1°C/seconds.
- Nuclease Digestion: Digest the heteroduplex DNA with T7 Endonuclease I (T7E1) or CEL-I nuclease for 30-60 minutes at 37°C. These enzymes cleave at mismatches in heteroduplex DNA.
- Analysis: Run the digested products on an agarose gel (2-3%). Cleavage products will appear as smaller bands. Editing frequency can be estimated by comparing band intensities.
Table 2: Comparison of Methods for Detecting Genome Editing Outcomes
| Method | Principle | Sensitivity | Detects Mutation Spectrum? | Key Limitation |
|---|---|---|---|---|
| T7E1 / CEL-I Assay [53] | Cleavage of heteroduplex DNA by mismatch-specific nucleases. | ~1-2% | No | Indirect measure; cannot reveal sequence of indels. |
| Sanger Sequencing [53] | Direct sequencing of PCR amplicons. | ~0.01% | Yes | Low-throughput; requires decomposition tools for analysis. |
| Next-Generation Sequencing (NGS) [53] | High-throughput sequencing of amplicons. | ~0.01% | Yes | Can miss large deletions that span PCR primer sites. |
| CIRCLE-Seq [54] | In vitro circularization and sequencing of Cas9-cleaved genomic DNA. | Very High (for in vitro cleavage) | Yes | Performed in vitro, may not reflect cellular context. |
Protocol 2: Next-Generation Sequencing (NGS) for Off-Target Analysis
For a comprehensive and quantitative analysis, NGS is the gold standard [53].
- Library Preparation:
- For targeted sites: Perform PCR using barcoded primers for the on-target and top predicted off-target loci. Pool the amplicons.
- For genome-wide unbiased discovery: Use methods like CIRCLE-Seq [54]. This involves fragmenting genomic DNA, circularizing the fragments, and performing in vitro Cas9 cleavage. Cleaved circles are linearized, amplified, and sequenced, providing a sensitive map of potential off-target sites.
- Sequencing: Run the pooled library on an NGS platform (e.g., Illumina MiSeq) to achieve high coverage (>1000x).
- Bioinformatic Analysis:
- Demultiplex and quality-filter the raw sequencing data.
- Align reads to the reference genome.
- Use specialized software (e.g., CRISPResso2, Cas-Analyzer) to quantify the frequency of insertions and deletions (indels) at each target site.
Strategies to Enhance Strain Specificity in Conjugative Delivery
Ensuring that the CRISPR system is delivered only to the target strain and acts specifically within it is a two-part challenge.
Enhancing CRISPR System Specificity
Table 3: Strategies to Minimize Off-Target Effects
| Strategy | Mechanism | Protocol Notes |
|---|---|---|
| High-Fidelity Cas Variants [54] [51] | Engineered Cas9 proteins (e.g., HiFi Cas9) with reduced affinity for DNA, lowering tolerance for gRNA mismatches. | Clone HiFi Cas9 into your conjugative plasmid backbone. Test on-target efficiency as it may be slightly reduced. |
| Truncated gRNAs (tru-gRNAs) [51] | Using gRNAs shorter than 20 nt reduces off-target binding energy without significantly affecting on-target activity. | Synthesize gRNAs with 17-18 nt spacers. Requires empirical testing for each target. |
| Optimized gRNA Design | Avoiding gRNAs with high sequence similarity to non-target genomic regions, especially in the seed region. | Use design tools that incorporate off-target prediction scores and exclude gRNAs with potential off-targets in essential genes. |
| Rationally Designed Cas9 Effectors | Using Cas9 versions with more stringent PAM requirements (e.g., SpCas9-NG) can reduce the genome-wide pool of potential off-target sites. | Choose a Cas nuclease whose PAM requirement best matches your specific target site to minimize genomic redundancy. |
| Avoid NHEJ Inhibitors | Inhibiting NHEJ repair (e.g., with DNA-PKcs inhibitors) to promote HDR can inadvertently increase large structural variations and chromosomal translocations [54]. | For antimicrobial applications aiming at bacterial killing, this is less relevant, but critical for therapeutic editing in eukaryotes. |
Ensuring Delivery Specificity with Conjugative Plasmids
The conjugative plasmid itself can be engineered to confer strain specificity.
- Leverage Narrow Host Range Plasmids: Use plasmids with a narrow host range limited to your target species. For example, pheromone-responsive plasmids (PRPs) in Enterococcus faecalis achieve highly efficient and species-specific conjugation [25].
- Incorporate Bacteriocins: Engineer the conjugative plasmid to encode bacteriocins (bacterial toxins) for which the plasmid also carries the immunity gene. Donor cells are immune, but recipient cells that fail to acquire the plasmid are killed, providing a strong selective pressure that enriches for the target population [25]. The pPD1 plasmid used in E. faecalis naturally employs this strategy [25].
- "Cis-Acting" Conjugative Systems: Construct a "cis-acting" plasmid where the genes for the conjugation machinery and the CRISPR antimicrobial are on the same plasmid. This ensures that any transconjugant (recipient cell that receives the plasmid) immediately becomes a new donor, leading to exponential spread within the target population and highly efficient depletion [14]. This system has been shown to be far more effective than "trans" systems where conjugation machinery and cargo are separate [14].
Application Note: Targeting an Antibiotic Resistance Gene in a Biofilm
Objective: Selectively remove the ermB gene (conferring erythromycin resistance) from a biofilm of Enterococcus faecalis using a conjugatively delivered CRISPR-Cas9 system.
Diagram 2: Sequence-specific killing of a target bacterium within a biofilm via conjugative CRISPR delivery.
Protocol: In Vitro Biofilm Killing Assay
Plasmid Construction:
- Engineer a conjugative plasmid (e.g., based on pPD1 for E. faecalis [25] or an IncP RK2 plasmid for broader host range [14]).
- Include a constitutively expressed cas9 gene and a gRNA expression cassette targeting the ermB gene.
- Ensure the plasmid retains all necessary conjugation genes (cis-acting system).
Biofilm Setup and Co-culture:
- Grow the recipient E. faecalis (with ermB) in a flow cell or microtiter plate to form a mature biofilm (24-48 hours).
- Introduce the E. coli donor strain harboring the CRISPR-conjugative plasmid to the biofilm system. Use a high donor-to-recipient ratio (e.g., 10:1) in low-salt LB media to enhance cell-to-cell contact and conjugation efficiency [14].
- Incubate under static or mild agitation conditions for 24-72 hours.
Assessment and Validation:
- Viable Counts: Disaggregate the biofilm and plate on selective media. A significant reduction in erythromycin-resistant CFUs compared to a control (e.g., non-targeting gRNA) indicates successful killing.
- PCR and Sequencing: Verify the loss or disruption of the ermB gene in the remaining population.
- Off-Target Check: Use the NGS protocol (Protocol 2) on the treated population to screen for mutations at the top predicted off-target sites in the E. faecalis genome.
The Scientist's Toolkit: Essential Reagents and Materials
Table 4: Key Research Reagent Solutions for Conjugative CRISPR Delivery
| Reagent / Material | Function | Example & Notes |
|---|---|---|
| High-Fidelity Cas9 | Reduces off-target cleavage while maintaining on-target activity. | HiFi Cas9 [54] is an engineered SpCas9 variant with point mutations (e.g., R691A) that reduce off-target effects. |
| Narrow Host Range Conjugative Plasmid | Enables species-specific delivery of the CRISPR payload. | Pheromone-Responsive Plasmids (PRPs) like pPD1 for Enterococcus faecalis [25]. |
| Broad Host Range Conjugative Plasmid | Allows delivery across a wider range of bacterial species. | IncP RK2-based plasmids [14]. Useful for targeting multiple Gram-negative pathogens. |
| T7 Endonuclease I (T7E1) | Enzyme for detecting mismatches in heteroduplex DNA in gel-based assays. | Available from suppliers like NEB. Part of Protocol 1 for initial, cost-effective off-target screening [53]. |
| CIRCLE-Seq Kit | For genome-wide, unbiased identification of off-target cleavage sites. | A highly sensitive in vitro method. Kits are available commercially to implement the unbiased screening in Protocol 2 [54]. |
| NGS Amplicon-Seq Kit | For preparing sequencing libraries from PCR amplicons of target sites. | Kits from Illumina or NEB are used in Protocol 2 for deep, quantitative analysis of editing efficiency at specific loci [53]. |
The escalating crisis of antimicrobial resistance (AMR), particularly the resilience of biofilm-associated infections, necessitates the development of next-generation antibacterial strategies [13] [1]. Biofilms, structured communities of bacteria encased in an extracellular polymeric substance (EPS), can exhibit up to 1,000-fold greater tolerance to antibiotics compared to their planktonic counterparts [13]. CRISPR/Cas9 gene-editing technology has emerged as a precision tool to combat this threat by enabling the sequence-specific disruption of antibiotic resistance genes, quorum-sensing pathways, and biofilm-regulating factors [13] [25]. However, the clinical translation of CRISPR-based antimicrobials is hampered by significant delivery challenges, including poor stability, inefficient bacterial uptake, and limited penetration through the protective biofilm matrix [13].
This application note details a synergistic hybrid approach that combines the strengths of two advanced delivery systems: the high efficiency and broad-host-range of conjugative plasmids with the enhanced biofilm-penetration and protective capabilities of nanoparticle carriers. Conjugative plasmids, such as pheromone-responsive plasmids (PRPs) in Enterococcus faecalis or the IncP RK2 system in Escherichia coli, facilitate robust inter-species transfer of CRISPR machinery [25] [14]. When integrated with engineered nanoparticles, these systems can overcome the physical and biological barriers that have traditionally limited antimicrobial efficacy, offering a powerful and precise method for controlling biofilm-forming, multidrug-resistant pathogens [13].
Key Performance Data of Individual and Hybrid Systems
The tables below summarize quantitative data from key studies, demonstrating the efficacy of conjugative plasmid and nanoparticle systems individually, and projecting the potential performance of their hybrid combination.
Table 1: Performance of Conjugative Plasmid Systems for CRISPR Delivery
| Conjugative System | Donor-Recipient Pair | Key Experimental Condition | Transfer Efficiency / Killing Outcome | Citation |
|---|---|---|---|---|
| pPD1 (PRP) derivative | E. faecalis CK135 → E. faecalis OG1SSp (pAM771/ermB) | In vitro mixing on agar plate | Significant reduction of erythromycin-resistant transconjugants [25] | |
| pPD1 (PRP) derivative | E. faecalis CK135 → E. faecalis V583 (MDR) | In vitro mixing on agar plate | Resolution of erythromycin resistance from clinical MDR isolate [25] | |
| pNuc-cis (IncP RK2) | E. coli → Salmonella enterica | Filter mating assay (24h) | Conjugation frequency of 1x10⁻² [14] | |
| pNuc-cis (IncP RK2) | E. coli → Salmonella enterica | Liquid culture with glass beads (enhanced contact) | Conjugation frequency of up to ~100% [14] | |
| pNuc-cis (IncP RK2) | E. coli → Salmonella enterica | Delivery of sgRNA targeting non-essential genes | High killing efficiencies demonstrated [14] |
Table 2: Performance of Nanoparticle Systems for CRISPR Delivery Against Biofilms
| Nanoparticle Type | Target / Application | Key Cargo | Efficacy / Outcome | Citation |
|---|---|---|---|---|
| Liposomal Nanoparticles | Pseudomonas aeruginosa biofilm | CRISPR-Cas9 | > 90% reduction in biofilm biomass in vitro [13] | |
| Gold Nanoparticles (AuNPs) | General CRISPR delivery enhancement | CRISPR-Cas9 | 3.5-fold increase in gene-editing efficiency vs. non-carrier systems [13] | |
| Ionizable Lipid Nanoparticles (LNPs) | Advanced CRISPR delivery in vivo | Nucleic Acids/CRISPR | Leading performance in clinical and pre-clinical delivery [55] |
Experimental Protocols
Protocol 1: Engineering a cis-Conjugative Plasmid for CRISPR Delivery
This protocol outlines the construction of a high-efficiency, self-transmissible plasmid for targeted bacterial killing [14].
Primary Objective: To create a conjugative plasmid that encodes all necessary components for its own transfer and for CRISPR-Cas9-mediated killing of a specific bacterial target.
Materials and Reagents:
- Plasmid Backbone: pTA-Mob or similar IncP RK2-based backbone containing the origin of transfer (oriT) and full conjugation machinery genes [14].
- Cas9 Nuclease Gene: A gene encoding Cas9, preferably a high-fidelity or dual nuclease variant (e.g., TevSpCas9), under the control of an inducible promoter (e.g., pBAD arabinose-inducible promoter) [14].
- sgRNA Cassette: A DNA sequence for the single-guide RNA (sgRNA), driven by a constitutive promoter (e.g., pTet) [14].
- Selectable Marker: An antibiotic resistance gene (e.g., chloramphenicol acetyltransferase, cat) for selection in the donor and transconjugant strains [25] [14].
- Molecular Biology Reagents: Restriction enzymes, DNA ligase, Gibson assembly master mix, competent E. coli (e.g., DH5α) for cloning.
Methodology:
- Clone the CRISPR-Cas9 Module: Insert the Cas9 nuclease gene and the sgRNA expression cassette into the multiple cloning site (MCS) of the pTA-Mob plasmid backbone using standard restriction-ligation or Gibson assembly techniques.
- Verify Plasmid Construction: Transform the assembled plasmid into a cloning-grade E. coli strain. Select positive clones on agar plates containing the appropriate antibiotic (e.g., chloramphenicol). Confirm the plasmid sequence via colony PCR and Sanger sequencing.
- Transform Donor Strain: Isolate the verified plasmid and transform it into the conjugative donor strain (e.g., E. coli for RK2 systems).
Critical Steps and Optimization:
Protocol 2: Conjugation and Killing Assay in Biofilm-Mimicking Conditions
This protocol measures the transfer efficiency and antibacterial efficacy of the engineered plasmid under conditions that promote cell-to-cell contact, mimicking a biofilm environment [14].
Primary Objective: To assess the conjugation frequency and target cell killing efficiency of the cis-conjugative CRISPR plasmid in a co-culture system designed to enhance cell-to-cell contact.
Materials and Reagents:
- Bacterial Strains: Donor strain (e.g., E. coli harboring the pNuc-cis plasmid) and recipient strain (e.g., Salmonella enterica or a biofilm-forming ESKAPE pathogen).
- Culture Media: Low-salt LB broth (LSLB, 0.25% w/v NaCl) and appropriate solid agar plates for selection and CFU enumeration [14].
- Contact Enhancement Substrate: 0.5 mm glass beads or similar solid surfaces to promote biofilm-like cell aggregation [14].
- Inducing Agent: L-Arabinose (e.g., 0.2% w/v) for induction of Cas9 expression from the pBAD promoter [14].
- Antibiotics: For selective plating of donors, recipients, and transconjugants.
Methodology:
- Co-culture Setup: Grow donor and recipient cultures to mid-log phase. Mix at a donor-to-recipient ratio of 10:1 in LSLB broth supplemented with 0.5 mm glass beads [14].
- Conjugation Incubation: Incubate the co-culture for up to 72 hours at 37°C with mild agitation (60 RPM) or static conditions [14].
- Induction of Killing: Add arabinose to the culture to induce Cas9 expression and initiate targeted killing in transconjugants.
- Enumeration and Analysis: After incubation, serially dilute the culture and plate on selective media to quantify the number of donors, recipients, and transconjugants. Calculate conjugation frequency as (number of transconjugants) / (total number of recipients).
Critical Steps and Optimization:
- Agitation Control: Mild agitation preserves cell-to-cell contacts formed around the beads, which is critical for high conjugation efficiency [14].
- Timing of Induction: The timing of arabinose induction can be varied to study its impact on the propagation of the plasmid and the efficiency of killing.
Protocol 3: Formulation of CRISPR-Nanoparticle Hybrid Complexes
This protocol describes the encapsulation of conjugative plasmids into lipid nanoparticles (LNPs), a leading nanocarrier platform, for enhanced biofilm penetration and delivery [13] [55].
Primary Objective: To formulate and characterize hybrid nanoparticles loaded with the engineered conjugative plasmid to create a multi-mechanistic anti-biofilm therapeutic.
Materials and Reagents:
- Lipid Components: Ionizable lipid, phospholipid, cholesterol, and PEG-lipid [55].
- Aqueous Buffer: Acetate buffer (pH 4.0-5.0).
- Genetic Cargo: Purified cis-conjugative CRISPR plasmid DNA.
- Formulation Equipment: Microfluidic mixer or tangential flow filtration system.
Methodology:
- Prepare Lipid Solution: Dissolve the lipid mixture (ionizable lipid, phospholipid, cholesterol, PEG-lipid) in ethanol.
- Prepare Aqueous Phase: Dilute the plasmid DNA in acidified acetate buffer.
- Nanoparticle Formation: Rapidly mix the lipid and aqueous phases using a microfluidic device. The change in pH upon mixing promotes the spontaneous formation of LNPs encapsulating the plasmid DNA.
- Buffer Exchange and Purification: Dialyze or use tangential flow filtration against PBS (pH 7.4) to remove ethanol and neutralize the LNPs.
- Characterization: Measure particle size and zeta potential using dynamic light scattering (DLS), and determine encapsulation efficiency using a dye exclusion assay.
Critical Steps and Optimization:
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents and Materials
| Reagent / Material | Function / Application | Example / Note |
|---|---|---|
| Pheromone-Responsive Plasmid (PRP) Backbone | Narrow-host-range conjugative delivery for Enterococcus faecalis; allows targeted decolonization of MDR strains [25]. | pPD1 derivative; encodes beneficial bacteriocins for competitive fitness [25]. |
| Broad-Host-Range Plasmid Backbone | Conjugative delivery across Gram-negative species; ideal for targeting ESKAPE pathogens like P. aeruginosa and K. pneumoniae [14]. | IncP RK2-based plasmid (e.g., pTA-Mob) for cis-conjugative systems [14]. |
| Ionizable Lipid Nanoparticles (LNPs) | Protect CRISPR cargo from degradation; enhance biofilm penetration and bacterial uptake; promote endosomal escape [13] [55]. | Leading nanocarrier platform; composition can be tuned for specific targeting and release kinetics [55]. |
| Gold Nanoparticles (AuNPs) | Inorganic carrier for CRISPR components; easily functionalized; enhances editing efficiency [13]. | Can be conjugated with plasmids or Cas9/sgRNA ribonucleoproteins (RNPs); shown to boost efficiency 3.5-fold [13]. |
| TevSpCas9 Nuclease | High-specificity dual nuclease; reduces the likelihood of off-target effects during targeted bacterial killing [14]. | I-TevI domain fused to SpCas9; requires specific protospacer adjacent motif (PAM) sites [14]. |
Integrated Hybrid Workflow and Data Projection
The ultimate application of this technology involves the sequential or simultaneous use of both delivery modalities. The nanoparticle, loaded with the conjugative plasmid, is applied to a mature biofilm. It first acts as a Trojan horse, penetrating the EPS and delivering its plasmid cargo to a subset of target cells. Once inside these initial "founder" bacteria, the conjugative machinery is expressed, initiating a second, self-amplifying wave of delivery via conjugation to neighboring cells within the biofilm, ultimately leading to widespread and precise genetic targeting.
Table 4: Projected Advantages of the Hybrid System over Individual Modalities
| Characteristic | Conjugative Plasmid Alone | Nanoparticle Alone | Hybrid System (Projected) |
|---|---|---|---|
| Biofilm Penetration | Limited by diffusion and cell contact within dense EPS [14]. | Enhanced passive diffusion; can be engineered for active penetration [13]. | Superior: NPs breach EPS, enabling initial plasmid delivery to founder cells. |
| Delivery Efficiency | Very high once initial contact is made; self-amplifying [14]. | High for initial transfection, but not self-propagating [13]. | Maximized: Combines efficient initial NP delivery with self-amplifying conjugation. |
| Breadth of Targeting | Host-range limited by plasmid specificity [25] [14]. | Can be engineered for broad targeting but may lack specificity [55]. | Potentially Broad & Specific: NP targets biofilm physically, plasmid targets species genetically. |
| Resistance Evolution | Potential for mutations in conjugation machinery or target site [25]. | Potential for changes in surface receptors or efflux pumps [13]. | Mitigated: Dual-mechanism delivery reduces the probability of concurrent resistance. |
Conjugative plasmid delivery of CRISPR-based systems represents a promising strategy for the precise manipulation of microbial communities and the targeting of antibiotic-resistant pathogens within biofilms. However, the efficacy of this approach is contingent upon overcoming the native defense systems of target bacteria, primarily Restriction-Modification (R-M) and CRISPR-Cas immunity. This Application Note details practical strategies and protocols for bypassing these barriers, enabling efficient plasmid transfer and CRISPR-mediated killing or genome editing. The content is framed within a broader research thesis on conjugative delivery for biofilm control, providing actionable methodologies for researchers and drug development professionals.
Strategic Approaches to Bypass Host Defenses
Bacterial targets possess multiple defense mechanisms that can cleave or reject incoming conjugative plasmids. The table below summarizes the primary barriers and the corresponding strategies to neutralize them.
Table 1: Key Bacterial Defense Mechanisms and Bypass Strategies
| Defense Mechanism | Challenge Posed | Proposed Bypass Strategy | Key Experimental Support |
|---|---|---|---|
| Restriction-Modification (R-M) Systems | Cleavage of unmethylated foreign DNA at specific recognition sites [56]. | Co-delivery of protective methyltransferases on the conjugative plasmid [56]. | LlaDCHI R-M system provided phage resistance in S. thermophilus; methylase activity protects DNA [56]. |
| CRISPR-Cas Immunity in Target | Cleavage of plasmid DNA if it contains a sequence matching the host's spacer [56]. | Leverage natural PAM requirements or use CRISPRi (dCas9) to avoid lethal double-strand breaks [9]. | CRISPR-Cas and R-M systems are compatible and can function independently [56]. |
| Inefficient Conjugation | Low transfer frequency of CRISPR machinery limits killing/editing efficiency [14]. | Use cis-acting conjugative plasmids that encode their own transfer machinery [14]. | Cis-plasmids increased conjugation frequency by ~1000-fold and achieved near 100% transfer in biofilms [14]. |
| Physical Biofilm Barrier | Extracellular Polymeric Substance (EPS) matrix limits diffusion and access to cells [6]. | Combine CRISPR with nanoparticle carriers to enhance penetration and delivery [6]. | Liposomal Cas9 reduced P. aeruginosa biofilm by >90%; gold NPs increased editing efficiency 3.5-fold [6]. |
Quantitative Data on Defense Bypass Efficacy
The following tables consolidate key quantitative findings from foundational studies, providing a reference for expected outcomes when implementing these strategies.
Table 2: Efficacy of Combined Defense Systems Against Phage Infection
| Bacterial Strain Defense Profile | Challenge Phage | Efficiency of Plating (EOP) | Observation |
|---|---|---|---|
| Wild-type S. thermophilus DGCC7710 (No active defense) | 2972 | 1 (Baseline) | Phage infection proceeds normally [56]. |
| With active CRISPR-Cas (One targeting spacer) | 2972 | 10⁻⁶ | High-level resistance from sequence-specific cleavage [56]. |
| With LlaDCHI R-M system (35 GATC sites) | 2972 | 10⁻⁵ | High-level resistance from multi-site restriction [56]. |
| With combined CRISPR-Cas & R-M systems | 2972 | <10⁻⁹ | Near-total resistance; systems are compatible and additive [56]. |
Table 3: Impact of Conjugative Plasmid Design on Transfer Efficiency
| Conjugative Plasmid Configuration | Donor-Recipient Pair | Experimental Condition | Conjugation Frequency |
|---|---|---|---|
| pNuc-trans (Transfer machinery in trans) | E. coli to S. enterica | Filter mating, 24 hours | ~1 × 10⁻⁵ [14] |
| pNuc-cis (Transfer machinery in cis) | E. coli to S. enterica | Filter mating, 24 hours | ~1 × 10⁻² [14] |
| pNuc-cis (Transfer machinery in cis) | E. coli to S. enterica | Liquid culture with glass beads (biofilm-like) | Up to ~100% [14] |
Detailed Experimental Protocols
Protocol: Bypassing R-M Systems via Plasmid-Borne Methyltransferases
This protocol describes the introduction of a heterologous R-M system into a conjugative plasmid to pre-protect it from restriction in the target bacterium.
Key Research Reagent Solutions Table 4: Essential Reagents for R-M Bypass
| Reagent / Material | Function / Explanation | Example (from Literature) |
|---|---|---|
| Broad-Host-Range Cloning Vector | Base plasmid with origin of transfer (oriT) and compatible replication origin. | pTA-Mob (IncP RK2) [14]. |
| Methyltransferase Genes | Provides in vivo methylation of the plasmid DNA to protect from restriction endonucleases. | LlaDCHI-A and LlaDCHI-B genes from Lactococcus lactis [56]. |
| Constitutive Promoter | Drives constant expression of methyltransferase genes in the host. | Native promoter of the methyltransferase operon [56]. |
| CRISPR-Cas9 Payload | The functional machinery for gene editing or killing in the target. | TevSpCas9 nuclease and sgRNA expression cassette [14]. |
Procedure:
- Cloning: Clone the genes for the restriction endonuclease and the corresponding methyltransferase (e.g., the LlaDCHI operon) into your broad-host-range conjugative plasmid backbone. Ensure the methyltransferase genes are under a strong, constitutive promoter active in the donor strain.
- Transformation: Transform the constructed plasmid into the donor E. coli strain.
- Validation of Methylation: Isolate the plasmid DNA from the donor E. coli and subject it to in vitro restriction digest using the cognate restriction enzyme (e.g., DpnII for LlaDCHI). Effective methylation will confer complete resistance to digestion.
- Conjugation: Proceed with the standard conjugation protocol between the donor and the target bacterial recipient that possesses the R-M system.
- Evaluation: Compare the conjugation efficiency of the methyltransferase-encoding plasmid against a control plasmid lacking the methyltransferase.
Protocol: High-Efficiency Delivery Using Cis-Acting Conjugative Plasmids
This protocol outlines the use of a self-transmissible plasmid to maximize delivery rates, which is critical for effective CRISPR-mediated killing before defenses are activated.
Procedure:
- Plasmid Construction: Assemble the final plasmid (e.g., pNuc-cis) where a single vector contains:
- The origin of transfer (oriT).
- The entire conjugation machinery (e.g., tra genes from IncP RK2 plasmid).
- The CRISPR nuclease (e.g., Cas9 or TevSpCas9) under a tightly regulated, inducible promoter (e.g., pBAD).
- The guide RNA (sgRNA) under a constitutive promoter.
- A selectable marker for transconjugants.
- Donor Preparation: Transform the cis-acting plasmid into the donor E. coli strain.
- Conjugation Setup:
- Method A (Liquid Culture with Beads): Mix donor and recipient cells at a 10:1 ratio in a low-salt LB medium (0.25% NaCl) containing 0.5 mm glass beads to enhance cell-to-cell contact. Incubate with mild agitation (60 RPM) for up to 72 hours [14].
- Method B (Filter Mating): Concentrate donor and recipient cells on a sterile filter membrane placed on a non-selective agar plate. Incubate for 4-24 hours.
- Selection and Analysis: Resuspend the conjugation mixture and plate on selective media to isolate transconjugants. Calculate the conjugation frequency as the number of transconjugants per total recipient cells.
Visualization of Strategies and Workflows
The following diagrams illustrate the core concepts and experimental workflows for bypassing bacterial defense systems.
Diagram 1: Strategic overview for bypassing bacterial defense systems during conjugative delivery.
Diagram 2: Experimental workflow for bypassing Restriction-Modification systems.
Diagram 3: Workflow for high-efficiency delivery using a cis-acting conjugative plasmid under biofilm-promoting conditions.
Assessing Efficacy, Specificity, and Clinical Potential
In Vitro and In Vivo Models for Quantifying Biofilm Disruption and Bacterial Killing
Bacterial biofilms present a significant challenge in treating infections due to their inherent resistance to conventional antibiotics. The development of innovative strategies, such as conjugative plasmid delivery of CRISPR-based antimicrobials, requires robust and quantifiable models to assess their efficacy both in laboratory settings and living organisms. This document provides detailed application notes and protocols for quantifying biofilm disruption and bacterial killing, specifically framed within research utilizing conjugative CRISPR systems. The methodologies are designed to provide researchers with reliable, reproducible techniques to evaluate novel anti-biofilm therapies from in vitro validation to in vivo confirmation.
Quantitative Data on Biofilm Disruption Modalities
The efficacy of anti-biofilm treatments varies significantly across bacterial species and delivery methods. The table below summarizes quantitative findings from key studies on non-antibiotic treatments and biological agents.
Table 1: Quantitative Efficacy of Various Biofilm Disruption Modalities
| Treatment / Delivery System | Target Bacteria | In Vitro Efficacy (CFU Reduction/Biomass Disruption) | In Vivo Model & Efficacy |
|---|---|---|---|
| N-acetylcysteine (3.3%) [57] | E. coli | Decreased biofilm biomass; killed bacteria within biofilms [57] | Not specified in the cited study [57] |
| P. aeruginosa, K. pneumoniae | Decreased recoverable CFU; no change in biofilm biomass [57] | Not specified in the cited study [57] | |
| Hydrogen Peroxide (1%) [57] | E. coli | Decreased biofilm biomass and reduced CFU [57] | Not specified in the cited study [57] |
| K. pneumoniae | Reduction in CFU [57] | Not specified in the cited study [57] | |
| P. aeruginosa | Minimal effects observed [57] | Not specified in the cited study [57] | |
| Blue Laser Light [58] | P. aeruginosa | Protocol A/B/C-high (120 J/cm²): Near-complete growth inhibition of planktonic cultures; significant biofilm biomass reduction [58] | Mouse skin wound infection model: Effective clearance of bacterial cells with minimal toxicity to mammalian tissues [58] |
| Conjugative CRISPR (pKH88[sp-ermB]) [25] | E. faecalis (with ermB gene) | Significant reduction of erythromycin-resistant transconjugants in vitro [25] | Murine intestine model: Reduced occurrence of antibiotic-resistant E. faecalis by several orders of magnitude [25] |
| Cis-Conjugative Plasmid (pNuc-cis) [14] | S. enterica | Conjugation frequency up to 100% under optimized conditions; high killing efficiency with targeted sgRNAs [14] | Not specified in the cited study [14] |
Experimental Protocols
Protocol 1: In Vitro Assessment of Non-Antibiotic Treatments on Pre-Formed Biofilms
This protocol is adapted from the study evaluating N-acetylcysteine, EDTA, and hydrogen peroxide on Gram-negative pathogens [57].
Key Materials:
- Bacterial Strains: Clinical isolates of E. coli, P. aeruginosa, and K. pneumoniae.
- Growth Medium: Appropriate broth and agar (e.g., Tryptic Soy Broth, Brain Heart Infusion).
- Biofilm Growth Substrate: 96-well microtiter plates (for biomass assays) or Calgary devices (for CFU enumeration).
- Test Agents: Solutions of N-acetylcysteine (3.3%), hydrogen peroxide (1%), and EDTA formulations in sterile water or buffer.
Procedure:
- Biofilm Formation: Grow bacterial cultures to mid-log phase. Dilute and inoculate wells of a 96-well plate or Calgary device. Incubate under static conditions for 24-48 hours at 37°C to allow biofilm formation.
- Treatment: Carefully remove the planktonic culture and rinse the established biofilms with a sterile buffer (e.g., PBS) to remove loosely attached cells. Add the prepared non-antibiotic treatments to the wells. Include an untreated control (buffer only).
- Incubation: Incubate the plates with the treatments for a specified period (e.g., 1-2 hours) at 37°C.
- Quantification:
- Biofilm Biomass (Crystal Violet Assay): After treatment, rinse the wells, fix the biofilms with methanol, and stain with 0.1% crystal violet. Elute the bound dye with acetic acid and measure the absorbance at 595 nm.
- Viable Bacterial Count (CFU Enumeration): After treatment, add a sterile buffer to the wells and disrupt the biofilms by vigorous pipetting or sonication. Serially dilute the suspension and plate on agar plates. Count the colonies after 24 hours of incubation.
Protocol 2: Conjugative Delivery of CRISPR-Cas9 for Selective Bacterial Killing
This protocol outlines the procedure for using pheromone-responsive conjugative plasmids to deliver CRISPR-Cas9 and target antibiotic resistance genes in Enterococcus faecalis [25].
Key Materials:
- Bacterial Strains:
- Donor: E. faecalis CK135 harboring the engineered pPD1 plasmid (e.g., pKH88[sp-ermB]) with constitutive cas9 expression and a guide RNA (e.g., targeting ermB or tetM).
- Recipient: Antibiotic-resistant E. faecalis (e.g., strain OG1SSp with pAM771 (ermB) or V583 with pTEF1 (ermB)).
- Media: Brain Heart Infusion (BHI) broth and agar, supplemented with appropriate antibiotics (e.g., chloramphenicol for plasmid maintenance).
- Equipment: Microcentrifuge tubes, agar plates, incubator.
- Bacterial Strains:
Procedure:
- Culture Preparation: Grow donor and recipient strains separately in BHI broth to mid-log phase.
- In Vitro Mating:
- Mix donor and recipient cells at a specific ratio (e.g., 1:1) and pellet by centrifugation.
- Resuspend the cell mixture in a small volume of broth and spot onto a BHI agar plate.
- Incubate for 18-24 hours at 37°C to allow conjugation.
- Quantification of Conjugation and Killing:
- Harvest the cells from the plate and resuspend in a known volume of buffer.
- Perform serial dilutions and plate on selective media:
- Media with chloramphenicol to count total transconjugants (recipients that received the plasmid).
- Media with erythromycin (or tetracycline) to count recipients that retained the antibiotic resistance gene.
- Compare the number of resistant recipients in the group that received the cognate CRISPR-targeting plasmid versus a non-cognate control. A significant reduction indicates sequence-specific killing.
Protocol 3: In Vivo Murine Model for Assessing Conjugative CRISPR Efficacy
This protocol describes the use of a mouse model to evaluate the decolonization of antibiotic-resistant E. faecalis from the intestine [25].
Key Materials:
- Animals: Specific pathogen-free (SPF) mice.
- Bacterial Strains: Donor and recipient E. faecalis strains as described in Protocol 2.
- Reagents: Antibiotics for selective plating, anaesthetics, sterile PBS.
Procedure:
- Colonization: Pre-treat mice with an antibiotic (e.g., streptomycin) in their drinking water to disrupt the native gut microbiota and facilitate colonization by the recipient E. faecalis strain. Introduce the recipient strain orally.
- Treatment/Conjugation: Once colonization is established, orally administer the donor E. faecalis strain harboring the conjugative CRISPR plasmid.
- Monitoring and Sample Collection: At designated time points post-treatment, collect fecal pellets from the mice.
- Quantification:
- Homogenize the fecal samples in PBS.
- Perform serial dilutions and plate on selective media to enumerate:
- Total E. faecalis (e.g., on media with a strain-specific antibiotic).
- Antibiotic-resistant E. faecalis (e.g., on media with erythromycin).
- A significant reduction in the counts of antibiotic-resistant E. faecalis in the treatment group compared to a control group (e.g., receiving a donor with a non-targeting plasmid) indicates successful in vivo conjugative delivery and killing.
Visualization of Experimental Workflows
Conjugative CRISPR Delivery and Killing Mechanism
Integrated In Vitro to In Vivo Evaluation Pipeline
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents and Materials for Conjugative CRISPR-Biofilm Studies
| Item | Function/Application | Specific Examples / Notes |
|---|---|---|
| Pheromone-Responsive Plasmids (PRPs) | High-efficiency conjugative delivery vector specific to E. faecalis; can be engineered to carry CRISPR machinery. | pPD1 derivative (e.g., pKH88); encodes Bac-21 bacteriocin for competitive fitness [25]. |
| Cis-Conjugative Plasmid Systems | Plasmid encoding both conjugation machinery and CRISPR nuclease, enabling exponential spread in bacterial populations. | pNuc-cis (based on IncP RK2 plasmid); significantly higher conjugation frequency than trans systems [14]. |
| Constitutive Promoters | Drives continuous expression of Cas9 nuclease within the target cell, essential for immediate killing post-conjugation. | bacA promoter used in pKH88 constructs [25]. |
| Selective Culture Media | Allows for isolation and quantification of specific bacterial types (donors, recipients, transconjugants) post-experiment. | BHI Agar supplemented with antibiotics (e.g., Chloramphenicol, Erythromycin, Tetracycline) [25]. |
| In Vivo Colonization Model | Provides a living system to test the efficacy and safety of conjugative CRISPR delivery in a complex biological environment. | Murine model with antibiotic-induced intestinal dysbiosis [25]. |
| Biofilm Growth Devices | Provides a surface and environment for robust, reproducible biofilm formation for in vitro disruption assays. | 96-well microtiter plates, Calgary Biofilm Devices, or flow cells for confocal microscopy [57] [58]. |
| Non-Antibiotic Biofilm Disruptors | Chemical agents used as positive controls or combination therapies to disrupt biofilm matrix. | N-acetylcysteine (3.3%), Hydrogen Peroxide (1%), EDTA [57]. |
The escalating crisis of antimicrobial resistance (AMR), driven significantly by biofilm-associated infections, demands innovative therapeutic strategies. A primary focus of contemporary research involves the precise delivery of CRISPR-Cas systems to target and eliminate antibiotic resistance genes within bacterial populations. This application note provides a comparative analysis of three primary delivery modalities: conjugative plasmids, bacteriophage vectors, and nanoparticle (NP) systems. Each platform offers distinct advantages and challenges in efficiency, payload capacity, and applicability against biofilms. Conjugative plasmids excel in delivering DNA-cleaving CRISPR systems and can exploit plasmid incompatibility for targeted removal of resistance genes [59]. Phage delivery offers high bacterial specificity and potent biofilm penetration through natural lytic cycles [60] [61]. Nanoparticle systems enhance stability and enable co-delivery of CRISPR components with conventional antibiotics, synergistically disrupting biofilm integrity and resensitizing bacteria to treatment [13] [62]. The optimal choice of delivery vehicle is context-dependent, influenced by the target pathogen, the nature of the biofilm, and the specific CRISPR mechanism employed (cleaving vs. silencing). The following sections detail quantitative comparisons, experimental protocols, and logistical guidance for implementing these technologies in biofilm control research.
Biofilms are structured communities of microorganisms encased in an extracellular polymeric substance (EPS), conferring inherent resistance to antibiotics and host immune responses [1] [63]. The EPS matrix limits antibiotic penetration and creates heterogeneous microenvironments where bacteria exhibit reduced metabolic activity, further complicating treatment [64]. The ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species) are particularly notorious for forming treatment-resistant biofilms on medical devices and tissues [1] [60].
CRISPR-Cas systems present a revolutionary approach to combat biofilm-associated AMR by enabling sequence-specific targeting of resistance genes, virulence factors, or genes essential for biofilm integrity [60] [65]. The effectiveness of this strategy is wholly dependent on the delivery vehicle. The three systems analyzed herein function through fundamentally different mechanisms:
- Conjugative Plasmids are self-transmissible DNA molecules that can transfer CRISPR-Cas machinery directly between bacterial cells via conjugation. Their efficacy is influenced by plasmid copy number and compatibility with the target resistance plasmid [59].
- Bacteriophage Vectors are viruses that naturally infect bacteria. They can be engineered to deliver CRISPR-Cas genes, leveraging the phage's ability to inject genetic material into specific bacterial hosts, often resulting in cell lysis and biofilm disruption [60] [61].
- Nanoparticle Systems are synthetic carriers, such as lipid-based or metallic nanoparticles, that encapsulate or complex with pre-assembled CRISPR-Cas ribonucleoproteins (RNPs) or encoding plasmids. They protect the payload and facilitate entry into bacterial cells within the biofilm [13] [62].
Comparative Performance Data
The following tables synthesize key performance metrics and characteristics of the three delivery systems, based on current experimental data.
Table 1: Quantitative Performance Metrics of Delivery Systems
| Performance Metric | Conjugative Plasmids | Phage Delivery Systems | Nanoparticle Systems |
|---|---|---|---|
| Delivery Efficiency | Varies with conjugation efficiency; can be high within receptive populations [59] | Highly efficient for specific host strains; narrow host range can be a limitation [60] | Enhanced editing efficiency (e.g., gold NPs show ~3.5-fold increase) [13] [62] |
| Biofilm Biomass Reduction | Effective when targeting resistance genes, leading to antibiotic sensitization [59] | Potent biofilm penetration and dispersal via lytic cycle | High (e.g., liposomal Cas9 reduces P. aeruginosa biofilm by >90% in vitro) [13] [62] |
| Payload Capacity | High; can deliver full CRISPR-Cas systems and multiple gRNAs | Limited by capsid size; constrains Cas protein choice | High; can be co-loaded with antibiotics or other agents [13] |
| Host Specificity | Broad among Gram-negative bacteria; depends on pilus type | Narrow and highly specific; determined by phage receptor binding | Broad; can be functionalized for targeting [63] |
| Immunogenicity | Low to moderate | Can be high; pre-existing antibodies may neutralize efficacy | Variable; depends on NP material (e.g., lipid vs. metal) |
Table 2: Strategic Advantages and Limitations
| Characteristic | Conjugative Plasmids | Phage Delivery Systems | Nanoparticle Systems |
|---|---|---|---|
| Key Advantages | • Capacity for large genetic payloads• Self-replicating within host• Can exploit plasmid incompatibility for targeted clearance [59] | • High natural specificity and infectivity• Self-amplifying at infection site• Innate ability to disrupt biofilms [60] | • Co-delivery with antibiotics [13]• Tunable physicochemical properties• Protects CRISPR payload from degradation [63] |
| Major Challenges | • Lower efficiency in dense biofilms• Horizontal gene transfer risks | • Limited host range• Rapid emergence of phage-resistant mutants• Potential for lysogeny in temperate phages [61] | • Potential cytotoxicity [13] [63]• Complex manufacturing and characterization• Inefficient uptake in some bacterial species |
| Optimal Use Case | Targeting plasmid-borne resistance in low-copy number plasmids using incompatible delivery vectors [59] | Targeted eradication of specific pathogens in mono-species biofilms; phage-antibiotic synergy (PAS) [64] | Complex, multi-species biofilm environments; synergistic therapy with antibiotics [13] [62] |
Detailed Experimental Protocols
Protocol 1: Sensitizing Bacteria Using Conjugative CRISPR Plasmids
This protocol outlines the use of conjugative plasmids to deliver a DNA-cleaving CRISPR-Cas system to target an antibiotic resistance plasmid within a bacterial population, subsequently sensitizing it to conventional antibiotics [59].
Research Reagent Solutions:
- Donor Strain: E. coli harboring the conjugative pCRISPR plasmid (e.g., pCleaving with Cas9 and gRNA targeting a specific resistance gene, e.g.,
blaNDM-1). - Recipient Strain: Target pathogen (e.g., K. pneumoniae) carrying the wild-type pAMR plasmid (e.g., pNDM-1).
- Growth Media: LB broth and LB agar plates.
- Selection Antibiotics: Use antibiotics to select for donor (e.g., Kanamycin), recipient (e.g., Ampicillin, due to pAMR), and transconjugants (e.g., Kanamycin + Ampicillin).
- Conjugation Buffer: Phosphate-buffered saline (PBS), pH 7.4.
Procedure:
- Culture Preparation: Grow donor and recipient strains overnight in LB broth with appropriate antibiotics to mid-exponential phase (OD600 ~0.5).
- Cell Harvesting: Harvest cells by centrifugation at 5,000 × g for 10 minutes and wash twice with PBS to remove residual antibiotics.
- Conjugation Mixture: Mix donor and recipient cells at a 1:1 ratio (e.g., 10^8 cells each) in a final volume of 1 mL PBS. A donor-only control is essential.
- Conjugation: Spot the entire mixture onto a pre-warmed, sterile 0.22 µm nitrocellulose filter placed on an LB agar plate without antibiotics. Incubate at 37°C for 4-6 hours to allow conjugation.
- Transconjugant Selection: Resuspend the cells from the filter in 1 mL PBS. Plate serial dilutions onto LB agar plates containing antibiotics that select for transconjugants (e.g., Kanamycin + Ampicillin). Incubate plates at 37°C for 24-48 hours.
- Efficiency Assessment: Count the colony-forming units (CFUs) of transconjugants. Calculate the conjugation frequency as (number of transconjugants) / (number of recipient cells).
- Antibiotic Sensitization Assay: Pick individual transconjugant colonies and grow them in liquid media with antibiotic selection for the pCRISPR plasmid. Perform a minimum inhibitory concentration (MIC) assay against the antibiotic whose resistance gene was targeted (e.g., Meropenem). A significant reduction in MIC compared to the wild-type recipient strain indicates successful sensitization.
Protocol 2: Biofilm Disruption via Phage-Delivered CRISPR-Cas
This protocol describes the use of engineered lytic bacteriophages to deliver a CRISPR-Cas system targeting a biofilm-related gene (e.g., pelA for EPS production in P. aeruginosa) [60] [61].
Research Reagent Solutions:
- Engineered Phage Stock: A lytic phage (e.g., a T4 or T7 derivative) engineered to carry a CRISPR-Cas cassette targeting the
pelAgene. - Bacterial Strain: P. aeruginosa biofilm-forming strain.
- Growth Media: Tryptic Soy Broth (TSB) or suitable medium for the target pathogen.
- Biofilm Substrate: 96-well polystyrene microtiter plates.
- Staining Solution: 0.1% (w/v) Crystal Violet solution.
- Destaining Solution: 30% (v/v) Acetic acid or 70% (v/v) Ethanol.
Procedure:
- Biofilm Formation: Grow the bacterial strain overnight and dilute 1:100 in fresh TSB. Dispense 200 µL per well into a 96-well plate. Incubate statically at 37°C for 24-48 hours to allow biofilm formation.
- Phage Application: Carefully remove the planktonic culture and gently wash the adhered biofilm twice with PBS. Add 200 µL of fresh TSB containing the engineered phage at a pre-optimized multiplicity of infection (MOI, e.g., 10). Include a no-phage control.
- Incubation: Incubate the plate for another 24 hours at 37°C.
- Biofilm Quantification (Crystal Violet Assay): a. Remove the treatment medium and wash the biofilm gently with PBS. b. Fix the biofilm by air-drying for 45 minutes. c. Stain with 200 µL of 0.1% Crystal Violet per well for 20 minutes. d. Wash thoroughly with water to remove unbound dye. e. Destain with 200 µL of 30% acetic acid for 30 minutes. f. Transfer 100 µL of the destained solution to a new plate and measure the absorbance at 595 nm. Lower absorbance indicates biofilm disruption.
- Confocal Microscopy Validation: For visual confirmation, form biofilms on glass-bottom dishes, treat with phages, and stain with a LIVE/DEAD BacLight bacterial viability kit. Image using confocal laser scanning microscopy (CLSM) to observe biofilm architecture and bacterial viability.
Protocol 3: Synergistic Therapy Using CRISPR-Loaded Nanoparticles
This protocol employs lipid nanoparticles (LNPs) co-loaded with CRISPR-Cas9 RNP targeting a resistance gene (e.g., mecA in MRSA) and an antibiotic (e.g., Oxacillin) to synergistically treat biofilms [13] [62].
Research Reagent Solutions:
- CRISPR RNP: Pre-complexed Cas9 protein and sgRNA (targeting
mecA). - Lipid Nanoparticles (LNPs): Comprising cationic lipids, phospholipids, cholesterol, and PEG-lipids.
- Antibiotic: Oxacillin sodium salt.
- Biofilm Model: Staphylococcus aureus (MRSA) biofilm grown in a 96-well plate.
- Viability Reagent: Resazurin (Alamar Blue) cell viability reagent.
Procedure:
- NP Formulation and Co-loading: a. Prepare the aqueous phase: Dilute the CRISPR RNP and Oxacillin in a citrate buffer (pH 4.0). b. Prepare the lipid phase: Dissolve lipid components in ethanol. c. Use microfluidics or rapid mixing to combine the two phases, forming LNPs that encapsulate both RNP and antibiotic. d. Dialyze the LNP formulation against PBS to remove ethanol and achieve buffer exchange.
- Biofilm Treatment: Grow MRSA biofilms in a 96-well plate as described in Protocol 4.2. After washing, treat the biofilms with:
- LNPs with RNP (anti-
mecA) + Oxacillin - Free Oxacillin (at the same concentration)
- LNPs with non-targeting RNP + Oxacillin
- PBS control
- LNPs with RNP (anti-
- Viability Assay: After 24 hours of incubation, remove the treatment, wash the biofilm, and add fresh medium containing Resazurin. Incubate for 2-4 hours and measure fluorescence (Excitation: 560 nm, Emission: 590 nm). The highest reduction in viability is expected in the group receiving the combination therapy.
- Gene Editing Confirmation: Extract genomic DNA from treated biofilms. Perform PCR amplification of the
mecAtarget region and analyze by Sanger sequencing or T7 Endonuclease I assay to confirm targeted mutation.
Experimental Workflows and System Schematics
The following diagrams, generated using Graphviz DOT language, illustrate the core workflows and mechanisms of the delivery systems.
Diagram 1: Conjugative Plasmid Workflow for Resistance Reversal
Diagram 2: Phage vs. Nanoparticle Delivery Mechanisms
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for CRISPR Delivery Experiments
| Reagent / Material | Function / Description | Example Application / Note |
|---|---|---|
| Conjugative pCRISPR Plasmid | A self-transmissible plasmid encoding Cas nuclease and guide RNA(s). | Should be designed to be incompatible with the target pAMR for enhanced clearance in low-copy number plasmids [59]. |
| Engineered Lytic Bacteriophage | A virus modified to carry CRISPR-Cas genetic payload. | Prefer lytic over temperate phages to avoid lysogeny and ensure bacterial lysis [60]. |
| Cationic Lipid Nanoparticles (LNPs) | Synthetic carriers for encapsulating CRISPR RNP and antibiotics. | Protect payload from degradation and enhance cellular uptake; allows for co-delivery strategies [13] [62]. |
| Gold Nanoparticles (AuNPs) | Metallic nanoparticles for complexing with or delivering CRISPR components. | Can enhance gene-editing efficiency (e.g., 3.5-fold increase reported) and exhibit intrinsic antibacterial properties [13]. |
| CRISPR Ribonucleoprotein (RNP) | Pre-assembled complex of Cas protein and guide RNA. | Offers rapid activity and reduced off-target effects compared to plasmid DNA delivery. |
| Selection Antibiotics | Antibiotics for selecting and maintaining plasmids in bacterial cultures. | Critical for maintaining selective pressure during conjugation and transconjugant culture. |
| Crystal Violet | Dye for staining and quantifying total biofilm biomass. | Standard, low-cost method for high-throughput screening of anti-biofilm agents [1]. |
| Resazurin Viability Reagent | Cell-permeant dye used to measure metabolic activity in biofilms. | Provides a fluorescence-based readout for cell viability after treatment. |
| Confocal Laser Scanning Microscope | Instrument for high-resolution 3D imaging of biofilms. | Essential for validating structural disruption of biofilms and spatial distribution of effects. |
The human gut microbiome, a complex ecosystem of trillions of microorganisms, plays a fundamental role in host health by regulating metabolism, immune function, and pathogen defense through a mechanism known as colonization resistance [66]. Traditional broad-spectrum antibiotic therapy, while effective against pathogenic bacteria, causes widespread collateral damage to these beneficial microbial communities. This disruption, termed dysbiosis, is characterized by a significant reduction in microbial diversity, a loss of key commensal taxa, and an increased susceptibility to opportunistic pathogens like Clostridioides difficile [67] [66]. Furthermore, antibiotic use intensifies selective pressures that promote the horizontal gene transfer of antibiotic resistance genes (ARGs), thereby fueling the crisis of multidrug-resistant organisms [13] [66].
In contrast, precision antimicrobials represent a paradigm shift towards targeted therapeutic strategies. These approaches are designed to selectively eliminate pathogenic bacteria or disable their virulence mechanisms while preserving the integrity of the commensal microbiota [68]. The deployment of CRISPR-Cas systems delivered via conjugative plasmids for the specific elimination of antibiotic resistance genes in biofilm-associated pathogens exemplifies this innovative approach [13] [69]. This application note provides a structured comparison of these two strategies and details experimental protocols for evaluating their impact on the microbiome within the context of a research thesis on conjugative plasmid delivery of CRISPR systems for biofilm control.
Quantitative Comparison of Microbiome Impact
The table below summarizes the key differential impacts of broad-spectrum antibiotics versus precision killing strategies on the gut microbiome, based on current literature.
Table 1: Comparative Impact of Broad-Spectrum Antibiotics and Precision Antimicrobials on the Gut Microbiome
| Parameter | Broad-Spectrum Antibiotics | Precision Antimicrobials (e.g., CRISPR-based) |
|---|---|---|
| Microbial Diversity | Significantly reduces alpha diversity; effects can persist for years [67] [66]. | Aims to preserve native community structure and diversity by targeting specific genetic elements [68]. |
| Colonization Resistance | Severely compromises, increasing risk of infections (e.g., C. difficile) and pathogen colonization (e.g., CRE) [70] [66]. | Seeks to maintain ecological barriers against pathogens by sparing commensals [68]. |
| Antibiotic Resistance Selection | Promotes horizontal gene transfer and enriches for antibiotic-resistant strains [13] [66]. | Directly targets and disrupts antibiotic resistance genes, potentially re-sensitizing bacteria [13] [69]. |
| Key Commensal Taxa | Depletes beneficial genera (e.g., Bifidobacterium, Faecalibacterium) [66]. | Hypothesized to have minimal impact on non-targeted commensals, though off-target effects require validation. |
| Recovery Trajectory | Recovery is variable and may be incomplete; influenced by drug spectrum, duration, and host factors [67] [66]. | Predictive models suggest a more rapid and stable return to pre-treatment state due to reduced collateral damage [71]. |
| Mechanism of Community Disruption | Non-selective bactericidal/bacteriostatic action; secondary nutrient reshuffling alters competitive dynamics [71]. | Precise genetic manipulation (e.g., gene knockout) or anti-virulence action; minimal perturbation to nutrient landscape. |
Experimental Protocols for Impact Assessment
Protocol 1: In Vitro Microbiome Model Assay for Ecodynamic Profiling
This protocol is designed to quantitatively assess the ecological impact of a precision antimicrobial versus a broad-spectrum antibiotic on a defined human gut microbial community.
I. Research Reagent Solutions
Table 2: Essential Reagents for In Vitro Microbiome Impact Studies
| Item | Function/Description |
|---|---|
| Fecal Sample Donors | Source of complex gut microbiota. Use pooled samples from multiple healthy donors to capture diversity [71]. |
| Anaerobic Chamber | Maintains a oxygen-free atmosphere (e.g., with 85% N₂, 10% CO₂, 5% H₂) for cultivating obligate anaerobic gut bacteria. |
| Gut Microbiota Medium (GMM) | A complex, chemically defined culture medium that supports the growth of a wide diversity of gut microbes. |
| Broad-Spectrum Antibiotic Control | e.g., Ciprofloxacin (a fluoroquinolone) or a β-lactam, known to cause significant dysbiosis [66]. |
| Conjugative Plasmid with CRISPR-Cas System | The precision therapeutic, carrying gRNAs targeting specific ARGs (e.g., NDM-1, KPC) in a model pathogen [13]. |
| Donor Strain | A bacterial strain (e.g., E. coli) capable of conjugatively transferring the CRISPR plasmid to the target pathogen. |
| Target Pathogen | A biofilm-forming, antibiotic-resistant strain (e.g., Carbapenem-resistant K. pneumoniae) spiked into the community. |
| DNA Extraction Kit | For high-yield, PCR-inhibitor-free microbial genomic DNA extraction from complex communities. |
| qPCR Assay for 16S rRNA Gene & ARGs | For absolute quantification of total bacterial load and specific resistance gene abundance. |
II. Procedure
- Community Inoculation: Inside an anaerobic chamber, inoculate 100 mL of pre-reduced GMM with 1% (w/v) of a pooled, fresh fecal slurry from healthy donors. Incubate at 37°C with gentle shaking for 48 hours to allow the community to stabilize.
- Experimental Arm Setup: After stabilization, aliquot the community into four separate bioreactors:
- Arm A (Control): No treatment.
- Arm B (Broad-Spectrum): Treat with a clinically relevant concentration of ciprofloxacin (e.g., 0.5 µg/mL).
- Arm C (Precision): Spike with the target pathogen (e.g., CRE at 10^5 CFU/mL) and the donor strain carrying the conjugative CRISPR plasmid.
- Arm D (Pathogen Control): Spike with the target pathogen only, to monitor its natural colonization.
- Sampling and Analysis: Collect samples (1-2 mL) from each arm at T=0, 4, 8, 24, 48, and 72 hours post-treatment.
- Microbial Composition: Perform 16S rRNA gene amplicon sequencing (V4 region) on all samples. Analyze data for alpha-diversity (Shannon Index) and beta-diversity (PCoA of Unifrac distances).
- Pathogen/ARG Burden: Use qPCR to track the abundance of the target pathogen (species-specific gene) and the targeted ARG.
- Community Metabolomics: Analyze supernatant via LC-MS to quantify changes in short-chain fatty acids (butyrate, acetate, propionate) and other key microbial metabolites.
Protocol 2: In Vivo Assessment in a Murine Colonization Model
This protocol evaluates the efficacy and ecological impact of the precision intervention in a live animal model of intestinal colonization.
I. Animal Model and Reagents
- Animals: 6-8 week old, C57BL/6 mice (n=8-10 per group).
- Antibiotic Pre-treatment: To enhance pathogen colonization, administer a cocktail of ampicillin, vancomycin, and metronidazole in drinking water for 3 days prior to pathogen challenge [70].
- Test Articles: The conjugative CRISPR plasmid system, delivered via donor strain. A broad-spectrum antibiotic control (e.g., meropenem) and a vehicle control.
II. Procedure
- Pathogen Challenge: On day 0, orally gavage all mice with a standardized inoculum (e.g., 10^8 CFU) of the target CRE strain.
- Treatment Phase: 24 hours post-challenge, begin treatments administered orally for 5 consecutive days:
- Group 1: Vehicle control.
- Group 2: Broad-spectrum antibiotic control.
- Group 3: Donor strain carrying the conjugative CRISPR plasmid.
- Sample Collection:
- Fecal Pellets: Collect fresh pellets daily throughout the study for CFU enumeration of the total bacteria, target pathogen, and donor strain, as well as for genomic DNA extraction.
- Cecal Content: At endpoint (e.g., day 7), euthanize animals and aseptically collect cecal content for 16S rRNA sequencing, metagenomic shotgun sequencing (to assess ARG repertoire), and metabolomic profiling.
- Outcome Measures:
- Primary Efficacy: Log-reduction in fecal target pathogen CFU counts over time compared to vehicle control.
- Microbiome Impact: Comparison of cecal microbial diversity and composition via 16S sequencing.
- Safety: Weight change, clinical signs, and histological analysis of colon tissue.
Conceptual Framework and Experimental Workflow
The following diagram illustrates the core conceptual contrast between the two therapeutic strategies and their differential ecological outcomes within the gut microbiome.
Diagram 1: Conceptual contrast of therapeutic strategies and ecological outcomes.
The experimental workflow for a comprehensive assessment, integrating both in vitro and in vivo models, is outlined below.
Diagram 2: Integrated experimental workflow for comprehensive assessment.
The global crisis of antimicrobial resistance (AMR) necessitates the development of innovative strategies that extend beyond conventional antibiotic discovery. Conjugative plasmid delivery of CRISPR systems represents a transformative approach for precisely targeting and eliminating antibiotic resistance genes in bacterial pathogens, thereby restoring their antibiotic susceptibility. This paradigm exploits the natural ability of plasmids to horizontally transfer genetic material between bacteria, but redirects this process to deliver molecular countermeasures against resistance mechanisms. By programming CRISPR systems to selectively disrupt resistance genes carried on plasmids or chromosomes, this technology enables the re-sensitization of multidrug-resistant pathogens, effectively reversing evolved resistance and restoring therapeutic efficacy to existing antibiotics [9] [72].
The integration of this approach with biofilm control is particularly valuable, as biofilms represent a critical nexus for AMR persistence and dissemination. Biofilms not only provide a physical barrier against antimicrobial penetration but also create microenvironments that enhance horizontal gene transfer, facilitating the spread of resistance genes among bacterial populations [13]. The extracellular polymeric substance (EPS) matrix of biofilms can reduce antibiotic efficacy by up to 1000-fold compared to planktonic cells, making biofilm-associated infections exceptionally difficult to treat [13]. By employing conjugative plasmids to deliver CRISPR-based countermeasures directly within biofilm communities, researchers can achieve targeted removal of resistance determinants while sparing commensal bacteria, offering a precision medicine approach to combat AMR [9].
Mechanisms of CRISPR-Mediated Re-sensitization
Molecular Foundations of Resistance Reversal
CRISPR systems restore antibiotic susceptibility through several mechanistically distinct pathways, each targeting different components of the bacterial resistance apparatus. The most direct approach involves precision cleavage of antibiotic resistance genes (ARGs) using Cas nucleases programmed with specific guide RNAs (gRNAs). When delivered via conjugative plasmids, these CRISPR components introduce double-strand breaks in target sequences such as β-lactamase genes (bla), methicillin resistance genes (mecA), or carbapenemase genes (ndm-1), permanently disrupting their function and resensitizing bacteria to the corresponding antibiotics [13] [72]. This gene disruption approach is particularly effective against plasmid-borne resistance, as it prevents both the expression of resistance proteins and the horizontal transfer of these genetic elements to neighboring cells.
An alternative strategy employs CRISPR interference (CRISPRi) using a catalytically inactive Cas9 (dCas9) that binds to target DNA without cleaving it, thereby physically blocking transcription. This approach allows for reversible suppression of resistance gene expression and is especially useful for targeting essential genes or evaluating gene function without permanent genetic damage [73]. CRISPRi systems can be designed to simultaneously target multiple resistance mechanisms by employing arrays of gRNAs, creating a multiplexed suppression approach that addresses the redundancy often present in bacterial resistance networks. For biofilm control, CRISPRi has been successfully deployed to silence genes regulating extracellular polymeric substance production (eps), quorum sensing (luxS), and stress response pathways, effectively weakening biofilm integrity and enhancing antibiotic penetration [9] [73].
Table 1: CRISPR Systems for Antibiotic Re-sensitization
| CRSystem Type | Key Components | Primary Mechanism | Target Examples | Advantages |
|---|---|---|---|---|
| CRISPR-Cas9 | Cas9 nuclease, gRNA | DNA cleavage and gene disruption | blaTEM, mecA, ndm-1 | Permanent resistance elimination |
| CRISPRi (dCas9) | dCas9, gRNA | Transcription blocking | ampC, armA, qnrB | Reversible, tunable suppression |
| CRISPRa | dCas9-activator, gRNA | Gene activation | Biofilm dispersion genes | Induces beneficial phenotypes |
| Cas13 | Cas13, crRNA | RNA degradation | mRNA of resistance genes | No genomic alteration |
Conjugative Plasmid Delivery Mechanisms
Conjugative plasmids serve as ideal vectors for CRISPR delivery due to their natural proficiency in horizontal gene transfer between diverse bacterial species. These self-transmissible plasmids encode all necessary machinery for establishing direct cell-to-cell contact (via sex pili) and transferring single-stranded DNA to recipient cells [74] [75]. In the context of CRISPR delivery, helper plasmids can facilitate the transfer of non-conjugative but mobilizable plasmids carrying CRISPR components, enabling their dissemination throughout mixed bacterial populations [74]. This biphasic delivery system allows for the precise engineering of donor strains that subsequently propagate CRISPR payloads to target pathogens in complex microbial communities.
The efficiency of conjugative transfer is influenced by multiple factors, including plasmid host range, recipient cell compatibility, and environmental conditions. Broad-host-range plasmids such as the RP4/RK2 origin plasmids offer particular utility for delivering CRISPR systems across taxonomic boundaries, potentially enabling re-sensitization of diverse pathogen populations [74]. Recent studies have demonstrated that plasmid transfer occurs even in the absence of antibiotic selection pressure, a concerning phenomenon for natural resistance spread but one that can be harnessed therapeutically for CRISPR delivery [74]. For biofilm applications, the conjugative transfer is enhanced within the structured microenvironment of biofilms, where high bacterial density and stabilized cell-cell contacts facilitate efficient plasmid dissemination [9] [13].
Quantitative Assessment of Re-sensitization Efficacy
Metrics for Susceptibility Restoration
Evaluating the success of CRISPR-mediated re-sensitization requires quantitative assessment across multiple parameters that collectively document the restoration of antibiotic efficacy. The most fundamental metric is the change in minimum inhibitory concentration (MIC), which directly measures the concentration of antibiotic required to inhibit bacterial growth. Successful re-sensitization typically demonstrates a significant reduction in MIC (often ≥4-fold decrease), potentially restoring clinically resistant isolates to the susceptible category according to established breakpoints (e.g., CLSI or EUCAST guidelines) [76] [77]. For example, studies targeting β-lactamase genes have reported MIC reductions of 8-16 fold for cephalosporins and carbapenems in formerly resistant Enterobacteriaceae [13].
Beyond MIC reduction, researchers should assess re-sensitization efficiency by quantifying the proportion of bacterial cells that regain antibiotic susceptibility following CRISPR treatment. This is typically measured by comparing colony-forming unit (CFU) counts on antibiotic-containing versus antibiotic-free media after CRISPR delivery. High-efficiency systems can achieve 85-99% re-sensitization in laboratory strains, though efficiency may vary in clinical isolates and within biofilm environments [9] [72]. Additional important metrics include plasmid cure rates (frequency of resistance plasmid elimination), collateral sensitivity (increased susceptibility to unrelated antibiotics), and fitness costs (impact on bacterial growth kinetics following resistance removal) [77].
Table 2: Key Metrics for Quantifying Re-sensitization
| Metric | Measurement Method | Interpretation | Typical Successful Outcome |
|---|---|---|---|
| MIC Reduction | Broth microdilution | Decreased antibiotic concentration needed for inhibition | ≥4-fold reduction, crossing clinical breakpoint |
| Re-sensitization Efficiency | CFU counting on selective vs. non-selective media | Proportion of population that regains susceptibility | >90% in pure culture; >70% in biofilms |
| Plasmid Cure Rate | Plasmid isolation and PCR | Frequency of resistance plasmid loss | >50% for targeted elimination |
| Biofilm Disruption | Crystal violet staining/confocal microscopy | Reduction in biofilm biomass and viability | ≥80% biomass reduction in targeted approaches |
| Collateral Sensitivity | Cross-screening against antibiotic panels | Increased susceptibility to other drug classes | 2-8 fold MIC reduction for collateral drugs |
Advanced Biofilm-Specific Assessment
For biofilm-associated pathogens, standard planktonic susceptibility testing provides an incomplete picture of re-sensitization efficacy. Instead, researchers should employ biofilm-specific susceptibility assays that account for the unique physiological state and protective matrix of biofilm communities. The minimum biofilm eradication concentration (MBEC) assay measures the antibiotic concentration required to eradicate biofilm-grown bacteria, typically yielding values 10-1000 times higher than MIC for planktonic cells [13]. Successful CRISPR-mediated re-sensitization should demonstrate a significant reduction in MBEC, indicating improved antibiotic penetration and efficacy within the biofilm architecture.
Additional biofilm-specific metrics include quantification of biofilm biomass (via crystal violet staining or optical density), assessment of biofilm viability (using live/dead staining coupled with confocal microscopy), and evaluation of biofilm structure alterations (through computational analysis of 3D biofilm images) [9] [73]. For example, studies combining CRISPR delivery with nanoparticle carriers have reported up to 90% reduction in Pseudomonas aeruginosa biofilm biomass and 3.5-fold enhancement in gene-editing efficiency compared to non-carrier systems [13]. These quantitative assessments provide comprehensive evaluation of how CRISPR-mediated re-sensitization impacts the structural integrity and antibiotic tolerance of biofilm communities.
Experimental Protocols
Protocol 1: Conjugative Plasmid Delivery of CRISPR Systems
This protocol describes the delivery of CRISPR re-sensitization constructs to antibiotic-resistant pathogens via conjugative plasmid transfer, with specific applications for biofilm-associated bacteria.
Materials:
- Donor strain: E. coli HB101 containing helper plasmid (e.g., pRK2013) and mobilizable CRISPR plasmid
- Recipient strain: Antibiotic-resistant target pathogen (e.g., MRSA, CRE)
- Growth media: LB broth, LB agar plates
- Antibiotics for selection: Kanamycin (50 µg/mL), Ampicillin (100 µg/mL), Chloramphenicol (25 µg/mL)
- Conjugation filters: 0.22µm cellulose nitrate membrane filters
- Phosphate buffered saline (PBS), pH 7.4
Procedure:
- Pre-culture preparation: Grow donor and recipient strains overnight in LB broth with appropriate antibiotics at 37°C with shaking (200 rpm).
- Cell harvesting: Centrifuge 1 mL of each culture at 5,000 × g for 5 min, wash twice with PBS to remove antibiotics.
- Cell mixing: Mix donor and recipient cells at a 1:3 ratio (donor:recipient) in 100 µL PBS.
- Filter mating: Pipette the cell mixture onto a sterile membrane filter placed on pre-warmed LB agar without antibiotics. Incubate at 37°C for 4-6 hours.
- Cell recovery: Transfer filter to 1 mL PBS and vortex to resuspend cells.
- Transconjugant selection: Plate serial dilutions on selective media containing antibiotics that select for the CRISPR plasmid while counter-selecting the donor strain (e.g., kanamycin + ampicillin for plasmid selection and donor counter-selection).
- Verification: Incubate plates at 37°C for 24-48 hours, then count transconjugant colonies to calculate conjugation frequency (transconjugants per recipient).
- CRISPR induction: For inducible systems, add inducer (aTc, 100 ng/mL or IPTG, 0.5 mM) to activate CRISPR function.
Troubleshooting:
- Low conjugation efficiency: Optimize donor:recipient ratio, extend mating time, or use fresh mid-log phase cultures.
- High donor background: Optimize counter-selection antibiotic concentrations.
- Variable re-sensitization: Verify guide RNA specificity and Cas expression levels.
Protocol 2: Assessment of Re-sensitization in Biofilm Models
This protocol details the quantification of antibiotic susceptibility restoration in biofilm-grown pathogens following CRISPR treatment.
Materials:
- CRISPR-treated and control bacterial strains
- 96-well polystyrene microtiter plates for biofilm formation
- Appropriate growth media (e.g., TSB, BHI)
- Antibiotic stock solutions for MIC/MBEC testing
- Crystal violet solution (0.1%)
- Phosphate buffered saline (PBS), pH 7.4
- Live/Dead BacLight Bacterial Viability Kit (or equivalent)
- Confocal microscopy equipment
Procedure: Biofilm formation:
- Dilute overnight cultures of CRISPR-treated and control strains to ~10⁶ CFU/mL in fresh media.
- Dispense 200 µL per well into 96-well microtiter plates.
- Incubate under appropriate conditions (e.g., 37°C for 24-48 hours) for biofilm development.
MBEC assay:
- Carefully aspirate planktonic cells and rinse established biofilms twice with PBS.
- Add serial dilutions of target antibiotic in fresh media (typically 2-256× MIC).
- Incubate for 24 hours at appropriate temperature.
- Aspirate antibiotic solutions and rinse biofilms with PBS.
- Disrupt biofilms by scraping and vortexing in PBS.
- Determine viable counts by serial dilution and plating.
Biofilm biomass quantification:
- After antibiotic treatment, fix biofilms with 99% methanol for 15 minutes.
- Stain with 0.1% crystal violet for 20 minutes.
- Rinse thoroughly with water to remove unbound dye.
- Extract bound dye with 33% acetic acid.
- Measure absorbance at 590 nm.
Confocal microscopy analysis:
- Grow biofilms on appropriate surfaces (e.g., glass coverslips).
- Stain with Live/Dead solution according to manufacturer's instructions.
- Image using confocal microscope with appropriate filters.
- Quantify biofilm thickness, biovolume, and live/dead ratio using image analysis software (e.g., ImageJ, COMSTAT).
Data analysis:
- Calculate MBEC as the lowest antibiotic concentration that reduces biofilm viability by ≥99.9%.
- Compare biofilm biomass between CRISPR-treated and control groups using statistical tests (t-test, ANOVA).
- Correlate re-sensitization with specific gene modifications using PCR and sequencing.
Visualization of Pathways and Workflows
CRISPR Re-sensitization Workflow: This diagram illustrates the complete experimental pathway from conjugative plasmid delivery through CRISPR-mediated re-sensitization to quantitative assessment of restored antibiotic susceptibility in biofilm models.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Conjugative CRISPR Delivery Studies
| Reagent Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| Conjugative Systems | RP4/RK2 origin plasmids, pRK2013 (helper), F-plasmid derivatives | Enable horizontal transfer of CRISPR constructs | Host range, transfer efficiency, stability in recipients |
| CRISPR Plasmids | pCas9, pCRISPR, dCas9-derived plasmids | Provide Cas nuclease and gRNA expression | Inducible vs. constitutive, gRNA multiplexing capacity |
| Selection Antibiotics | Kanamycin, Ampicillin, Chloramphenicol, Tetracycline | Maintain plasmids and select for transconjugants | Compatibility with recipient strain resistance profiles |
| Inducers | Anhydrotetracycline (aTc), IPTG, Arabinose | Regulate CRISPR system activation | Concentration optimization, timing of induction |
| Biofilm Growth Surfaces | Polystyrene microtiter plates, Silicone coupons, Glass coverslips | Support biofilm development for testing | Surface properties influencing attachment |
| Viability Assays | Live/Dead BacLight, CTC-DAPI, Resazurin | Differentiate live vs. dead cells in biofilms | Compatibility with imaging methods, quantification approach |
| Imaging Tools | Confocal laser scanning microscopy, SEM, Epifluorescence | Visualize biofilm architecture and composition | Resolution, 3D reconstruction capability |
The integration of conjugative plasmid delivery with CRISPR-based gene editing represents a paradigm shift in our approach to combating antibiotic resistance. By directly targeting the genetic basis of resistance while simultaneously disrupting protective biofilm structures, this technology offers a path to restore the efficacy of existing antibiotics against currently untreatable infections. The experimental frameworks and quantitative assessment protocols outlined here provide researchers with robust methodologies to evaluate and optimize these approaches across diverse bacterial pathogens and resistance mechanisms.
Future developments in this field will likely focus on enhancing the precision and specificity of CRISPR delivery systems, expanding the host range of conjugative vectors, and developing regulatory frameworks for potential clinical applications. The combination of CRISPR-mediated re-sensitization with conventional antibiotics holds particular promise for addressing biofilm-associated infections, potentially extending the useful lifespan of our current antimicrobial arsenal while new therapeutic strategies are developed. As this technology advances, it will be essential to maintain rigorous assessment protocols and safety standards to ensure effective translation from laboratory research to clinical application.
Economic and Scalability Considerations for Clinical Translation
The translation of conjugative plasmid delivery systems for CRISPR-based biofilm control from laboratory research to clinical application presents a complex interplay of technical efficacy, economic viability, and scalable manufacturing. These systems, which harness bacterial conjugation to deliver DNA-targeting CRISPR nucleases specifically against antibiotic-resistant pathogens, represent a promising alternative to conventional antibiotics [25] [14]. However, their pathway to clinical deployment requires careful consideration of production costs, process scalability, and regulatory hurdles. This application note examines these considerations within the context of a research thesis on conjugative CRISPR systems, providing structured quantitative data, detailed protocols for key experiments, and analysis of scaling challenges to inform researchers and drug development professionals.
Key Performance and Economic Data
The assessment of clinical potential relies on key performance metrics and their associated costs. The tables below summarize experimental efficacy data and economic considerations for conjugative CRISPR systems.
Table 1: Experimental Efficacy of Conjugative CRISPR Systems Against Bacterial Targets
| Target Organism | Delivery Plasmid | Conjugation Efficiency | Killing/Resistance Reduction | Key Resistance Gene Targeted | Citation |
|---|---|---|---|---|---|
| Enterococcus faecalis | pKH88 (pPD1-based) | High (in murine intestine) | Several orders of magnitude | ermB (erythromycin), tetM (tetracycline) | [25] |
| Salmonella enterica | pNuc-cis (RK2-based) | Up to ~100% (with beads) | High killing efficiency | Non-essential and essential genes | [14] |
| Escherichia coli (Donor) to S. enterica (Recipient) | pNuc-trans (RK2-based) | ~1x10⁻⁵ to ~1x10⁻³ | Demonstrated | Various chromosomal sites | [14] |
Table 2: Economic and Scalability Analysis of Conjugative CRISPR Systems
| Consideration Factor | Technical/Scalability Challenge | Potential Economic Impact | Possible Mitigation Strategy |
|---|---|---|---|
| Delivery Efficiency | Low conjugation frequency in complex microbiomes [14]. | Increased dose requirements, higher cost of goods (COGs). | Engineer high-efficiency cis-conjugative plasmids [14]. |
| Manufacturing & Purification | Production of pure, clinical-grade plasmid DNA and delivery vehicles. | Significant capital investment and process development costs. | Leverage existing Good Manufacturing Practice (GMP) platforms for nucleic acid production. |
| Specificity & Off-Targets | Potential cytotoxicity associated with Cas9 [78]. | Pre-clinical safety testing costs, risk of trial failure. | Utilize high-fidelity Cas variants and rigorous bioinformatics guide RNA design. |
| Regulatory Pathway | Ambiguity for live biocontainment vehicles and engineered plasmids [78]. | Protracted timeline, uncertain regulatory requirements. | Early engagement with regulatory agencies, establish robust characterization and tracking methods. |
Detailed Experimental Protocols
Protocol: In Vitro Assessment of Conjugative Transfer and Killing Efficiency
This protocol is adapted from methods used to demonstrate high-frequency conjugation and targeted killing of Salmonella enterica [14].
I. Research Reagent Solutions
| Item | Function/Description |
|---|---|
| pNuc-cis Plasmid | Conjugative plasmid encoding both CRISPR nuclease (e.g., TevSpCas9) and guide RNA(s) under inducible promoter (e.g., pBAD) [14]. |
| Donor Strain | E. coli DH5α or similar strain harboring the pNuc-cis plasmid. |
| Recipient Strain | Target pathogen (e.g., S. enterica, E. faecalis) with a defined chromosomal target. |
| Low-Salt LB (LSLB) Media | Enhances conjugation efficiency; contains 0.25% w/v NaCl [14]. |
| Glass Beads (0.5mm) | Provides solid surface to enhance cell-to-cell contact and biofilm-like conditions, boosting conjugation [14]. |
| Inducer Solution | L-Arabinose solution to induce nuclease expression from the pBAD promoter post-conjugation. |
| Selective Agar Plates | Contains antibiotics to selectively count donor, recipient, and transconjugant colonies. |
II. Methodology
Strain Preparation:
- Inoculate donor (E. coli with pNuc-cis) and recipient (target pathogen) strains in separate liquid cultures and grow overnight at 37°C with appropriate antibiotics.
- Sub-culture the strains to mid-log phase (OD₆₀₀ ~0.5).
Conjugation in Liquid Culture with Beads:
- Mix donor and recipient cells in a 10:1 ratio in a tube containing LSLB media and 0.5mm glass beads.
- Incubate the tube at 37°C for up to 72 hours with mild agitation (60 RPM) [14].
Induction of CRISPR Nuclease:
- After conjugation, induce the culture with a predetermined concentration of L-arabinose (e.g., 0.2%) to express the TevSpCas9 nuclease.
Enumeration and Efficiency Calculation:
- Serially dilute the conjugation mixture at various time points.
- Plate on selective agar to differentiate and count donors, recipients, and transconjugants.
- Conjugation Frequency = (Number of Transconjugants CFU) / (Number of Recipients CFU).
- Killing Efficiency: Compare the viable count of recipients in the induced culture versus a non-induced control or a control with a non-targeting guide RNA.
Protocol: Validating CRISPR Editing and Essential Controls
This protocol outlines critical controls to confirm that observed phenotypic effects are due to specific CRISPR-mediated killing [79].
I. Research Reagent Solutions
| Item | Function/Description |
|---|---|
| Positive Editing Control | A validated guide RNA with known high editing efficiency in the target strain (e.g., targeting a non-essential gene like lacZ or a known essential gene) [79]. |
| Negative Editing Control (Scramble) | A guide RNA with no complementary sequence in the host genome, controlling for non-specific effects of Cas9 or RNA delivery [79]. |
| Mock Control | Cells subjected to the same transfection/conjugation protocol but without delivery of Cas9 or guide RNA, controlling for stress induced by the experimental procedure [79]. |
| Donor-Only Control | Culture containing only the donor strain to confirm the selectivity of the agar plates used for transconjugant enumeration. |
II. Methodology
Control Setup:
- In parallel with the main experiment, perform separate conjugation assays using:
- Donor strain with pNuc-cis plasmid containing the positive control guide RNA.
- Donor strain with pNuc-cis plasmid containing a scrambled guide RNA.
- A mock control where recipients are processed identically but without exposure to donors.
- In parallel with the main experiment, perform separate conjugation assays using:
Analysis:
- For the positive control: A high killing efficiency validates that the conjugation and CRISPR machinery are functional under the experimental conditions.
- For the negative control (scramble): The absence of significant killing indicates that observed effects in the main experiment are sequence-specific.
- For the mock control: Establishes the baseline viability of recipient cells without any conjugative pressure.
Visualization of Workflows and Pathways
The following diagrams illustrate the core experimental workflow and the decision-making pathway for clinical translation, highlighting key economic and scalability checkpoints.
Conclusion
The integration of conjugative plasmid delivery with CRISPR-Cas technology represents a paradigm shift in our approach to combating biofilm-mediated antimicrobial resistance. This strategy offers unparalleled precision in targeting genetic resistance determinants and key virulence pathways directly within the challenging biofilm microenvironment. While significant progress has been made—demonstrated by high-efficiency conjugation systems and successful biofilm disruption in models—key challenges in optimal delivery, safety, and regulatory approval remain. Future directions must focus on refining in vivo delivery platforms, developing robust resistance management strategies, and advancing towards targeted clinical trials for chronic infections. Success in this field promises to unlock a new class of 'smart' antimicrobials that precisely edit pathogenic traits, restoring the efficacy of our existing antibiotic arsenal and bolstering the global fight against AMR.
References
- 1. Biofilms: structure, resistance mechanism, emerging ... [https://pubs.rsc.org/en/content/articlehtml/2025/pm/d5pm00094g]
- 2. Mechanisms of antimicrobial resistance in biofilms [https://www.nature.com/articles/s44259-024-00046-3]
- 3. The Role of Bacterial Biofilms in Antimicrobial Resistance [https://asm.org/articles/2023/march/the-role-of-bacterial-biofilms-in-antimicrobial-re]
- 4. Biofilm Resilience: Molecular Mechanisms Driving Antibiotic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11852148/]
- 5. A comprehensive review on biofilm-associated infections [https://www.sciencedirect.com/science/article/pii/S2950194625002043]
- 6. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-025-04323-4]
- 7. The Role of Bacterial Biofilm in Antibiotic Resistance and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7468660/]
- 8. Antibiotics versus biofilm: an emerging battleground in ... [https://aricjournal.biomedcentral.com/articles/10.1186/s13756-019-0533-3]
- 9. CRISPR–Cas systems as emerging tools for precision ... [https://www.sciencedirect.com/science/article/abs/pii/S0963996925021416]
- 10. CRISPR-Cas systems as emerging tools for precision ... [https://pubmed.ncbi.nlm.nih.gov/41271362/?utm_source=FeedFetcher&utm_medium=rss&utm_campaign=None&utm_content=14MpKbWO9zvAeasdR__ANU13Wb_0XmzdDkWFWNOZzBCk4IB5C&fc=None&ff=20251122111023&v=2.18.0.post22+67771e2]
- 11. Biofilms: Understanding the structure and contribution ... [https://www.sciencedirect.com/science/article/pii/S2590097823000095]
- 12. Bacterial Persister Cells and Development of Antibiotic ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11335562/]
- 13. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366227/]
- 14. Efficient inter-species conjugative transfer of a CRISPR ... [https://www.nature.com/articles/s41467-019-12448-3]
- 15. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm- ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2024.1408569/full]
- 16. CRISPR-Cas Systems: Bridging Bacterial Immunity and ... [https://www.mdpi.com/2673-8007/5/4/118]
- 17. CRISPR-Cas Systems: Transformative Precision Tools to ... [https://www.genesispub.org/crispr-cas-systems-transformative-precision-tools-to-combat-antimicrobial-resistance-in-multidrug-resistant-gram-negative-pathogens]
- 18. An updated evolutionary classification of CRISPR–Cas ... [https://www.nature.com/articles/s41564-025-02180-8]
- 19. BaeR and H-NS control CRISPR-Cas-mediated immunity and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12625773/]
- 20. Cas3 of type I-Fa CRISPR-Cas system upregulates ... [https://www.nature.com/articles/s42003-025-08124-6]
- 21. CRISPR-Cas9 and Its Bioinformatics Tools: A Systematic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12109910/]
- 22. Plasmid Transfer by Conjugation in Gram-Negative Bacteria [https://pmc.ncbi.nlm.nih.gov/articles/PMC7690428/]
- 23. Bacterial conjugation [https://en.wikipedia.org/wiki/Bacterial_conjugation]
- 24. Bacterial Conjugation – Conjugation Biology Explained [https://www.technologynetworks.com/immunology/articles/bacterial-conjugation-conjugation-biology-explained-397228]
- 25. Conjugative Delivery of CRISPR-Cas9 for the Selective ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6811441/]
- 26. Bacterial Conjugation- Definition, Principle, Process, ... [https://microbenotes.com/bacterial-conjugation/]
- 27. Harnessing bacterial immunity: CRISPR-Cas system as a ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1588446/full]
- 28. CRISPR-Cas-Based Antimicrobials: Design, Challenges, ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10353011/]
- 29. Harnessing the CRISPR-Cas Systems to Combat ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.716064/full]
- 30. Plasmids, a Molecular Cornerstone of Antimicrobial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12440250/]
- 31. Engineered phage with antibacterial CRISPR–Cas ... [https://www.nature.com/articles/s41587-023-01759-y]
- 32. A CRISPR-Cas9 system protecting E. coli against ... [https://www.nature.com/articles/s41598-025-85334-2]
- 33. Technologies and Computational Analysis Strategies for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7497852/]
- 34. How To Use ICE: A Detailed Guide for Analyzing CRISPR ... [https://www.synthego.com/guide/how-to-use-crispr/ice-analysis-guide/]
- 35. Step-by-Step Guide for Analyzing CRISPR Editing Results ... [https://www.editco.bio/step-by-step-ice-guide]
- 36. A mathematician's guide to plasmids [https://pmc.ncbi.nlm.nih.gov/articles/PMC10433428/]
- 37. Comparative Genomics of the Conjugation Region of F-like ... [https://www.frontiersin.org/journals/molecular-biosciences/articles/10.3389/fmolb.2016.00071/full]
- 38. Single-cell measurement of plasmid copy number and ... [https://www.nature.com/articles/s41467-021-21734-y]
- 39. Removal of AMR plasmids using a mobile, broad host- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10268836/]
- 40. Superior Conjugative Plasmids Delivered by Bacteria to ... [https://www.sciencedirect.com/science/article/abs/pii/S2693125724000700]
- 41. A three-plasmid-containing CRISPR-Cas9 platform to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12542745/]
- 42. Biofilm architecture determines the dissemination of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12054802/]
- 43. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11257871/]
- 44. Current Knowledge on CRISPR Strategies Against ... [https://www.mdpi.com/2079-6382/13/12/1141]
- 45. Quantification and Comparison of Different Biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12388762/]
- 46. Innovative approaches to combat antibiotic resistance ... [https://www.springermedicine.com/antibiotic/innovative-approaches-to-combat-antibiotic-resistance-integratin/51360102]
- 47. Efficient inter-species conjugative transfer of a CRISPR nuclease for... [https://www.nature.com/articles/s41467-019-12448-3?error=cookies_not_supported&code=c9a7c9ec-53a7-4bfa-ada7-c96ef7a42a3f]
- 48. formation and control Biofilm of foodborne pathogens: food... strategies [https://pubs.rsc.org/en/content/articlehtml/2017/ra/c7ra02497e]
- 49. Bacteria Gang Together in Killer Biofilms , but... | Scientific American [https://www.scientificamerican.com/article/bacteria-gang-together-in-killer-biofilms-but-scientists-can-disrupt-gang-communications/]
- 50. Off-target interactions in the CRISPR-Cas9 Machinery [https://www.sciencedirect.com/science/article/pii/S2405580825002213]
- 51. Enhancing Specificity, Precision, Accessibility, Flexibility ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12302564/]
- 52. A quantitative model for the dynamics of target recognition ... [https://www.nature.com/articles/s41467-022-35116-5]
- 53. Quantifying On and Off-Target Genome Editing - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4308725/]
- 54. The hidden risks of CRISPR/Cas: structural variations and ... [https://www.nature.com/articles/s41467-025-62606-z]
- 55. Advances in Nanomaterial‐Mediated CRISPR/Cas ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1669104/abstract]
- 56. CRISPR-Cas and restriction–modification systems are ... [https://www.nature.com/articles/ncomms3087]
- 57. In Vitro Efficacy of Nonantibiotic Treatments on Biofilm ... [https://pubmed.ncbi.nlm.nih.gov/26719448/]
- 58. Blue laser light inhibits biofilm formation in vitro and in vivo ... [https://www.nature.com/articles/s41522-019-0102-9]
- 59. Effectiveness of CRISPR-Cas in sensitizing bacterial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12606438/]
- 60. CRISPR-Cas mediated targeting of resistance genes for ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813025097375]
- 61. (PDF) Phage engineering and phage ‐assisted CRISPR ‐Cas delivery ... [https://www.academia.edu/95282324/Phage_engineering_and_phage_assisted_CRISPR_Cas_delivery_to_combat_multidrug_resistant_pathogens]
- 62. Innovative approaches to combat antibiotic resistance [https://pubmed.ncbi.nlm.nih.gov/40830872/]
- 63. Targeting bacterial biofilm-related genes with nanoparticle ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2024.1387114/full]
- 64. Frontiers | Biofilms and multidrug resistance: an emerging crisis and... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1625356/full]
- 65. Combating Antimicrobial Resistance: Innovative Strategies ... [https://www.mdpi.com/1424-8247/18/8/1119]
- 66. The Impact of Antibiotic Therapy on Intestinal Microbiota [https://pmc.ncbi.nlm.nih.gov/articles/PMC12024230/]
- 67. The Impact of Antibiotic Use on the Human Gut Microbiome [https://gmr.scholasticahq.com/article/143628-the-impact-of-antibiotic-use-on-the-human-gut-microbiome-a-review]
- 68. Precision antimicrobial therapeutics: the path of least ... [https://www.nature.com/articles/s41522-018-0048-3]
- 69. Harnessing the Microbiome: CRISPR-Based Gene Editing ... [https://link.springer.com/article/10.1007/s12602-025-10573-8]
- 70. The potential of the microbiome as a target for prevention and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12540337/]
- 71. Medications change our gut microbiome in predictable ways [https://news.stanford.edu/stories/2025/11/medications-gut-microbiome-bacteria-digestive-system-research]
- 72. Precision targeting of food biofilm-forming genes by microbial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9396135/]
- 73. CRISPR interference to interrogate genes that control ... [https://www.nature.com/articles/s41598-019-52400-5]
- 74. Plasmids and the Spread of Antibiotic Resistance Genes [https://asm.org/articles/2023/january/plasmids-and-the-spread-of-antibiotic-resistance-g]
- 75. acquisition and transfer of antibiotic resistance genes in bacteria [https://pmc.ncbi.nlm.nih.gov/articles/PMC2268074/]
- 76. Current and Future Technologies for the Detection of Antibiotic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10606640/]
- 77. Systematic mapping of antibiotic cross-resistance and ... [https://www.nature.com/articles/s41564-024-01857-w]
- 78. Microbial Genome Editing with CRISPR–Cas9 [https://www.mdpi.com/2311-5637/11/7/410]
- 79. Ensure Proper Controls in Your CRISPR Experiments [https://www.synthego.com/blog/controls-crispr-experiments/]