CRISPR vs. Culture: A New Paradigm for Rapid and Sensitive Biofilm Pathogen Detection
This article provides a comprehensive evaluation of CRISPR-based diagnostic technologies against traditional culture methods for detecting pathogens within biofilms.
CRISPR vs. Culture: A New Paradigm for Rapid and Sensitive Biofilm Pathogen Detection
Abstract
This article provides a comprehensive evaluation of CRISPR-based diagnostic technologies against traditional culture methods for detecting pathogens within biofilms. Aimed at researchers, scientists, and drug development professionals, it explores the foundational principles of biofilm complexity and the limitations of conventional diagnostics. The scope extends to the molecular mechanisms of key Cas proteins (Cas9, Cas12, Cas13), their application in detecting polymicrobial infections, and integration with platforms like SHERLOCK and DETECTR. It further addresses critical troubleshooting aspects, such as inhibitor interference and delivery challenges in extracellular polymeric substances (EPS), and presents validation data from clinical and food safety studies. The review concludes by synthesizing performance comparisons and outlining future trajectories, including AI integration and point-of-care device development, for transforming biofilm-related infection management.
The Biofilm Diagnostic Challenge: Why Traditional Culture Methods Fall Short
Bacterial biofilms represent a predominant mode of life for microorganisms in both environmental and clinical settings, characterized by structured communities of cells encased in a self-produced matrix and adherent to surfaces. The intricate architecture of biofilms confers significant survival advantages, including enhanced tolerance to antimicrobial agents and host immune responses, making biofilm-associated infections particularly challenging to diagnose and treat [1]. This architectural fortress is primarily composed of the extracellular polymeric substance (EPS) matrix and harbors subpopulations of dormant cells known as persisters, which together facilitate recurrent infections and complicate clinical management [2] [3].
Within diagnostic microbiology, the protective nature of biofilms presents substantial obstacles for pathogen detection. Conventional culture methods, long considered the gold standard, often fail to accurately detect and identify biofilm-associated pathogens due to the matrix barrier and the heterogeneous metabolic states of embedded cells. Emerging technologies, particularly CRISPR-based diagnostic systems, offer promising alternatives with potential to overcome these limitations through their high sensitivity and specificity for nucleic acid targets, even within complex biofilm samples [4]. This guide provides a comprehensive comparison of these diagnostic approaches within the context of biofilm architecture, empowering researchers and drug development professionals to select appropriate methodologies for their investigative needs.
Architectural Foundations: EPS Matrix and Persister Cells
The EPS Matrix: More Than Just "Slime"
The extracellular polymeric substance (EPS) matrix represents the fundamental scaffolding of biofilms, forming a highly hydrated and biologically active environment that determines the immediate conditions of life for embedded cells. Often metaphorically described as the "house of biofilm cells," the EPS comprises a diverse assortment of biopolymers of microbial origin, including polysaccharides, proteins, glycoproteins, glycolipids, and surprisingly abundant amounts of extracellular DNA (e-DNA) [2]. This complex composition contrasts with the historical perception of biofilms as primarily polysaccharide-based structures.
The matrix functions as a dynamic, multifunctional component that directly contributes to antimicrobial tolerance through several mechanisms:
- Physical barrier: The EPS matrix creates a diffusional barrier that can retard antibiotic penetration into the deeper layers of the biofilm, potentially allowing for antibiotic degradation or neutralization before reaching target cells [1].
- Sorption properties: Charged polymers within the matrix can bind antimicrobial agents, particularly positively charged aminoglycosides, through interactions with negatively charged components like e-DNA, effectively reducing the concentration reaching bacterial cells [1].
- Enzyme retention: The matrix retains extracellular enzymes close to the cells that produced them, creating an "activated matrix" capable of degrading polymers and providing nutrients while potentially inactivating antimicrobial compounds [2].
- Mechanical stability: Interactions including hydrophobic associations, cross-linking by multivalent cations, and polymer entanglements provide structural integrity to maintain spatial arrangements for metabolic cooperation between different species within biofilm microconsortia [2].
Table 1: Key Components of the Biofilm EPS Matrix and Their Functional Roles
| EPS Component | Chemical Nature | Primary Functions | Role in Antimicrobial Tolerance |
|---|---|---|---|
| Polysaccharides | Neutral or charged polymers | Structural scaffolding, water retention, adhesion | Diffusion barrier, antibiotic binding |
| Extracellular DNA (e-DNA) | Double-stranded DNA | Structural integrity, genetic information | Cationic antibiotic sequestration, matrix stability |
| Proteins/Amyloids | Proteinaceous fibrils | Adhesion, structural reinforcement | Matrix stabilization, enzyme retention |
| Membrane Vesicles | Lipid nanostructures | Enzyme transport, communication | Virulence factor delivery, gene exchange |
| Lipids | Hydrophobic compounds | Surface interaction, hydrophobicity | Interface interactions, compound sorption |
Persister Cells: The Dormant Reservoirs
Persister cells represent a non-genetic, phenotypic variant within bacterial populations characterized by a transient, dormant state that confers tolerance to antimicrobial treatment. These cells are genetically identical to their susceptible counterparts but exist in a metabolically inactive or slow-growing state that protects them from antibiotics that typically target active cellular processes [5] [3]. Unlike resistant bacteria that grow in the presence of antibiotics, persister cells do not grow but survive antibiotic exposure and can resume growth once the antibiotic pressure is removed, potentially leading to recurrent infections [5].
The formation and maintenance of persister cells involve several molecular mechanisms:
- Toxin-Antitoxin (TA) Systems: These genetic modules typically consist of a stable toxin that disrupts essential cellular processes and a labile antitoxin that prevents toxicity. Under stress conditions, activation of toxins such as MqsR and TisB can induce dormancy by diminishing translation or reducing proton motive force and ATP levels [5].
- Stringent Response: The alarmone ppGpp, produced during nutrient limitation and other stresses, mediates a global transcriptional reprogramming that promotes persistence through interactions with RNA polymerase and activation of stress response sigma factors [5].
- Metabolic Dormancy: A fundamental characteristic of persisters is their reduced metabolic activity, which protects them from antibiotics that corrupt active cellular processes. This dormancy exists on a continuum from shallow to deep persistence states [3].
The biofilm environment particularly enhances persister formation, with biofilms containing up to 1% persister cells compared to exponentially growing planktonic cultures [5]. This enrichment occurs because the heterogeneous conditions within biofilms, including nutrient and oxygen gradients, create microenvironments conducive to the induction of dormant states. The combination of physical matrix barriers and physiological dormancy makes biofilm-associated infections exceptionally difficult to eradicate with conventional antibiotic therapies [3] [1].
Diagnostic Modalities: CRISPR vs. Culture Methods
Traditional Culture-Based Methods
Culture methods represent the historical foundation of microbiological diagnosis and continue to serve as reference standards in many clinical laboratories. These approaches rely on the growth and propagation of microorganisms on various media, followed by morphological, biochemical, and sometimes molecular characterization. For biofilm-associated pathogens, specific culture techniques have been developed to assess biofilm-forming capacity and study biofilm-related characteristics.
Table 2: Conventional Culture Methods for Biofilm Detection and Analysis
| Method | Experimental Protocol | Key Performance Metrics | Advantages | Limitations |
|---|---|---|---|---|
| Tissue Culture Plate (TCP) Method | Grow bacteria in 96-well plates with media, stain with crystal violet, measure OD570 [6] [7] | Sensitivity: 82-100%, Specificity: 61.5-100% [6] [7] | Quantitative, high-throughput, considered gold standard | Labor-intensive, requires specialized equipment |
| Tube Adherence Method | Inoculate bacteria in tubes with media, stain with crystal violet, visualize biofilm [6] | Sensitivity: 72.7-82%, Specificity: 46.2% [6] [7] | Simple, inexpensive, no special equipment | Semi-quantitative, subjective interpretation |
| Congo Red Agar (CRA) Method | Culture on CRA medium, observe black colony formation [6] [7] | Sensitivity: 78-81.8%, Specificity: 61.5% [6] [7] | Easy to perform, low cost | Variable specificity, qualitative results |
The fundamental limitation of culture methods for biofilm diagnostics lies in their reliance on bacterial growth, which is inherently problematic when dealing with persister cells that exist in a dormant, non-growing state [3]. Additionally, the EPS matrix can impede the release of bacteria from biofilm specimens into culture media, potentially leading to false-negative results. Culture methods also require significant time—typically 24-48 hours for initial results and longer for full identification and susceptibility testing—which delays appropriate therapeutic interventions [4] [6].
CRISPR-Based Diagnostic Systems
CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) systems have evolved from bacterial adaptive immune mechanisms into powerful tools for molecular diagnostics. These systems leverage the programmable recognition and cleavage capabilities of Cas proteins to achieve highly specific and sensitive detection of pathogen nucleic acids, offering distinct advantages for detecting biofilm-associated pathogens that may be difficult to culture [4].
The core mechanism involves two fundamental processes:
- Target Recognition: CRISPR RNA (crRNA) acts as a guide molecule that recognizes specific sequences of target nucleic acids through complementary base pairing. Artificially designed crRNAs can precisely target conserved regions of pathogen nucleic acids, such as bacterial 16S rRNA genes or antibiotic resistance determinants [4].
- Enzymatic Activity Trigger: Upon target recognition, Cas proteins undergo conformational changes that activate their nuclease activities. Cas12 and Cas13 exhibit trans-cleavage activity (non-specific cleavage of surrounding nucleic acids) after binding their target DNA or RNA, respectively, enabling signal amplification through the cleavage of reporter molecules [4].
Several CRISPR-based platforms have been developed for diagnostic applications:
- SHERLOCK (Specific High Sensitivity Enzyme Reporter Unlocking): Utilizes Cas13 for RNA detection with attomolar sensitivity, enabling identification of specific pathogens and antimicrobial resistance markers [4].
- DETECTR (DNA Endonuclease Targeted CRISPR Trans Reporter): Employs Cas12a for DNA detection, achieving sensitivity comparable to PCR-based methods with faster turnaround times [4].
- HOLMES (One-Hour Low-Cost Multipurpose Highly Efficient System): A rapid, multiplexable platform that combines Cas12 with amplification methods for sensitive detection [4].
Table 3: Performance Comparison of CRISPR Systems for Pathogen Detection
| CRISPR System | Target | Sensitivity | Time to Result | Key Advantages for Biofilm Diagnostics | |
|---|---|---|---|---|---|
| Cas9-Based | DNA | ~pM- nM | 2-4 hours | High specificity, programmable targeting | Direct detection without culture |
| Cas12 (DETECTR) | DNA | aM levels | 30-90 minutes | Trans-cleavage activity, rapid detection | Identifies pathogens despite dormancy |
| Cas13 (SHERLOCK) | RNA | aM levels | 1-2 hours | RNA targeting, high sensitivity | Detects viable cells through RNA |
CRISPR diagnostics directly address several limitations of culture methods for biofilm-associated pathogens. Their ability to detect nucleic acids independent of bacterial viability enables identification of persister cells that would not grow in culture [4] [3]. Additionally, the high sensitivity of CRISPR systems allows detection of low bacterial burdens within biofilm specimens, potentially improving diagnostic yield from clinical samples with sparse or difficult-to-culture pathogens.
Comparative Analysis: Diagnostic Performance for Biofilm Pathogens
Technical and Performance Parameters
The selection between CRISPR and culture-based diagnostic approaches requires careful consideration of multiple performance parameters, particularly in the context of biofilm-associated pathogens where both matrix barriers and bacterial dormancy present unique challenges.
Table 4: Comprehensive Comparison of Diagnostic Modalities for Biofilm Pathogens
| Parameter | Culture Methods | CRISPR Diagnostics | Implications for Biofilm Applications |
|---|---|---|---|
| Sensitivity | 10^2-10^3 CFU/mL [6] | aM levels (single molecule detection) [4] | CRISPR superior for low bacterial burden in chronic biofilm infections |
| Time to Result | 24-48 hours (up to weeks for slow-growers) [6] | 30 minutes - 4 hours [4] | CRISPR enables same-day treatment decisions |
| Viability Requirement | Required (limitation for persisters) [3] | Not required | CRISPR detects dormant persisters that evade culture |
| EPS Matrix Interference | Significant (impedes bacterial release) [1] | Minimal with proper sample processing | CRISPR less affected by physical biofilm barriers |
| Species Identification | Required subsequent testing (24-48 additional hours) [6] | Built-in specificity through crRNA design [4] | CRISPR enables simultaneous detection and identification |
| Point-of-Care Applicability | Limited (requires specialized equipment, trained personnel) [4] | High (lateral flow readouts, minimal equipment) [4] | CRISPR suitable for resource-limited settings |
| Antimicrobial Susceptibility | Provides phenotypic AST | Requires separate genetic marker detection | Culture provides direct resistance profiling |
| Cost | Low-moderate (reagent costs) | Moderate (enzyme production, development) | Culture remains more accessible in low-resource settings |
Practical Considerations for Research and Clinical Applications
The optimal diagnostic approach varies depending on the specific application, research question, and available resources. For fundamental biofilm research investigating mechanisms of persistence or matrix composition, CRISPR systems offer powerful tools for probing genetic determinants and metabolic states without the biases introduced by culture. However, for antimicrobial susceptibility testing or when targeting organisms with unknown genetic markers, culture methods remain indispensable [6] [7].
An emerging approach involves strategic integration of both methodologies, using CRISPR for rapid screening and culture for subsequent phenotypic characterization. This hybrid model leverages the sensitivity and speed of nucleic acid detection while maintaining the comprehensive information provided by culture-based antimicrobial susceptibility testing [4] [7].
Experimental Protocols for Biofilm Diagnostics
Standardized TCP Method for Biofilm Detection
The Tissue Culture Plate (TCP) method represents the gold standard for in vitro assessment of biofilm formation and is widely used in both research and clinical settings for evaluating biofilm-producing capacity of bacterial isolates [6] [7].
Materials and Reagents:
- Trypticase Soy Broth (TSB) supplemented with 1% glucose
- Sterile 96-well flat-bottom polystyrene tissue culture plates
- Phosphate Buffered Saline (PBS), pH 7.3
- Crystal violet stain (0.1% w/v)
- Sodium acetate (2% w/v)
- Microplate ELISA reader capable of measuring optical density at 570 nm
Procedure:
- Prepare bacterial suspensions by inoculating a loopful of freshly cultured bacteria into 10 mL of TSB with 1% glucose.
- Incubate the suspension for 18-24 hours at 37°C under appropriate atmospheric conditions.
- Dilute the cultured suspension 1:100 in fresh TSB with 1% glucose.
- Dispense 180 µL of sterile TSB into each well of the tissue culture plate.
- Add 20 µL of the diluted bacterial suspension to each test well, creating a 1:10 final dilution.
- Include appropriate controls: sterile broth (negative control) and known biofilm-forming and non-forming strains (positive and negative controls).
- Seal the plate with Parafilm to prevent evaporation and incubate at 37°C for 24 hours.
- After incubation, gently shake the plate and carefully remove the contents by inversion.
- Wash each well three to four times with sterile distilled water (200 µL per wash) to remove non-adherent cells.
- Air-dry the inverted plate completely.
- Add 200 µL of 2% sodium acetate to each well and incubate for 30 minutes for fixation.
- Wash the fixed biofilm three to four times with sterile distilled water.
- Stain with 200 µL of 0.1% crystal violet for 15 minutes.
- Wash extensively with distilled water until the negative control wells show no residual stain.
- Air-dry the plate completely.
- Quantify biofilm formation by measuring optical density at 570 nm using a microplate reader.
Interpretation: Biofilm formation is classified based on the optical density (OD) measurements:
- Non-biofilm producer: OD ≤ ODC
- Weak biofilm producer: ODC < OD ≤ 2×ODC
- Moderate biofilm producer: 2×ODC < OD ≤ 4×ODC
- Strong biofilm producer: 4×ODC < OD
Where ODC represents the average OD of the negative control wells [6] [7].
CRISPR-Cas12a Diagnostic Protocol for Biofilm Pathogens
This protocol adapts the DETECTR system for detection of specific pathogens within biofilm samples, leveraging the collateral cleavage activity of Cas12a for highly sensitive detection [4].
Materials and Reagents:
- Cas12a enzyme (purified)
- Custom crRNA targeting specific pathogen DNA sequence
- Single-stranded DNA (ssDNA) fluorescent reporter (e.g., FAM-TTATT-BHQ1)
- Isothermal amplification reagents (RPA or LAMP)
- Nucleic acid extraction kit
- Lateral flow strips or fluorometer for detection
- Appropriate positive and negative control templates
Procedure:
- Sample Processing:
- Collect biofilm sample (catheter tip, tissue biopsy, or microbial colony).
- For adherent biofilms, sonicate or enzymatically treat (e.g., with DNase I or dispersin B) to disaggregate matrix and release cells.
- Extract nucleic acids using standardized protocols, ensuring adequate purification to remove PCR inhibitors.
Target Amplification:
- Prepare recombinase polymerase amplification (RPA) or loop-mediated isothermal amplification (LAMP) reaction mix according to manufacturer's instructions.
- Add extracted nucleic acid template to amplification reaction.
- Incubate at optimal temperature (typically 37-42°C) for 15-30 minutes for amplification.
CRISPR Detection:
- Prepare Cas12a detection mixture containing:
- 50 nM Cas12a enzyme
- 50 nM crRNA (designed against target pathogen gene)
- 100 nM ssDNA fluorescent reporter
- Appropriate reaction buffer
- Combine 5 µL of amplified product with 15 µL of Cas12a detection mixture.
- Incubate at 37°C for 10-30 minutes to allow target recognition and collateral cleavage.
- Prepare Cas12a detection mixture containing:
Signal Detection:
- Fluorometric Method: Measure fluorescence intensity using a plate reader or portable fluorometer.
- Lateral Flow Readout: Apply reaction mixture to lateral flow strip containing immobilized reporters for visual detection.
Interpretation:
- Positive result: Significant increase in fluorescence or visible test line on lateral flow strip compared to negative controls.
- Negative result: No fluorescence increase or absent test line.
- The assay should include appropriate controls: no-template control, amplification control, and known positive control [4].
Visualization of Diagnostic Workflows
The diagnostic pathways for biofilm pathogens differ significantly between culture and CRISPR-based approaches, each with distinct advantages and limitations as illustrated below.
Diagnostic Workflow Comparison
The visualization highlights the significant time differential between these approaches, with CRISPR systems compressing the diagnostic timeline from several days to under two hours. This acceleration is particularly valuable for biofilm-associated infections where timely, targeted therapy is essential for successful outcomes.
The Scientist's Toolkit: Essential Research Reagents
Table 5: Key Reagents for Biofilm Diagnostic Research
| Reagent/Category | Specific Examples | Research Application | Considerations for Biofilm Studies |
|---|---|---|---|
| Culture Media | Trypticase Soy Broth (TSB) with 1% glucose, Congo Red Agar | Biofilm formation assessment, phenotypic characterization | Glucose enhances EPS production; CRA differentiates producers |
| Molecular Enzymes | Cas12a/Cas13 proteins, recombinase polymerase amplification (RPA) enzymes | CRISPR diagnostics, isothermal amplification | Enable rapid detection without culturalbility requirement |
| Nucleic Acid Reporters | FAM-TTATT-BHQ1 ssDNA reporters, biotin-labeled RNA probes | Signal generation in CRISPR assays | Collateral cleavage detection for ultrasensitive readout |
| Matrix Disruption Agents | DNase I, proteinase K, dispersin B | EPS breakdown for cell release or component analysis | Enhance nucleic acid extraction efficiency from biofilms |
| Staining Reagents | Crystal violet (0.1%), SYTO fluorescent dyes | Biofilm visualization and quantification | CV stains matrix components; fluorescent dyes label cells |
| Reference Strains | S. epidermidis ATCC 35984 (biofilm-positive), S. epidermidis ATCC 12228 (biofilm-negative) | Method validation and quality control | Essential for standardized assay performance assessment |
The architectural complexity of biofilms, characterized by the protective EPS matrix and heterogeneous bacterial populations containing persister cells, presents significant challenges for conventional diagnostic approaches. Culture methods, while providing valuable phenotypic information including antimicrobial susceptibility profiles, face limitations in detecting dormant persisters and require extended timeframes that delay appropriate therapy. CRISPR-based diagnostics offer a promising alternative with rapid turnaround times, exceptional sensitivity, and the ability to detect pathogens independent of viability or growth state.
The optimal diagnostic strategy depends on the specific clinical or research context. For routine susceptibility testing and when dealing with unknown pathogens, culture methods remain relevant. However, for rapid screening, detection of fastidious or slow-growing organisms, and identification of specific genetic markers, CRISPR systems provide distinct advantages. Future directions point toward integrated approaches that leverage the strengths of both methodologies, potentially combining CRISPR-based rapid identification with culture-derived phenotypic information to optimize the management of biofilm-associated infections.
As CRISPR technologies continue to evolve with improvements in multiplexing, quantification, and point-of-care applicability, they are poised to transform our approach to diagnosing and monitoring biofilm-associated infections, ultimately contributing to more effective antimicrobial stewardship and improved patient outcomes.
In the field of clinical microbiology, culture-based methods have long been the cornerstone for pathogen identification. However, the evolving landscape of infectious diseases, particularly the prevalence of polymicrobial and biofilm-associated infections, has exposed critical limitations in these traditional techniques [8]. The inherent constraints of culture-based methods—notably their prolonged turnaround times and inadequate sensitivity for complex microbial communities—can directly impact patient outcomes by delaying the implementation of targeted therapeutic interventions [9] [10]. This review objectively examines these limitations through quantitative data and explores how emerging CRISPR-based diagnostic platforms address these challenges within biofilm pathogen detection research.
Fundamental Limitations of Culture-Based Methods
Extended Turnaround Times and Technical Demands
Traditional culture-based identification requires significant time to yield results, creating critical delays in clinical decision-making. Microbial culture, considered the laboratory detection "gold standard," typically takes 2–10 days to complete due to the necessity for microbial growth and subsequent biochemical analysis [11] [12]. This extended timeframe contrasts sharply with the urgent need for rapid diagnosis in clinical settings, particularly for immunocompromised patients where treatment delays can be life-threatening [10].
The technical demands of culture methods further limit their utility. These techniques require specialized equipment, controlled laboratory facilities, and skilled personnel, making them less accessible and cost-effective, especially in resource-limited settings [4]. The reliance on viable, fast-growing organisms means that slow-growing, fastidious, or unculturable pathogens are frequently overlooked, resulting in incomplete diagnostic information [8].
Critical Sensitivity Gaps in Polymicrobial and Biofilm-Associated Infections
The diagnostic challenges of culture-based methods become particularly pronounced in the context of polymicrobial and biofilm-associated infections, where their sensitivity limitations are markedly evident.
Table 1: Documented Culture Negative Rates in Clinical Infections
| Infection Type | Reported Culture-Negative Rate | Primary Contributing Factors |
|---|---|---|
| Periprosthetic Joint Infections (PJIs) | Up to 20% [9] | Heterogeneous bacterial distribution in biofilms [9] [13] |
| Osteomyelitis (OM) | "Alarming" rates using conventional culture [9] | Bacterial aggregation and biofilm formation [9] |
| Polymicrobial Infections (Overall) | 30-40% of co-pathogens missed [8] | Focus on dominant, fast-growing organisms [8] |
Polymicrobial infections (PMIs) represent an estimated 20-50% of severe clinical infection cases, with biofilm-associated and device-related infections reaching 60-80% in hospitalized patients [8]. Despite this prevalence, conventional culture-based methods demonstrate significant shortcomings as they predominantly detect fast-growing, dominant microbes while frequently missing slow-growing, anaerobic, or difficult-to-culture organisms [8].
The underlying issue often stems from the biological nature of biofilms themselves. Bacteria within biofilms exist in aggregates of various sizes (5 to 1,000 µm) distributed heterogeneously throughout tissues rather than as uniform suspensions [9]. This aggregated distribution creates substantial sampling challenges, as the probability of capturing bacterial colonies in a tissue biopsy decreases dramatically as aggregate size increases [9]. Mathematical modeling reveals that beyond a critical aggregation parameter, increasing the number of tissue specimens provides limited benefit for detection, leading to culture-negative diagnoses despite the presence of active infection [9].
Table 2: Impact of Bacterial Aggregation on Detection Probability
| Aggregation Level | Recommended Specimen Number | Expected Detection Outcome |
|---|---|---|
| Below Critical Aggregation | 5 tissue specimens | High probability of detection |
| Above Critical Aggregation | Increasing specimen count provides limited benefit | High false-negative rate persists |
Additionally, the phenotypic state of biofilm-associated bacteria further complicates detection. Environmental stressors, including antibiotic exposure or oxygen availability, can induce bacterial growth restriction or dormancy, leading to viable but non-culturable (VBNC) bacteria that evade traditional culture methods despite their continued pathogenicity [9]. This phenomenon has been confirmed through direct visualization techniques that identify bacteria in samples where culture methods showed no growth [9].
CRISPR-Based Diagnostics: Emerging Alternatives
Fundamental Mechanisms and Workflows
CRISPR-based diagnostics represent a paradigm shift from culture-based methods by directly targeting pathogen nucleic acids rather than relying on microbial growth. These systems utilize Cas proteins (e.g., Cas9, Cas12, Cas13) that, upon recognition of specific pathogen DNA or RNA sequences through guide RNA (crRNA), exhibit both specific cleavage of the target and non-specific trans-cleavage activity against surrounding reporter molecules [4] [11] [12]. This collateral cleavage of fluorescent or colorimetric reporter probes enables highly sensitive signal amplification and visual detection of pathogen nucleic acids [4].
The CRISPR diagnostic workflow eliminates the dependency on bacterial viability and growth kinetics, instead leveraging programmable nucleic acid recognition to achieve pathogen identification. Two primary approaches have emerged:
- Amplification-based CRISPR: Combines isothermal amplification techniques (RPA, LAMP) with CRISPR detection to enhance sensitivity [11] [12]
- Amplification-free CRISPR: Utilizes direct detection strategies (cascade CRISPR, sensor technologies) to reduce operational complexity and contamination risk [11] [12]
Performance Comparison: Quantitative Experimental Data
Substantial experimental evidence demonstrates the superior performance characteristics of CRISPR-based diagnostics compared to traditional methods, particularly for challenging infection types.
Table 3: Experimental Comparison of Detection Technologies for Pathogen Identification
| Methodology | Sensitivity | Time to Result | Key Advantages | Representative Experimental Findings |
|---|---|---|---|---|
| Traditional Culture | Variable (misses 30-40% of co-pathogens in PMIs) [8] | 2-10 days [11] [12] | Determines microbial viability, antibiotic susceptibility testing | 40% detection rate in myelosuppressed patients with respiratory infections [10] |
| CRISPR-based Detection | attomolar (aM) level [4]; single-copy detection [11] | <2 hours to ~30 minutes [8] [11] | Rapid, high sensitivity/specificity, equipment-free potential | 100% pathogen detection rate in respiratory infections; 94.74% sensitivity [10] |
| Targeted NGS (tNGS) | Broad-spectrum pathogen identification | ~24 hours [10] | Comprehensive pathogen profiling | Directly influenced treatment changes in 75% of cases [10] |
Recent clinical studies provide compelling evidence for the superior diagnostic performance of molecular methods. In a prospective study comparing diagnostic efficacy for respiratory infections in myelosuppressed hematology patients, targeted next-generation sequencing (tNGS) demonstrated a 100% pathogen detection rate compared to just 40% with traditional culture-based methods (p < 0.001) [10]. The same study reported that tNGS identified a broader spectrum of pathogens, including bacteria, viruses, and fungi that were missed by traditional methods, with results directly influencing treatment adjustments in 75% of cases [10].
Beyond sequencing technologies, CRISPR platforms have demonstrated remarkable sensitivity in controlled experiments. Bead-based CRISPR approaches have achieved sensitivity reaching as low as 2.5 copies per µl of input RNA through a novel split-luciferase reporter system that provides up to 20× higher sensitivity compared to standard fluorescence-based reporter designs [14]. Similarly, amplification-free CRISPR-Cas13a platforms have successfully detected SARS-CoV-2 down to 470 aM within 30 minutes, demonstrating potential for rapid, highly sensitive pathogen identification without the need for target amplification [11] [12].
Experimental Protocols for Key Comparisons
Protocol: Evaluating Sensitivity in Polymicrobial Detection
Objective: Compare the detection sensitivity of culture-based methods versus CRISPR-based diagnostics for identifying multiple bacterial species in synthetic polymicrobial samples.
Sample Preparation:
- Create standardized samples containing known ratios of common polymicrobial infection pathogens (e.g., Staphylococcus aureus, Pseudomonas aeruginosa, Escherichia coli, and Bacteroides fragilis) in both planktonic and biofilm states [8]
- For biofilm samples, cultivate communities on relevant substrates (e.g., medical device materials) for 48-72 hours to establish mature biofilms [13]
- Serially dilute samples across a concentration range from 10^8 to 10^1 CFU/mL
Culture-Based Method:
- Plate samples on appropriate culture media (general and selective)
- Incubate under aerobic and anaerobic conditions at 37°C for up to 14 days [9]
- Record time-to-detection for each species and calculate minimum detection limits
CRISPR-Based Detection:
- Extract nucleic acids using commercial kits (e.g., MagPure Pathogen DNA/RNA Kit) [10]
- Employ multiplexed crRNAs targeting species-specific genetic markers
- Use Cas12 or Cas13 systems with fluorescent reporters
- Perform detection with and without pre-amplification (RPA/LAMP)
- Measure detection limits and time-to-result
Analysis: Compare detection thresholds, time requirements, and species identification completeness across methods [8] [10].
Protocol: Assessing Detection of Heterogeneously Distributed Biofilms
Objective: Evaluate the ability of culture versus CRISPR methods to detect bacteria distributed in aggregates simulating in vivo biofilm infection architecture.
Sample Preparation:
- Generate bacterial aggregates of controlled sizes (5-500 µm) using centrifugation-based methods [9]
- Incorporate aggregates into tissue homogenates at defined bacterial loads (10^2-10^6 CFU/g)
- Prepare samples with varying aggregation parameters (c) relative to the detection limit (ηℓ × mB)
Sampling and Detection:
- Collect multiple tissue specimens (e.g., 0.1 g punch biopsies) from homogenized samples
- Process specimens by:
- Calculate probability of detection based on positive samples and compare to mathematical models [9]
Mathematical Modeling: Apply probability formula: P(at least one positive biopsy) ≈ 1 - Q(1 + (ηℓ·mB)/c, (η·mB)/c)^M Where: η = bacterial load, ηℓ = detection limit, mB = biopsy size, c = aggregation parameter, M = number of biopsies [9]
Essential Research Reagent Solutions
The implementation of comparative studies between culture-based and CRISPR-based detection methods requires specific research reagents and platforms.
Table 4: Essential Research Reagents for Diagnostic Comparison Studies
| Reagent Category | Specific Examples | Research Application | Key Considerations |
|---|---|---|---|
| CRISPR Enzymes | Cas12a, Cas13a, Cas14 [11] [12] | Nucleic acid detection core | Varying PAM requirements, target preferences (DNA/RNA) [12] |
| Nucleic Acid Amplification | RPA, LAMP kits [11] [12] | Sensitivity enhancement | Compatibility with CRISPR systems; one-pot vs. two-step protocols [11] |
| Reporter Systems | Fluorescent (FAM/ROX with BHQ), lateral flow strips [11] [12] | Detection signal generation | Equipment requirements; visual vs. instrumental readout [11] |
| crRNA Design | Custom pathogen-specific crRNAs [4] | Target recognition | Specificity validation; multiplexing capacity [4] |
| Bead-Based Platforms | Split-luciferase reporters, color-coded crRNA beads [14] | Enhanced sensitivity and multiplexing | Signal-to-noise improvement; parallel target detection [14] |
The limitations of culture-based methods—particularly their extended turnaround times and inadequate sensitivity for polymicrobial and biofilm-associated infections—represent significant challenges in clinical diagnostics. Quantitative evidence demonstrates that culture methods miss 30-40% of co-pathogens in polymicrobial infections and exhibit false-negative rates up to 20% in biofilm-associated orthopedic infections [9] [8]. CRISPR-based diagnostic platforms address these limitations through their rapid detection capabilities (<2 hours), exceptional sensitivity (attomolar level), and ability to identify complex microbial communities without dependence on bacterial viability [4] [11]. While culture methods remain valuable for antibiotic susceptibility testing, the integration of CRISPR technologies into diagnostic pipelines offers researchers and clinicians powerful tools for comprehensive pathogen detection, particularly in the context of biofilm research where traditional methods consistently underperform. The continued refinement of CRISPR-based platforms promises to further bridge the diagnostic gaps in managing complex infectious disease presentations.
Microbial biofilms are structured communities of bacteria, fungi, or other microorganisms embedded within a self-produced extracellular polymeric substance (EPS) matrix that adheres to biological or inert surfaces [13] [15]. This complex architecture creates formidable barriers in both healthcare and food industry settings, enabling pathogens to persist despite aggressive cleaning and antimicrobial interventions. In healthcare-associated infections (HAIs), biofilms form on medical devices like catheters, implants, and ventilators, causing persistent infections that exhibit dramatically increased antibiotic tolerance—up to 1,000-fold greater than their planktonic (free-floating) counterparts [13]. Similarly, in the food industry, biofilms establish on processing equipment, conveyor belts, and food contact surfaces, serving as persistent contamination sources that lead to spoilage and foodborne illness outbreaks [16] [17].
The World Health Organization estimates that foodborne and waterborne bacterial diseases cause 2.2 million deaths annually worldwide, with biofilm-associated contamination representing a significant contributing factor [18]. The economic burden is equally staggering, with biofilm-related losses in the global agrifood sector recently estimated at approximately $324 billion annually, while foodborne illnesses in the U.S. alone cost about $17.6 billion per year [16]. This persistent public health burden has stimulated the development of novel diagnostic and control strategies, with CRISPR-based technologies emerging as particularly promising alternatives to conventional culture-based methods for detecting and characterizing biofilm-forming pathogens.
Conventional Culture Methods: The Established Paradigm
Methodological Principles and Workflows
Traditional culture-based methods remain the cornerstone of microbial detection in most clinical and industrial settings, relying on the growth and phenotypic characterization of microorganisms. The standard workflow involves sample collection from suspected biofilm contamination sites, inoculation onto selective and non-selective culture media, incubation under appropriate conditions (typically 24-48 hours for initial growth), followed by morphological examination and biochemical testing for identification [11] [17]. For antimicrobial susceptibility testing (AST) of biofilm-associated organisms, the Calgary Biofilm Device or similar systems are employed to grow standardized biofilms before exposing them to antimicrobial agents, with results typically available within 72-96 hours from specimen collection [19].
Performance Limitations in Biofilm Detection
While culture methods provide the historical "gold standard" for pathogen identification and enable antimicrobial susceptibility profiling, they face significant limitations when applied to biofilm-associated infections and contamination. The heterogeneous nature of biofilms, with subpopulations of dormant "persister" cells and variable metabolic states, leads to potential false negatives as some viable organisms may not grow under standard laboratory conditions [15]. The extensive time-to-result (often 3-5 days for complete identification and AST) represents another critical limitation, particularly in clinical settings where delayed appropriate antibiotic therapy correlates with increased mortality in sepsis [19]. Additionally, culture-based approaches cannot readily distinguish between planktonic and biofilm-growing cells in mixed samples, potentially underestimating the true burden of biofilm-associated pathogens [15].
CRISPR-Based Diagnostics: An Emerging Technological Paradigm
Fundamental Mechanisms and Molecular Principles
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) proteins represent a revolutionary tool for nucleic acid detection, derived from adaptive immune systems in bacteria and archaea [4] [11]. Unlike culture methods that detect viable organisms through growth, CRISPR-based diagnostics identify pathogen-specific DNA or RNA sequences through programmable molecular recognition. The system utilizes Cas nucleases (such as Cas9, Cas12, Cas13) guided by CRISPR RNA (crRNA) to recognize and cleave specific nucleic acid targets [4]. Upon target recognition, certain Cas enzymes (notably Cas12 and Cas13) exhibit "collateral cleavage" or trans-activity, non-specifically degrading reporter molecules that generate detectable fluorescent or colorimetric signals [11]. This combination of specific recognition and nonspecific signal amplification enables sensitive detection of pathogen signatures without requiring microbial viability or growth.
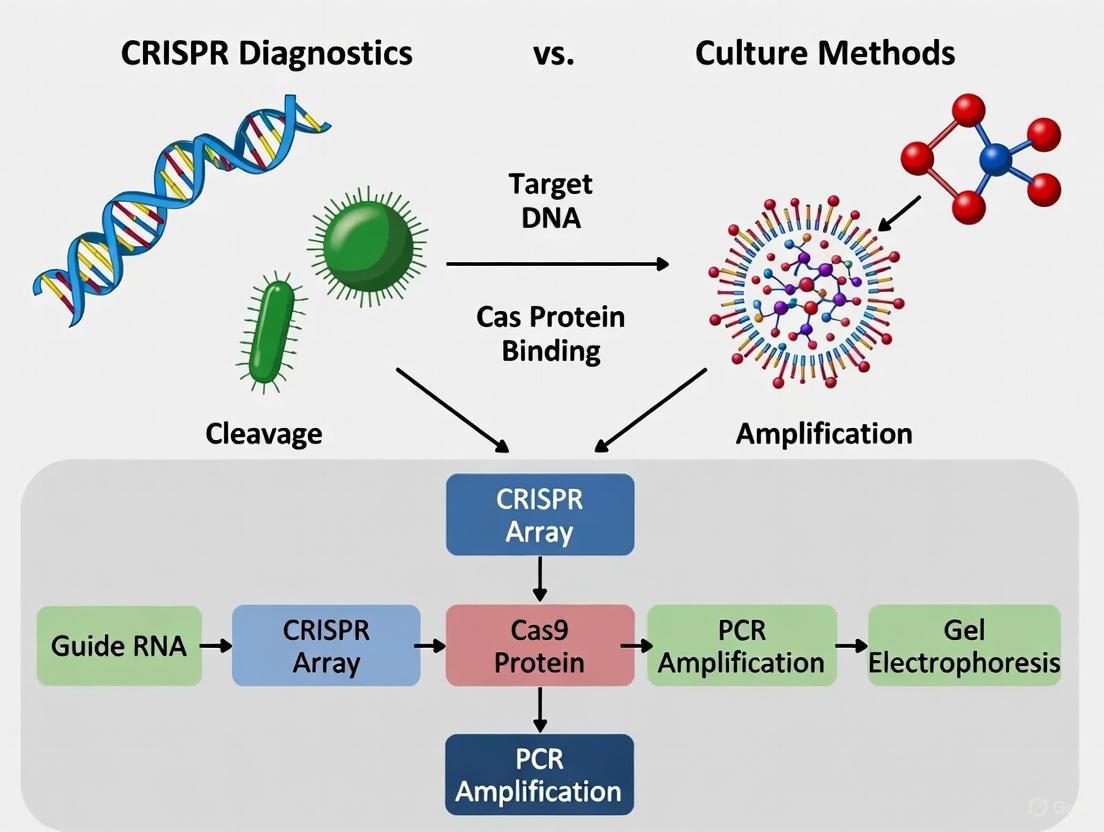
Figure 1: CRISPR-Cas Diagnostic Mechanism. The workflow illustrates how Cas enzymes complexed with guide RNAs recognize specific pathogen DNA/RNA sequences, activating collateral cleavage that amplifies detectable signals.
CRISPR Platform Variants and Applications
Several CRISPR systems have been adapted for diagnostic applications, each with distinct targeting preferences and operational characteristics. Cas9-based systems provide precise DNA targeting but typically lack trans-cleavage activity, limiting their signal amplification capacity. In contrast, Cas12a targets DNA and exhibits robust trans-cleavage of single-stranded DNA reporters, enabling platforms like DNA Endonuclease Targeted CRISPR Trans Reporter (DETECTR) [4]. Meanwhile, Cas13 targets RNA sequences and cleaves single-stranded RNA reporters, forming the basis for Specific High-Sensitivity Enzymatic Reporter Unlocking (SHERLOCK) [11]. These platforms can be further enhanced through integration with pre-amplification techniques like recombinase polymerase amplification (RPA) or loop-mediated isothermal amplification (LAMP), enabling detection sensitivities approaching attomolar (aM) levels—capable of identifying single copies of target genes in some applications [11]. For biofilm-specific applications, CRISPR systems can be programmed to target conserved regions of bacterial genomes, virulence genes, or antibiotic resistance determinants associated with biofilm-forming pathogens.
Comparative Performance Analysis: CRISPR Versus Culture Methods
Analytical Sensitivity and Specificity
Recent advances in CRISPR-based diagnostics have demonstrated exceptional sensitivity and specificity for pathogen detection, often surpassing traditional culture methods, particularly for slow-growing or fastidious organisms. As shown in Table 1, CRISPR platforms consistently achieve limits of detection at the attomolar level for purified nucleic acids, translating to single-copy sensitivity in optimized systems [11]. The intrinsic sequence specificity of crRNA-guided recognition enables single-nucleotide discrimination, allowing differentiation between closely related bacterial strains or specific detection of antibiotic resistance mutations [4]. This precision facilitates the identification of specific virulence genes (such as biofilm-associated genes) rather than relying on phenotypic characteristics alone.
Table 1: Analytical Performance Comparison of Diagnostic Methods for Biofilm-Associated Pathogens
| Parameter | Conventional Culture Methods | CRISPR-Based Diagnostics | References |
|---|---|---|---|
| Limit of Detection | 10¹-10³ CFU/mL (varies by organism) | ~1 copy/μL (aM levels) with pre-amplification | [19] [11] |
| Time-to-Result | 2-5 days (identification + AST) | 15 minutes to 4 hours | [19] [11] |
| Specificity | Morphological and biochemical differentiation | Single-nucleotide discrimination possible | [4] [11] |
| Viability Requirement | Requires viable, culturable organisms | Detects DNA/RNA regardless of viability | [15] [11] |
| Throughput Potential | Low to moderate | High (potential for multiplexing) | [4] [16] |
| Point-of-Care Suitability | Limited (requires specialized equipment and training) | High (lyophilized reagents, portable readers) | [4] [11] |
Time-to-Result and Clinical Impact
The dramatically reduced time-to-result represents perhaps the most significant advantage of CRISPR-based diagnostics over culture methods. While conventional culture requires 24-72 hours for pathogen identification and an additional 24 hours for antimicrobial susceptibility testing, CRISPR platforms can deliver specific identification in 15 minutes to 4 hours, including sample preparation [19] [11]. This accelerated timeline has profound implications for clinical management of biofilm-associated infections, as delayed appropriate antimicrobial therapy directly correlates with increased mortality in septic patients. One study analyzing rapid AST technologies reported that each hour of delay in effective antibiotic administration for bloodstream infections was associated with a 7.6% increase in mortality [19]. Similar benefits apply to food industry settings, where rapid detection enables timely intervention before contaminated products advance through the supply chain, potentially preventing costly recalls and outbreak investigations.
Practical Implementation Considerations
Despite its promising analytical performance, CRISPR diagnostics face distinct implementation challenges. The requirement for target nucleic acid extraction and potential inhibition by complex sample matrices (like sputum, food homogenates, or biofilm debris) can affect assay robustness [4]. Additionally, the technology currently lacks the comprehensive databases that support matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry or genomic methods for unidentified organisms [4]. Culture methods retain the advantage of being hypothesis-free, enabling detection of unexpected pathogens without prior genetic knowledge, and providing viable isolates for further characterization, outbreak investigation, or epidemiological tracking [19]. Furthermore, regulatory approval and standardization of CRISPR-based diagnostics for clinical use remain in development, while culture methods represent established, validated approaches with well-defined quality control procedures.
Table 2: Functional Characteristics in Applied Settings
| Application Context | Culture Methods | CRISPR-Based Diagnostics | References |
|---|---|---|---|
| Clinical Diagnostics | Gold standard; provides live isolates for further testing | Rapid guiding of targeted therapy; no isolate obtained | [19] [11] |
| Food Industry Monitoring | Detects viable pathogens; established regulatory acceptance | Rapid screening; identifies specific virulence or resistance genes | [16] [17] |
| Biofilm Research | Determines viable counts; antibiotic susceptibility profiling | Targets specific biofilm-associated genes; spatial mapping potential | [16] [15] |
| Antimicrobial Resistance Detection | Phenotypic confirmation of resistance expression | Direct detection of resistance genes (e.g., mecA, bla, ndm-1) | [4] [13] |
| Resource-Limited Settings | Requires sustained infrastructure, stable power, trained personnel | Point-of-care potential; lyophilized reagents; minimal equipment | [4] [19] |
Experimental Approaches and Research Applications
Standardized Methodologies for Comparative Studies
Robust comparison of diagnostic platforms requires standardized experimental protocols. For biofilm studies, researchers typically establish model systems using common biofilm-forming pathogens like Pseudomonas aeruginosa, Staphylococcus aureus, or Escherichia coli on relevant surfaces (catheter segments, stainless steel coupons, or microtiter plates) [16] [13]. Biofilm maturity is quantified using crystal violet staining for total biomass, confocal microscopy for structural analysis, and viable counts for traditional quantification. For CRISPR detection, nucleic acids are extracted using commercial kits optimized for complex matrices, with incorporation of sample processing controls to monitor inhibition [11]. Validation against reference standards and inter-laboratory reproducibility testing are essential for both established and emerging technologies.
Integrated Approaches for Biofilm Research
The most comprehensive understanding of biofilm-associated pathogens emerges from integrated approaches that leverage the complementary strengths of both culture-based and molecular methods. Culture provides confirmation of viability and phenotypic characteristics, while CRISPR-based methods offer rapid detection and specific genotyping. This integration is particularly valuable for investigating biofilm resistance mechanisms, where conventional MIC testing combined with CRISPR-based detection of resistance genes provides a complete picture of both genotypic potential and phenotypic expression [13] [15]. Additionally, culture-based isolation enables subsequent studies of biofilm formation kinetics, dispersal mechanisms, and transcriptomic profiling under controlled conditions, generating biological insights that complement diagnostic applications.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Research Reagents for Biofilm and CRISPR Diagnostic Studies
| Reagent/Material | Function/Application | Examples/Specifications | References |
|---|---|---|---|
| Cas Enzymes | CRISPR nucleic acid recognition and cleavage | Cas12a, Cas13a (purified recombinant proteins) | [4] [11] |
| crRNA Guides | Target-specific sequence recognition | Synthetic RNA designed for pathogen-specific genes | [4] [11] |
| Reporter Molecules | Signal generation upon collateral cleavage | Fluorescent (FAM-quencher) or lateral flow reporters | [11] [20] |
| Nucleic Acid Amplification Reagents | Pre-amplification of target sequences | RPA, LAMP, or PCR kits with appropriate primers | [11] |
| Biofilm Matrix Disruption Reagents | Nucleic acid extraction from biofilms | DNase-RNase-free enzymes for EPS degradation | [16] [15] |
| Microfluidic Platforms | Automated sample processing and detection | Integrated "sample-to-answer" cartridges | [4] [19] |
Future Directions and Translational Applications
The evolving landscape of biofilm diagnostics points toward increased integration of CRISPR technologies with complementary advanced methodologies. Combination with microfluidic systems enables automated "sample-to-answer" platforms that minimize hands-on time and reduce contamination risk [4] [19]. Integration with artificial intelligence and machine learning algorithms facilitates pattern recognition for outbreak detection and resistance prediction [16]. Emerging amplification-free CRISPR strategies further simplify workflows and reduce costs, potentially improving accessibility in resource-limited settings [11]. For biofilm-specific applications, researchers are developing CRISPR-based approaches that not only detect pathogens but also modulate biofilm formation through targeted interference with quorum-sensing or adhesion genes [16] [13].
Figure 2: Emerging Applications of CRISPR Technology. Future developments focus on platform integration, AI-enhanced analysis, and therapeutic applications that extend beyond diagnostics.
The translation of CRISPR diagnostics from research laboratories to clinical and industrial settings will require addressing several remaining challenges. Efficient delivery of CRISPR components through complex biofilm matrices, standardization and quality control for regulatory approval, and demonstrated cost-effectiveness in real-world applications represent active areas of investigation [4] [16]. Furthermore, ethical considerations regarding biosecurity and appropriate use must be addressed through established frameworks and guidelines. Despite these challenges, the exceptional sensitivity, rapid turnaround, and programmability of CRISPR-based systems position them as transformative tools for addressing the persistent public health burden of biofilm-associated infections across healthcare and food industry sectors.
The comparison between established culture methods and emerging CRISPR-based diagnostics reveals a compelling evolution in approaches to biofilm detection and characterization. Conventional culture techniques provide the irreplaceable benefit of yielding viable isolates for phenotypic testing and remain the gold standard for antimicrobial susceptibility profiling. However, their extended time-to-result and dependence on microbial cultivability present significant limitations for rapid intervention. CRISPR technologies offer transformative potential through exceptional sensitivity, dramatically reduced detection timelines, and precise sequence-specific identification of pathogens and resistance markers. The optimal approach for biofilm research and diagnostics likely involves strategic integration of both methodologies, leveraging their complementary strengths to advance both fundamental understanding and applied management of biofilm-associated public health challenges. As CRISPR platforms continue to mature through ongoing research and development, they promise to significantly enhance our capacity to detect, monitor, and ultimately mitigate the substantial disease and economic burdens imposed by biofilms across healthcare and food industry environments.
In resource-limited settings, the absence of advanced laboratory infrastructure, reliable electricity, and trained personnel creates a critical need for diagnostic tools that can operate independently of these constraints. The World Health Organization (WHO) has established the ASSURED criteria (Affordable, Sensitive, Specific, User-friendly, Rapid and robust, Equipment-free, and Deliverable to end-users) as a benchmark for ideal point-of-care (POC) tests [21] [22]. These criteria ensure that diagnostics are not only effective but also practical and accessible in the most challenging environments. Recently, this framework has been expanded to REASSURED to include Real-time connectivity, Ease of specimen collection, and Sustainability, reflecting evolving technological and operational priorities [22].
This review evaluates two divergent technological paths for detecting biofilm-associated pathogens—a major cause of persistent infections—against this ideal benchmark: traditional culture-based methods and emerging CRISPR-based diagnostics. Biofilms, which are structured communities of microorganisms encased in a protective matrix, account for an estimated 60–80% of all microbial infections in humans and are notoriously difficult to diagnose and treat due to their inherent resistance to antibiotics and host immune responses [23] [24] [8]. Their detection is critical for managing conditions such as catheter-associated urinary tract infections (CAUTIs), diabetic foot infections, and infections related to other indwelling medical devices [23] [8]. We objectively compare the performance of these two diagnostic approaches, provide supporting experimental data, and detail the essential methodologies and reagents that constitute the modern scientist's toolkit for biofilm pathogen detection.
Performance Comparison: Culture-Based Methods vs. CRISPR Diagnostics
The following tables provide a quantitative comparison of traditional culture-based methods and novel CRISPR-based diagnostics for detecting biofilm-forming pathogens, evaluated against the WHO's ASSURED criteria and key performance metrics.
Table 1: Evaluation Against WHO ASSURED Criteria
| WHO ASSURED Criteria | Culture-Based Methods | CRISPR-Based Diagnostics |
|---|---|---|
| Affordable | Low cost per test; high overall system cost due to lab infrastructure and trained personnel [21] | Potentially low-cost; dependent on lyophilized reagents and simplified readouts [4] [25] |
| Sensitive | Variable; can miss low-abundance or slow-growing pathogens in polymicrobial biofilms [8] | High; attomolar (aM) sensitivity for nucleic acids, especially when combined with pre-amplification [4] [11] |
| Specific | High for identified colonies; misses non-culturable organisms [23] | Extremely high; single-base pair specificity guided by crRNA [4] [25] |
| User-friendly | Low; requires multiple complex steps and significant technical expertise [23] | Moderate to High; minimal steps, but requires careful reagent handling; results often visible on lateral flow strips [4] [22] |
| Rapid & Robust | Slow (24-72 hours); not robust for urgent care [23] [11] | Very rapid (30 min - 2 hours); robust in controlled conditions, though enzymes can be fragile (e.g., 63% performance drop in high humidity) [4] [8] |
| Equipment-free | No; requires incubators, biosafety cabinets, etc. [21] | Yes for some formats; can be equipment-free with lateral flow readouts [4] [25] |
| Deliverable | Challenging; cold chain for media, bulky equipment [21] | Promising; reagents can be lyophilized for ambient temperature storage and transport [4] |
Table 2: Technical Performance and Operational Characteristics
| Characteristic | Culture-Based Methods | CRISPR-Based Diagnostics |
|---|---|---|
| Time-to-Result | 2-10 days [11] | 30 minutes - 2 hours [4] [8] |
| Limit of Detection | ~104 CFU/mL for some pathogens in biofilms [23] | As low as 1 copy/µL (with amplification) [11] |
| Multiplexing Capability | Low; requires multiple selective media and processing [8] | High; inherent potential for multiplexing different crRNAs [4] [16] |
| Sample Preparation | Centrifugation, rinsing, enrichment broths [23] | Often requires sample lysis and nucleic acid extraction; some direct detection formats emerging [4] [11] |
| Key Applications | Gold standard for viability; biofilm phenotypic characterization (e.g., TCP, Tube method) [23] | Rapid pathogen ID, resistance gene detection, and potentially precision biofilm disruption [4] [16] |
| Key Limitation | Lengthy turnaround, inability to detect viable but non-culturable bacteria [23] [8] | Susceptibility to inhibitors in complex samples, delivery barriers in biofilm matrix [4] [16] |
Experimental Protocols for Biofilm Pathogen Detection
Protocol for Culture-Based Phenotypic Detection (TCP Method)
The Tissue Culture Plate (TCP) method is considered the reference standard for quantitative biofilm detection [23].
Methodology:
- Inoculation: A loopful of a freshly cultured bacterial isolate is inoculated into 10 mL of trypticase soy broth supplemented with 1% glucose (TSBg) [23].
- Dilution: A 1:100 dilution of the bacterial suspension is prepared. Then, 180 µL of sterile TSBg is dispensed into wells of a sterile 96-well flat-bottom polystyrene tissue culture plate, followed by 20 µL of the bacterial suspension [23].
- Incubation: The plate is sealed with Parafilm and incubated at 37°C for 24 hours to allow biofilm formation [23].
- Washing: After incubation, the plate's contents are discarded to remove planktonic (non-attached) cells. Each well is washed three to four times with sterile distilled water to remove loosely associated bacteria and inverted to dry [23].
- Fixing and Staining: Biofilms are fixed by adding 200 µL of 2% sodium acetate to each well for 30 minutes. After washing again, 200 µL of 0.1% crystal violet solution is added to each well and left for 15 minutes to stain the adherent biofilm [23].
- Quantification: The optical density (OD) of the stained biofilm is measured using a microtiter plate reader at a specific wavelength (e.g., 570-595 nm). The OD values are compared to a control well and a predetermined cutoff to classify isolates as strong, moderate, or weak biofilm producers [23].
Protocol for CRISPR-Cas12a-Based Pathogen Detection (DETECTR-like Assay)
This protocol outlines a CRISPR-based method for detecting specific nucleic acid sequences from pathogens, which can be coupled with a pre-amplification step for maximum sensitivity [4] [11] [25].
Methodology:
- Sample Preparation and Nucleic Acid Extraction: Bacterial samples are collected and lysed to release genomic DNA or RNA. For biofilm samples, mechanical disruption may be necessary. Nucleic acids are then purified using standard commercial kits or rapid lysis buffers [11].
- Pre-Amplification (Optional but common): To enhance sensitivity, the extracted DNA is amplified using an isothermal amplification method like Recombinase Polymerase Amplification (RPA) or Loop-Mediated Isothermal Amplification (LAMP). This step can increase the abundance of the target sequence, enabling detection of very few pathogen copies [4] [11].
- CRISPR-Cas12a Reaction:
- The reaction mixture is prepared containing the Cas12a enzyme, a specific crRNA designed to be complementary to the target pathogen's DNA sequence, and single-stranded DNA (ssDNA) reporter molecules. The reporters are typically labeled with a fluorophore and a quencher (for fluorescence readouts) or with biotin and FAM (for lateral flow readouts) [4] [25].
- The pre-amplified product (or extracted nucleic acid for amplification-free methods) is added to the reaction mix.
- The reaction is incubated at a constant temperature (e.g., 37°C) for 15-60 minutes.
- Signal Detection:
- Fluorescence: If the target DNA is present, the Cas12a-crRNA complex binds to it, activating its "collateral cleavage" activity. This results in the non-specific cleavage of the ssDNA reporters, separating the fluorophore from the quencher and generating a fluorescent signal that can be measured with a portable fluorometer [25].
- Lateral Flow: For the lateral flow format, cleavage of the reporter modifies its mobility on the strip. A visible test line appears, indicating a positive result. This format is entirely equipment-free [4] [11].
Visualizing the CRISPR-Cas12a Diagnostic Mechanism
The following diagram illustrates the molecular mechanism of the CRISPR-Cas12a system (as used in the DETECTR assay), which is fundamental to its function as a diagnostic tool.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Reagents and Materials for Featured Experiments
| Item | Function/Description | Relevance to Experiment |
|---|---|---|
| Trypticase Soy Broth (TSB) with 1% Glucose | Growth medium that promotes biofilm formation by providing nutrients and enhancing polysaccharide production [23]. | Essential for culturing bacteria and promoting biofilm formation in the Tissue Culture Plate (TCP) method [23]. |
| Crystal Violet (0.1%) | A dye that binds to polysaccharides and other negatively charged molecules within the biofilm matrix, allowing for visual and spectrophotometric quantification [23]. | Used for staining adherent biofilms in the TCP method; the intensity of staining correlates with the amount of biofilm [23]. |
| Cas12a Enzyme (e.g., LbCas12a) | A CRISPR-associated nuclease that, upon recognition of a specific target DNA sequence, exhibits non-specific single-stranded DNA (ssDNA) cleavage (trans-cleavage) activity [4] [25]. | The core enzyme in the diagnostic assay; its programmable targeting and collateral activity enable specific and sensitive detection [25]. |
| crRNA (CRISPR RNA) | A short, custom-designed RNA molecule that guides the Cas12a enzyme to a complementary target DNA sequence with high specificity [4] [25]. | Determines the specificity of the assay; designed to uniquely identify the pathogen of interest by targeting a conserved genomic region [11]. |
| ssDNA Fluorescent Reporter | A short single-stranded DNA oligonucleotide labeled with a fluorophore and a quencher; cleavage separates the pair, generating a fluorescent signal [25]. | Serves as the signal-generating molecule in fluorescence-based CRISPR assays; its cleavage indicates a positive detection event [25]. |
| RPA/LAMP Reagents | Isothermal nucleic acid amplification kits that rapidly multiply a specific DNA target at a constant temperature, without the need for a thermal cycler [11]. | Used for pre-amplification of target DNA from samples, dramatically increasing the sensitivity of the CRISPR assay to attomolar levels [4] [11]. |
| Lateral Flow Strips | Paper-based devices that can visually display the result of a biochemical reaction, such as the capture of a cleaved reporter molecule [4] [11]. | Provide an equipment-free readout for CRISPR assays, making them suitable for point-of-care use in resource-limited settings [4]. |
The comparative analysis presented here clearly delineates the divergent profiles of culture-based and CRISPR-based diagnostics. Culture methods, while affordable and providing vital information on viability and phenotype, fall short of the ASSURED ideal in critical areas such as speed, equipment needs, and deliverability [21] [23]. In contrast, CRISPR diagnostics excel in sensitivity, specificity, and rapidity, showing immense potential to meet the demands of POC testing [4] [25]. However, their current vulnerability to environmental conditions and sample inhibitors highlights that the path to a truly "ideal" diagnostic is not yet complete [4].
The future of biofilm pathogen detection in resource-limited settings lies in the continued refinement of molecular tools like CRISPR. Overcoming delivery barriers within the biofilm matrix, developing robust sample preparation methods, and integrating these systems with digital and connectivity solutions (as envisioned in the REASSURED framework) are the next critical steps [22] [16]. By focusing development efforts on these challenges, the scientific community can bridge the gap between revolutionary diagnostic potential and the practical, equitable application required to improve patient outcomes in the world's most challenging healthcare environments.
The CRISPR Diagnostic Toolkit: Mechanisms and Platforms for Precision Biofilm Detection
The CRISPR-Cas system, originally identified as an adaptive immune mechanism in bacteria and archaea, has been repurposed as a revolutionary tool for molecular diagnostics [4] [25]. This technology leverages the precise nucleic acid targeting capabilities of Cas proteins, which are guided by short RNA sequences to identify and cleave specific pathogen DNA or RNA [11]. For biofilm pathogen detection, CRISPR diagnostics offer a promising alternative to traditional culture methods, providing rapid, sensitive, and specific identification of antibiotic-resistant bacteria embedded in complex biofilm matrices [26] [27]. The core principle enabling this diagnostic capability revolves around two key molecular activities: crRNA-guided target recognition and trans-cleavage activity, which form the foundation for detecting even minimal amounts of pathogen nucleic acids within challenging sample types [28] [29].
This guide objectively compares the performance of CRISPR-based diagnostic systems against traditional culture methods, with a specific focus on detecting biofilm-forming pathogens. We present experimental data, detailed methodologies, and analytical frameworks to help researchers select appropriate detection strategies for their specific applications in clinical diagnostics and drug development.
Molecular Mechanisms of CRISPR Diagnostics
crRNA-Guided Target Recognition
The CRISPR-Cas system's exceptional specificity originates from its guide RNA component, known as CRISPR RNA (crRNA) [4]. The crRNA acts as a molecular homing device, carrying a spacer sequence that is complementary to the target nucleic acid (DNA or RNA) of the pathogen [11]. This spacer sequence enables the crRNA to recognize and bind to specific pathogen sequences through Watson-Crick base pairing [4]. In diagnostic applications, artificially designed crRNAs can precisely target conserved regions of pathogen nucleic acids, such as bacterial 16S rRNA genes, virulence factors, or drug-resistant genes, to achieve specific recognition [4]. This targeting mechanism is highly programmable and can be adapted to different pathogens by modifying crRNA sequences [4].
The target recognition process requires the presence of a specific protospacer adjacent motif (PAM) sequence adjacent to the target region for Cas12a systems [28]. The PAM sequence varies depending on the Cas protein used; for Cas12a, it is typically a 5' TTTN motif [28]. Only upon recognition of the PAM sequence by the Cas protein does the Cas protein unwind the target DNA, allowing complementarity-dependent cleavage activity to be activated [28].
Trans-Cleavage Activity
Upon successful recognition and binding to the target nucleic acid, several Cas proteins exhibit a collateral cleavage activity, known as trans-cleavage [25]. This phenomenon is particularly prominent in Cas12a (which targets DNA) and Cas13a (which targets RNA) systems [25] [11]. Once activated by target recognition, these Cas proteins undergo conformational changes that activate their non-specific nuclease activity, enabling them to indiscriminately cleave nearby single-stranded DNA (for Cas12a) or single-stranded RNA (for Cas13a) reporter molecules [4] [25].
This collateral cleavage activity serves as a powerful signal amplification mechanism for diagnostic applications [25]. When the ssDNA or ssRNA reporter is labeled with a fluorophore and a quencher, its cleavage releases the fluorophore, generating a detectable fluorescent signal [25]. Similarly, electrochemical signals can be generated when the reporter is cleaved from an electrode surface [28]. This "amplification-by-cleavage" allows for detectable signals from minute quantities of target nucleic acids without relying on complex thermal cycling equipment [25].
Table 1: Key Cas Proteins and Their Diagnostic Mechanisms
| Cas Protein | Target Type | Trans-Cleavage Substrate | PAM Requirement | Primary Diagnostic Applications |
|---|---|---|---|---|
| Cas12a (Type V) | dsDNA, ssDNA | ssDNA | 5' TTTN | DNA virus detection, bacterial pathogens [25] |
| Cas13a (Type VI) | RNA | ssRNA | None | RNA virus detection, gene expression monitoring [25] |
| Cas9 (Type II) | dsDNA | None (cis-cleavage only) | 5' NGG | Specific sequence identification, dCas9 for binding [25] |
| Cas12b (Type V) | dsDNA | ssDNA | 5' TTN | One-pot detection systems [25] |
Experimental Comparison: CRISPR vs. Culture Methods
Performance Metrics and Experimental Data
The following quantitative comparison summarizes experimental data from multiple studies directly or indirectly comparing CRISPR-based detection methods with traditional culture techniques for pathogen detection.
Table 2: Performance Comparison: CRISPR Diagnostics vs. Culture Methods
| Parameter | CRISPR-Based Detection | Traditional Culture Methods | Experimental Support |
|---|---|---|---|
| Detection Time | 30 minutes - 2 hours [30] | 24 hours - 10 days [11] | CRISPR: 40-60 min for SARS-CoV-2 with LFA [29]; Culture: 2-10 days for microbial culture [11] |
| Sensitivity | aM (10⁻¹⁸ M) to pM (10⁻¹² M) levels [4]; As low as 1 copy/μL for DNA targets [11] | Viable bacteria only, typically 10³-10⁴ CFU/mL [27] | E-CRISPR: pM level for HPV-16 without amplification [28]; CRISPR-Cas12a with RPA: 1 copy for Mpox DNA [11] |
| Specificity | Single-base discrimination possible [30] | Morphological and biochemical differentiation | SHERLOCK: 100% sensitivity and specificity in clinical samples [11] |
| Automation Potential | High (integrated systems with microfluidics) [30] | Low (multiple manual steps) | CRISPR reagents can be lyophilized, integrated into microfluidic platforms [4] |
| Biofilm Penetration | Detects released nucleic acids, not affected by biofilm matrix | Limited (requires disruption for effective culture) | Biofilm EPS can hinder antibiotic penetration and protect bacteria [26] |
| Viability Assessment | Cannot distinguish live/dead bacteria without pretreatment | Specifically detects viable organisms | CRISPR can target genes specific to viable cells (e.g., mRNA) but typically detects DNA regardless of viability [4] |
Experimental Protocols for CRISPR-Based Detection
E-CRISPR Protocol for Nucleic Acid Detection
The E-CRISPR (electrochemical CRISPR) platform provides a cost-effective and portable detection system [28]. The detailed methodology is as follows:
Sensor Preparation: A disposable, micro-fabricated gold-based three-electrode sensor with gold as working and counter electrodes and Ag/AgCl as the reference electrode is used [28].
Reporter Immobilization: A nonspecific ssDNA reporter is designed with a methylene blue (MB) electrochemical tag for signal transduction and a thiol moiety to tether on the sensor surface [28]. The surface density of the ssDNA reporter is optimized by controlling the concentration of the ssDNA reporter incubation solution to ensure accessibility for Cas12a cleavage [28].
Cas12a-crRNA Complex Formation: LbCas12a or AsCas12a is complexed with target-specific crRNA in a buffer containing optimized Mg²⁺ concentration (up to 15 mM), which enhances trans-cleavage activity [28].
Target Recognition and Cleavage: The Cas12a-crRNA complex is incubated with the sample. If the target DNA is present, the complex binds and activates trans-cleavage activity, cleaving the MB-ssDNA reporter off the electrode surface [28].
Signal Detection: Square wave voltammetry (SWV) is applied to evaluate the MB signal. A decreased MB signal indicates target presence and successful cleavage [28].
Key optimization parameters include selection of Cas12a analog (LbCas12a demonstrated more robust trans-cleavage than AsCas12a), trans-cleavage period (activity remains active for up to 3 hours), and Mg²⁺ concentration (15 mM optimal) [28].
ENHANCE Protocol with Engineered crRNAs
The ENHANCE (ENHanced Analysis of Nucleic acids with CrRNA Extensions) system improves detection sensitivity through engineered crRNAs [29]:
crRNA Engineering: Wild-type crRNAs are extended with a 7-mer ssDNA on the 3'-end, which augments the rate of LbCas12a-mediated collateral cleavage activity by approximately 3.5-fold compared to wild-type crRNA [29].
Reporter Design: A FRET-based reporter composed of a fluorophore (FAM or HEX) and a quencher (3IABkFQ) connected by a 5-nucleotide sequence (TTATT) is used, with TA-rich sequences providing maximal trans-cleavage activity [29].
Kinetic Enhancement: The 7-mer DNA extension on the 3'-end of crRNA increases the Kcat/Km ratio by 3.2-fold compared to unmodified crGFP, enhancing the rate of trans-cleavage without affecting binary complex formation between LbCas12a and crRNA [29].
Detection Integration: The system can be incorporated into paper-based lateral flow assays, detecting targets with up to 23-fold higher sensitivity within 40-60 minutes when combined with isothermal amplification [29].
Diagnostic Workflows: Traditional vs. CRISPR-Based Methods
The following diagrams illustrate the fundamental differences in workflow between traditional culture methods and CRISPR-based diagnostics for biofilm pathogen detection.
Diagram 1: Traditional Culture Method Workflow
Diagram 2: CRISPR-Based Detection Workflow
Research Reagent Solutions
The following table details essential materials and reagents required for implementing CRISPR-based diagnostic assays, particularly for biofilm pathogen detection.
Table 3: Essential Research Reagents for CRISPR Diagnostics
| Reagent/Category | Specific Examples | Function/Purpose | Optimization Notes |
|---|---|---|---|
| Cas Proteins | LbCas12a, AsCas12a, LbuCas13a | Target recognition and trans-cleavage activation | LbCas12a shows more robust trans-cleavage than AsCas12a; selection depends on PAM preferences [28] |
| crRNA Guides | Wild-type crRNA, Engineered crRNA with 3' DNA extensions | Target-specific recognition | 7-mer DNA extension on 3'-end enhances trans-cleavage activity 3.5-fold; spacer sequence determines target specificity [29] |
| Reporter Molecules | FAM/TTATT/BHQ-1 ssDNA reporters (for Cas12a), Methyleneblue-tagged ssDNA with thiol moiety | Signal generation through cleavage | TA-rich sequences maximize cleavage efficiency; electrochemical tags enable E-CRISPR platforms [28] [29] |
| Buffer Components | Mg²⁺, DTT, NEBuffer | Optimal enzymatic activity | Mg²⁺ concentration of 15 mM significantly enhances trans-cleavage; divalent cations essential for RuvC domain function [28] |
| Signal Detection Systems | Fluorescent plate readers, Electrochemical sensors, Lateral flow strips | Result readout | Lateral flow enables POC applications; electrochemical systems offer cost-effective portability [28] [30] |
| Nucleic Acid Amplification | RPA, LAMP kits | Pre-amplification for enhanced sensitivity | Enables attomolar sensitivity when combined with CRISPR; can be integrated in one-pot systems [11] [30] |
CRISPR-based diagnostic systems demonstrate clear advantages over traditional culture methods for detecting biofilm pathogens in terms of speed, sensitivity, and potential for automation. The molecular mechanisms of crRNA-guided target recognition and trans-cleavage activity provide a robust foundation for specific pathogen identification, with detection times reduced from days to hours or even minutes [28] [11] [30].
However, traditional culture methods remain essential for assessing bacterial viability and obtaining isolates for antibiotic susceptibility testing [27]. The optimal diagnostic approach depends on the specific application requirements: CRISPR diagnostics for rapid screening and culture methods for comprehensive phenotypic characterization. Future developments in CRISPR technology, including engineered crRNAs with enhanced sensitivity [29] and integrated sample-to-result systems [30], will further solidify its position as a transformative tool for biofilm pathogen detection in both clinical and research settings.
The persistent challenge of diagnosing polymicrobial biofilm-associated infections, which account for an estimated 60–80% of clinical infections in hospitalized patients, has exposed critical limitations in conventional culture-based methods [8]. Traditional diagnostic techniques often require several days, lack the sensitivity to detect low-abundance or slow-growing pathogens, and struggle to resolve complex microbial communities within biofilms [8] [31]. Within this diagnostic landscape, CRISPR-Cas systems have emerged as transformative tools for molecular diagnostics, offering unprecedented specificity, sensitivity, and rapid detection capabilities [4]. This guide provides a comprehensive comparative analysis of three principal Cas effectors—Cas12a, Cas13, and Cas14—specifically contextualized for application in biofilm pathogen detection research. We evaluate their mechanistic actions, performance parameters against traditional culture methods, and provide detailed experimental protocols to facilitate their implementation in diagnostic development.
Comparative Analysis of Cas Effector Mechanisms and Performance
The diagnostic application of Cas effectors leverages their unique nucleic acid recognition and collateral cleavage activities, enabling the development of highly sensitive detection platforms. The table below summarizes the core characteristics and diagnostic performance of Cas12a, Cas13, and Cas14.
Table 1: Comparative Analysis of Cas Effectors for Diagnostic Applications
| Feature | Cas12a (Cpf1) | Cas13a (C2c2) | Cas14 (Cas12f) |
|---|---|---|---|
| Primary Target | Double-stranded DNA (dsDNA) | Single-stranded RNA (ssRNA) | Single-stranded DNA (ssDNA) [32] |
| Collateral Cleavage Substrate | Single-stranded DNA (ssDNA) [4] | Single-stranded RNA (ssRNA) [4] | Single-stranded DNA (ssDNA) [33] |
| PAM Requirement | T-rich (e.g., TTTN) [4] | None for RNA target; PFS for some variants | None for ssDNA target [32] |
| Key Diagnostic Platform | DETECTR, HOLMES [4] [32] | SHERLOCK [4] [32] | Cas14-DETECTR [32] |
| Specificity | High; can distinguish bacterial species and strains [34] | Single-base mismatch specificity [32] | Extremely high; capable of SNP detection without pre-amplification [32] |
| Reported Sensitivity | ~100 fg DNA (Nocardia detection) [34] | Attomolar (aM) level [4] [32] | Attomolar (aM) level [32] |
| Key Advantage for Biofilms | Direct detection of bacterial DNA and resistance genes | Direct targeting of RNA, useful for viability assessment | Superior capability for identifying SNPs and minor genetic variations |
Cas12a: The DNA-Targeting Workhorse for Bacterial Identification
Cas12a is highly effective for detecting double-stranded DNA targets, making it ideal for identifying bacterial pathogens and antibiotic resistance genes (ARGs) within biofilm samples. Upon recognizing its target DNA sequence, which is guided by a CRISPR RNA (crRNA) and a T-rich Protospacer Adjacent Motif (PAM), Cas12a undergoes a conformational change that activates its non-specific trans-cleavage activity, indiscriminately degrading single-stranded DNA (ssDNA) reporters [4]. This collateral cleavage is harnessed for signal generation in diagnostic assays. For instance, a CRISPR-Cas12a platform combined with LAMP pre-amplification was developed to detect Nocardia farcinica with a limit of detection (LoD) of 100 fg DNA per reaction and no cross-reactivity with 132 non-target strains, demonstrating high specificity for a single bacterial species within a complex sample [34]. This precision is critical for pinpointing specific pathogens in polymicrobial biofilms.
Cas13: The RNA-Sensing Specialist for Transcriptional Profiling
Cas13 systems are uniquely designed to target single-stranded RNA molecules. After binding to its target RNA sequence guided by crRNA (without requiring a PAM sequence), Cas13 exhibits collateral cleavage of surrounding single-stranded RNA reporters [4] [11]. This activity is the foundation of the SHERLOCK platform. A key application in biofilm research is the potential for viability assessment, as RNA is typically more labile than DNA and its presence can correlate with metabolically active cells [11]. The Cas13-based SHERLOCKv2 platform, which utilizes multiple orthogonal Cas proteins, has demonstrated sensitivities as low as 8.0 x 10^-3 attomolar (aM), enabling the detection of extremely low levels of viral RNA, a performance that can be translated to bacterial mRNA targets in biofilms [32].
Cas14: The SNP-Discriminating Powerhouse for Resistance Mutations
Cas14 is a hyper-compact Cas effector that targets and cleaves single-stranded DNA (ssDNA) without requiring a PAM sequence for its activity [33] [32]. Similar to Cas12a, it exhibits promiscuous trans-cleavage of ssDNA reporters upon target recognition. Its most distinguishing feature is its exceptional specificity; Cas14 can discriminate single-nucleotide polymorphisms (SNPs) with high fidelity, making it exceptionally suited for identifying point mutations associated with antibiotic resistance [32]. The Cas14-DETECTR platform has been applied for high-fidelity genotyping, achieving attomolar sensitivity [32]. This capability allows researchers to track specific resistance alleles, such as SNPs in gyrA or rpoB genes, within the complex genetic landscape of a biofilm microbiome.
Experimental Protocols for CRISPR-Based Biofilm Pathogen Detection
Protocol 1: Cas12a-based Detection of Bacterial DNA from Biofilm Samples
This protocol, adapted from a study detecting Nocardia farcinica, outlines the steps for identifying a specific bacterial pathogen and its resistance genes from a biofilm sample using the CRISPR-CLA (CRISPR/Cas12a-based Detection with LAMP) method [34].
- Workflow Overview: The process involves DNA extraction from the biofilm, isothermal pre-amplification of the target gene, CRISPR/Cas12a detection, and result readout.
- Step-by-Step Procedure:
- Sample Preparation and DNA Extraction (20 min): Homogenize the biofilm sample in sterile PBS. Extract genomic DNA using a commercial kit (e.g., Wizard Genomic DNA Purification Kit). Quantify DNA using a microspectrophotometer [34].
- LAMP Preamplification (40 min): Design LAMP primers targeting a species-specific gene or antibiotic resistance gene. The forward inner primer (FIP) must be engineered to include the Cas12a PAM site (TTTA) in its linker region.
- Reaction Setup: Use a commercial DNA isothermal amplification kit. Assemble a 25 µL reaction containing LAMP premix, target DNA (e.g., 100 fg to 1 ng), and LAMP primer mix.
- Amplification: Incubate at 70°C for 40 minutes in a thermal cycler or isothermal bath, then halt the reaction at 80°C for 5 min [34].
- CRISPR/Cas12a-mediated detection (8 min):
- Reporter Preparation: Design an ssDNA reporter molecule labeled with a fluorophore (e.g., FAM) and a quencher (e.g., BHQ).
- crRNA Design: Design crRNAs of varying lengths complementary to the target LAMP amplicon and screen for the most efficient one.
- Detection Reaction: In a separate tube, mix EnGen Lba Cas12a, the selected crRNA, and the ssDNA reporter. Add 2 µL of the LAMP product to this mixture.
- Incubation: Incubate at 37°C for 8 minutes [34].
- Result Readout: Results can be read using a real-time fluorescence reader for quantitative analysis or a lateral flow biosensor for a visual, binary (yes/no) output. The cleavage of the reporter by activated Cas12a produces a fluorescent signal or a test line on the strip.
Protocol 2: Cas14-based Detection of Single-Nucleotide Polymorphisms
This protocol utilizes the high specificity of Cas14 to identify single-nucleotide changes, such as those conferring antibiotic resistance in biofilm bacteria [32].
- Workflow Overview: The process involves DNA extraction, optional PCR pre-amplification of the target region containing the SNP, and Cas14-mediated detection with result readout.
- Step-by-Step Procedure:
- DNA Extraction and Target Amplification: Extract DNA from the biofilm sample. Amplify the target region containing the SNP of interest using a specific PCR assay. This step increases the concentration of the target ssDNA for detection.
- Cas14-DETECTR Assay:
- crRNA Design: Design a crRNA that is perfectly complementary to the target sequence containing the SNP of interest. The high fidelity of Cas14 ensures that even a single mismatch will significantly reduce cleavage activity.
- Detection Reaction: In a single tube, combine the following:
- Cas14 protein
- Specific crRNA
- Fluorescently labeled ssDNA reporter (e.g., FAM-TTATT-BHQ-1)
- The PCR-amplified target DNA
- Incubation: Incubate the reaction at 37°C for 15-30 minutes [32].
- Result Readout: Measure the fluorescence signal in real-time or at the endpoint. A positive signal indicates the presence of the specific SNP in the sample. The attomolar sensitivity of the Cas14 system allows for the detection of low-abundance resistance mutations within a mixed population [32].
Diagram 1: CRISPR-Cas12a Diagnostic Workflow for Biofilm DNA Detection
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of CRISPR diagnostics for biofilm research requires a suite of specific reagents and tools. The following table details the essential components.
Table 2: Key Research Reagent Solutions for CRISPR-Cas Diagnostics
| Reagent / Tool | Function | Example & Notes |
|---|---|---|
| Cas Nuclease | The effector protein that executes target cleavage and collateral activity. | Recombinant EnGen Lba Cas12a (NEB), LwaCas13a, or purified Cas14 protein. Selection depends on the target (DNA/RNA) [34] [32]. |
| crRNA | Guides the Cas complex to the specific target nucleic acid sequence. | Synthetic crRNA designed to be complementary to a pathogen-specific gene (e.g., 16S rRNA, virulence factor) or an antibiotic resistance gene (e.g., mecA, blaKPC). Requires screening for optimal length and efficiency [4] [34]. |
| ssDNA/ssRNA Reporter | The molecule cleaved collateraly to generate a detectable signal. | For Cas12a/Cas14: ssDNA oligo labeled with Fluorophore (e.g., FAM) and Quencher (e.g., BHQ1). For Cas13: ssRNA reporter with similar labels [4] [34]. |
| Isothermal Amplification Kit | Preamplifies target nucleic acids to enhance detection sensitivity. | LAMP Kit (e.g., from HuiDeXin) or RPA Kit (e.g., TwistAmp). Crucial for detecting low-biomass targets in biofilms [34] [11]. |
| Lateral Flow Biosensor | Provides equipment-free, visual readout of results. | Commercial strips (e.g., from Milenia) that detect labeled reporters, often using FAM and biotin labels. Ideal for point-of-care application [34] [32]. |
| Fluorescence Reader | Enables quantitative, real-time measurement of the reaction signal. | Real-time qPCR instrument (e.g., QuantStudio) or portable fluorometer. Used for kinetic monitoring and obtaining quantitative data [34]. |
The strategic selection of Cas effectors—Cas12a for DNA targets, Cas13 for RNA, and Cas14 for SNPs—provides researchers with a powerful, versatile toolkit that fundamentally outperforms traditional culture methods in speed, specificity, and sensitivity for biofilm pathogen detection. While culture remains the historical gold standard, its prolonged turnaround time and inability to resolve complex communities are significant drawbacks in managing biofilm-associated infections [8] [31]. The integration of these CRISPR-based assays into streamlined workflows, including pre-amplification and diverse readout systems, enables the rapid (<70 minutes), accurate, and on-site identification of pathogens and their resistance profiles [4] [34]. This technological advancement paves the way for improved diagnostic stewardship and personalized treatment strategies against resilient polymicrobial biofilm infections.
In the critical field of biofilm pathogen detection, the limitations of traditional culture-based methods—including prolonged incubation times of 2-10 days and extensive technical requirements—have created an urgent need for rapid, precise diagnostic alternatives [11] [12]. Molecular methods such as quantitative PCR (qPCR) offer improved sensitivity but remain dependent on sophisticated thermal cycling equipment, restricting their use in point-of-care testing (POCT) scenarios [4]. The emergence of CRISPR-Cas systems has catalyzed a transformation in molecular diagnostics, enabling unprecedented sequence-specific detection capabilities. When integrated with isothermal amplification techniques—primarily Recombinase Polymerase Amplification (RPA) and Loop-Mediated Isothermal Amplification (LAMP)—these systems achieve ultimate sensitivity while maintaining operational simplicity [11] [35]. This review objectively evaluates integrated RPA-CRISPR and LAMP-CRISPR platforms, comparing their performance metrics, experimental protocols, and practical applications within biofilm pathogen detection research, providing researchers with critical insights for diagnostic development.
Fundamental Principles: CRISPR-Cas Systems and Isothermal Amplification
CRISPR-Cas Mechanism: From Bacterial Immunity to Diagnostic Tool
CRISPR-Cas systems function as adaptive immune mechanisms in prokaryotes, utilizing CRISPR RNA (crRNA) to guide Cas nucleases to complementary nucleic acid targets [36]. For diagnostic applications, Class 2 CRISPR systems—particularly Cas12 and Cas13—are most valuable due to their collateral trans-cleavage activity. Upon recognizing its target DNA, Cas12 exhibits nonspecific single-stranded DNA (ssDNA) cleavage, while Cas13 cleaves single-stranded RNA (ssRNA) after target recognition [11] [12]. Researchers leverage this trans-cleavage activity by designing reporter molecules—typically nucleic acid strands labeled with a fluorophore-quencher pair—that generate detectable signals when cleaved [12]. This mechanism provides the foundation for highly specific pathogen detection platforms such as DNA Endonuclease Targeted CRISPR Trans Reporter (DETECTR) utilizing Cas12 and Specific High Sensitivity Enzyme Reporter Unlocking (SHERLOCK) employing Cas13 [4].
Isothermal Amplification: RPA and LAMP
Isothermal amplification techniques enable rapid nucleic acid amplification at constant temperatures, eliminating dependence on thermal cyclers and facilitating point-of-care applications [35]. The table below compares the fundamental characteristics of RPA and LAMP:
Table 1: Comparison of Isothermal Amplification Techniques
| Characteristic | RPA (Recombinase Polymerase Amplification) | LAMP (Loop-Mediated Isothermal Amplification) |
|---|---|---|
| Reaction Temperature | 37-42°C | 60-65°C |
| Time to Result | 10-30 minutes | 15-60 minutes |
| Key Enzymes | Recombinase, SSB, DNA polymerase | Bst DNA polymerase |
| Primer Design | 2 primers (30-35 bp) | 4-6 primers (15-25 bp) |
| Sensitivity | 1-100 copies/μL | 1-100 copies/μL |
| Equipment Cost | Low | Medium |
| Advantages | Rapid, low temperature, simple design | Robust, highly specific, visual detection |
| Disadvantages | Primer design critical, limited throughput | Complex primer design, non-specific products |
RPA employs three core enzymes—recombinase, single-stranded binding protein (SSB), and strand-displacing DNA polymerase—that collaboratively enable exponential amplification of target sequences at 37-42°C within 10-30 minutes [35] [37]. The recombinase forms complexes with primers that scan double-stranded DNA for homologous sequences, initiating strand displacement and primer extension.
LAMP utilizes 4-6 specifically designed primers that recognize 6-8 distinct regions of the target DNA and the highly processive Bst DNA polymerase with strong strand-displacing activity [11]. The reaction occurs at 60-65°C, generating stem-loop DNA structures that enable continuous amplification with exceptional speed and efficiency.
Integrated Platform Performance: Comparative Analysis
RPA-CRISPR Integration
The combination of RPA with CRISPR-Cas12a creates a powerful diagnostic platform that leverages the rapid amplification capability of RPA with the precise recognition and signal generation of CRISPR [35]. This integrated system typically operates at 37°C, making it ideal for resource-limited settings. Performance data demonstrate that RPA-CRISPR-Cas12a achieves exceptional results in detecting biofilm-forming pathogens:
Table 2: Performance Metrics of RPA-CRISPR Platforms
| Pathogen Target | CRISPR System | Sensitivity | Detection Time | Key Performance Findings |
|---|---|---|---|---|
| Mpox Virus | Cas12a | 1 copy/μL | 30 minutes | Specific detection with minimal equipment [11] |
| Staphylococcus aureus | Cas12a | Single copy | <60 minutes | Effective for Gram-positive biofilm pathogens [35] |
| Pseudomonas aeruginosa | Cas9 | N/A | N/A | 90% biofilm biomass reduction with nanoparticle delivery [13] |
| Human Papillomavirus | Cas12a | attomolar level | <30 minutes | Clinical validation with high accuracy [37] |
The DETECTR platform, developed by Jennifer Doudna's team, exemplifies the RPA-CRISPR integration, combining efficient isothermal amplification with specific nucleic acid cleavage capability for real-time pathogen detection [35]. The system demonstrates particular strength in detecting bacterial pathogens with single-copy sensitivity, crucial for identifying low-abundance pathogens within complex biofilm matrices.
LAMP-CRISPR Integration
LAMP-CRISPR platforms leverage the robust amplification power of LAMP with CRISPR's specificity, creating systems capable of detecting pathogens directly from complex samples including biofilm isolates. The higher operating temperature of LAMP (60-65°C) provides inherent stringency that reduces non-specific amplification, while CRISPR confirmation eliminates false positives common with LAMP alone [11]. Research demonstrates that LAMP-CRISPR-Cas12a exhibits excellent performance in detecting bacterial pathogens with sensitivity comparable to gold-standard methods but with significantly reduced operational complexity [11] [12]. This integration is particularly valuable for biofilm research where sample processing may yield inhibitors that affect other amplification methods.
Figure 1: LAMP-CRISPR Integrated Workflow. The process begins with nucleic acid extraction from biofilm samples, followed by LAMP amplification at 60-65°C for 15-60 minutes. Amplified products are detected using CRISPR-Cas12a at 37°C, generating signals via fluorescent reporters or lateral flow assays (LFA).
Experimental Protocols: Methodology for Research Applications
RPA-CRISPR Protocol for Bacterial Pathogen Detection
Sample Preparation:
- Resuspend biofilm samples in digestion buffer containing proteinase K and lysozyme
- Incubate at 56°C for 30 minutes to degrade extracellular polymeric substances
- Extract nucleic acids using magnetic bead-based purification
- Elute in nuclease-free water and quantify using spectrophotometry
RPA Amplification:
- Prepare 50 μL reaction containing: 25 μL rehydration buffer, 5 μL template DNA, 2.5 μL forward primer (10 μM), 2.5 μL reverse primer (10 μM), 9 μL nuclease-free water
- Add 2.5 μL magnesium acetate (280 mM) to initiate reaction
- Incubate at 39°C for 15-30 minutes
- Terminate reaction by heating at 85°C for 5 minutes
CRISPR-Cas12a Detection:
- Prepare detection mix: 2 μL Cas12a (100 nM), 2 μL crRNA (100 nM), 2 μL reporter probe (500 nM FAM/TAMRA-BHQ quencher), 9 μL NEBuffer 2.1
- Add 5 μL RPA product to detection mix
- Incubate at 37°C for 10-30 minutes
- Measure fluorescence using plate reader or visual detection with blue light illuminator
Validation:
- Compare results with culture methods on selective media
- Confirm specificity with sequencing of representative samples
- Determine limit of detection using serial dilutions of quantified genomic DNA [35] [37]
LAMP-CRISPR Protocol for Multiplex Pathogen Detection
LAMP Reaction Setup:
- Design LAMP primers targeting conserved virulence or resistance genes in biofilm-forming pathogens
- Prepare 25 μL reaction: 1.6 μM each FIP/BIP primers, 0.2 μM each F3/B3 primers, 0.4 μM each LF/LB primers, 1.4 mM dNTPs, 0.8 M betaine, 20 mM Tris-HCl, 10 mM KCl, 8 mM MgSO₄, 10 mM (NH₄)₂SO₄, 0.1% Tween-20, 8 U Bst DNA polymerase
- Add 5 μL template DNA and incubate at 65°C for 30-45 minutes
- Heat-inactivate at 80°C for 5 minutes
CRISPR Detection with Lateral Flow Readout:
- Prepare Cas12a detection cocktail: 3 μL Cas12a (50 nM), 3 μL specific crRNA (50 nM), 3 μL reporter (500 nM FAM/Biotin-labeled ssDNA), 16 μL reaction buffer
- Combine 5 μL LAMP product with detection cocktail
- Incubate at 37°C for 15 minutes
- Apply 10 μL reaction to lateral flow strip
- Read results after 5-10 minutes development [11] [12]
Advanced Applications: Enhancing Sensitivity and Specificity
Signal Enhancement Strategies
Recent innovations focus on improving detection sensitivity through advanced reporter systems. Bead-based luciferase reporters represent a significant advancement, demonstrating up to 20× higher sensitivity compared to conventional fluorescent reporters in amplification-free CRISPR assays [14]. These systems utilize HiBiT and LgBiT peptide subunits coupled via Cas13-cleavable RNA linkers to separate beads, generating luminescent signals only upon target-activated collateral cleavage. For biofilm applications where pathogen concentrations may be low, such enhancement enables detection limits as low as 2.5 copies/μL of input RNA [14].
Nanoparticle-mediated delivery of CRISPR components further improves biofilm penetration and intracellular editing efficiency. Liposomal Cas9 formulations reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro, while gold nanoparticle carriers enhance editing efficiency up to 3.5-fold compared to non-carrier systems [13]. These approaches address the critical challenge of delivering molecular tools through the protective extracellular polymeric substance matrix of biofilms.
Multiplex Detection Platforms
Bead-based color-coded systems enable highly multiplexed pathogen detection, essential for identifying polymicrobial biofilms. The Bead-Based CARMEN (bbCARMEN) platform utilizes fluorescently color-coded beads coupled to distinct crRNAs, enabling parallelized detection of nine distinct viral targets with sensitivity reaching 2.5 copies/μL [14]. This approach permits comprehensive pathogen profiling within complex biofilm communities without significant infrastructure requirements.
Figure 2: Multiplex Detection Platform. Biofilm samples undergo nucleic acid extraction followed by multiplex RPA or LAMP amplification. Color-coded beads with specific crRNAs are combined with amplified products and partitioned into droplets for parallelized CRISPR detection, enabling simultaneous identification of multiple pathogens.
Research Reagent Solutions: Essential Materials for Implementation
Successful implementation of RPA-CRISPR and LAMP-CRISPR platforms requires specific reagent systems optimized for integrated assays:
Table 3: Essential Research Reagents for RPA/LAMP-CRISPR Assays
| Reagent Category | Specific Products | Application Function | Considerations for Biofilm Research |
|---|---|---|---|
| Amplification Kits | TwistAmp Basic RPA Kit | Provides core recombinase/polymerase enzymes | Optimize magnesium concentration for biofilm inhibitors |
| CRISPR Enzymes | LbCas12a, AsCas12a, LwCas13a | Target recognition and trans-cleavage | Cas12a for DNA targets, Cas13 for RNA targets |
| crRNA Design | Custom synthetic crRNAs | Target-specific guidance | Design against conserved biofilm genes (16S rRNA, virulence factors) |
| Reporter Systems | FAM-TTATT-BHQ-1 ssDNA | Fluorescent signal generation | Validate against sample autofluorescence |
| Lateral Flow Strips | Milenia HybriDetect | Visual readout | Ideal for resource-limited settings |
| Sample Preparation | Proteinase K, lysozyme | Biofilm matrix degradation | Critical for efficient nucleic acid release |
| Nucleic Acid Purification | Magnetic bead-based kits | Inhibitor removal | Essential for complex biofilm samples |
RPA-CRISPR and LAMP-CRISPR platforms represent significant advancements over traditional culture methods for biofilm pathogen detection, offering rapid results (30-60 minutes versus 2-10 days), exceptional sensitivity (single-copy detection), and minimal equipment requirements [11] [12]. While RPA-CRISPR excels in speed and low-temperature operation, LAMP-CRISPR provides robust amplification and higher reaction stringency. The integration of nanoparticle delivery systems and enhanced reporter technologies continues to address challenges related to biofilm penetration and signal detection [13] [14]. For researchers developing diagnostic applications for biofilm-associated pathogens, these amplification-based CRISPR strategies provide powerful, adaptable tools that balance ultimate sensitivity with practical implementation requirements, potentially revolutionizing how we detect and monitor complex microbial communities in clinical, industrial, and environmental settings.
In the field of pathogen detection research, particularly for biofilm-associated infections, the scientific community has long relied on traditional culture methods as the "gold standard." While these methods provide valuable information on microbial viability, they are notoriously time-consuming, often requiring 2-10 days for completion, and demand significant technical expertise and biosafety precautions [12] [11]. The limitations of these conventional approaches have become increasingly apparent in the face of urgent diagnostic needs during pathogen outbreaks, driving the development of innovative molecular diagnostics that offer rapid, sensitive, and specific alternatives.
Among these innovations, Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) technology has emerged as a revolutionary tool for molecular diagnostics. CRISPR-based detection methods are primarily categorized as amplification-based and amplification-free approaches [12]. While amplification-based CRISPR methods (such as those combined with RPA or LAMP) offer exceptional sensitivity, they introduce operational complexity and potential contamination risks from the nucleic acid amplification step [12] [4]. In response to these challenges, amplification-free CRISPR strategies have gained significant attention for their ability to enable direct pathogen detection without target amplification, thereby reducing operational complexity, minimizing contamination risks, and shortening detection time [12]. This review focuses specifically on cascade systems and sensor technologies within amplification-free CRISPR, evaluating their performance against traditional culture methods and other molecular alternatives in the context of biofilm pathogen detection research.
Principles of Amplification-Free CRISPR Detection
Fundamental Mechanisms of CRISPR-Cas Systems
The CRISPR-Cas system functions as an adaptive immune mechanism in bacteria and archaea, capable of recognizing and cleaving specific nucleic acid sequences [38]. This system comprises two key components: the Cas protein (nuclease) and a guide RNA (gRNA) that directs the Cas protein to target DNA or RNA sequences through complementary base pairing [12] [38]. For detection applications, researchers leverage the trans-cleavage activity exhibited by certain Cas proteins such as Cas12, Cas13, and Cas14 – a phenomenon where upon recognizing and binding to its specific target sequence, the Cas protein becomes activated to non-specifically cleave surrounding nucleic acid molecules [12] [4].
This trans-cleavage activity serves as the fundamental basis for CRISPR-based detection technologies. By labeling reporter molecules (single-stranded nucleic acid fragments) with fluorescent signals (e.g., FAM, ROX) and quenching groups (e.g., BHQ), target recognition can be converted into a detectable signal: when the Cas protein is activated by its target, it cleaves the reporter molecule, separating the fluorophore from the quencher and generating a fluorescent signal [12]. Alternatively, reporters labeled with biotin and FAM or digoxigenin can be used with lateral flow assay strips for visual detection without specialized equipment, significantly expanding the point-of-care application potential of CRISPR technology [12].
The Amplification-Free Advantage
Amplification-free CRISPR detection emphasizes the direct detection of pathogen nucleic acids without prior amplification steps [12]. This approach offers several distinct advantages for biofilm pathogen detection research:
- Reduced operational complexity: By eliminating nucleic acid amplification steps, the detection process becomes simpler and more streamlined
- Decreased contamination risk: Without amplification reactions, the potential for false positives from amplicon contamination is significantly reduced
- Faster detection times: Direct detection can yield results more quickly, often within 30-40 minutes [12] [39]
- Quantitative potential: Without amplification biases, results may more accurately reflect original target concentrations
- Point-of-care suitability: Simplified workflows are more adaptable to resource-limited settings
The development of amplification-free CRISPR represents a significant advancement toward creating more efficient and convenient pathogen detection tools, particularly for challenging applications like biofilm-associated infections where traditional culture methods are slow and molecular methods can be complex [12].
Cascade CRISPR Systems for Direct Detection
Principles of Cascade Systems
Cascade CRISPR systems represent a sophisticated amplification-free approach that employs multiple Cas complexes in a sequential activation mechanism to achieve signal amplification without target nucleic acid amplification [12]. These systems are designed to overcome the sensitivity limitations of simple CRISPR detection by creating a cascading effect where the initial target recognition triggers a series of subsequent reactions that collectively amplify the detection signal.
The fundamental principle involves an initial Cas complex (RNP1) that recognizes the original pathogen-derived target sequence. Upon activation, this complex cleaves not only the reporter molecules but also specific activator molecules or blocking domains that normally suppress secondary Cas complexes. The removal of these suppression elements activates secondary Cas complexes (RNP2), which then contribute additional reporter cleavage, thereby amplifying the detection signal without requiring target amplification [39]. This elegant molecular circuitry enables extremely sensitive detection while maintaining the specificity inherent to CRISPR systems.
Experimental Protocol: TCC Method
A groundbreaking example of cascade CRISPR technology is the Target-amplification-free Collateral-cleavage-enhancing CRISPR-CasΦ method (TCC), which achieves exceptional sensitivity through an ingeniously designed DNA amplifier [39]. The experimental workflow proceeds as follows:
Sample Preparation: Target pathogens are lysed to release genomic DNA using thermal or chemical lysis methods. For biofilm samples, additional steps may be required to disrupt the extracellular polymeric substance matrix before pathogen lysis.
Reaction Setup: The one-pot reaction mixture is assembled containing:
- CasΦ protein (a compact member of the Cas12 family)
- Two guide RNAs (gRNA1 for target recognition, gRNA2 for amplifier recognition)
- TCC amplifier (a dual stem-loop DNA structure)
- Fluorescent reporter (ssDNA with fluorophore and quencher)
- Microbial lysate containing target DNA
Reaction Incubation: The mixture is incubated at 37°C for 40 minutes to allow the cascading reaction to proceed.
Signal Detection: Fluorescence is measured in real-time or at endpoint using a plate reader or portable fluorescence detector.
In the TCC mechanism, RNP1 (CasΦ + gRNA1) first recognizes the target pathogen DNA, activating its trans-cleavage activity. The activated RNP1 then cleaves the stem-loops in the TCC amplifier, generating a product that activates RNP2 (CasΦ + gRNA2). The activated RNP2 subsequently cleaves numerous fluorescent reporters, generating a detectable signal while simultaneously cleaving more TCC amplifiers to perpetuate the cascade [39].
The following diagram illustrates the sophisticated molecular circuitry of this cascade system:
Figure 1: Cascade mechanism of the TCC amplification-free CRISPR detection system
Performance Data and Comparison
The TCC method demonstrates exceptional performance characteristics for pathogen detection [39]:
Table 1: Performance metrics of the TCC cascade CRISPR system
| Parameter | Performance | Comparative Context |
|---|---|---|
| Detection Limit | 0.11 copies/μL (0.18 aM) | 100-1000x more sensitive than qPCR |
| Detection Time | 40 minutes | ~5x faster than culture methods |
| Sensitivity (Clinical Samples) | 1.2 CFU/mL in serum | Superior to blood culture for low pathogen loads |
| Specificity | High | Comparable to other CRISPR methods |
| Sample Processing | One-pot reaction | Simplified workflow |
| Multiplexing Potential | Demonstrated for multiple pathogens | S. aureus, P. aeruginosa, K. pneumoniae, E. coli |
This exceptional sensitivity enables detection of pathogenic bacteria at concentrations as low as 1.2 CFU/mL in clinical serum samples, surpassing both qPCR and traditional culture methods for low-abundance pathogens [39]. For biofilm pathogen research, this level of sensitivity is particularly valuable for detecting residual pathogens after treatment or for identifying early-stage biofilm formation.
Sensor-Integrated CRISPR Technologies
The integration of CRISPR-based detection with advanced sensor technologies represents another innovative approach to amplification-free pathogen detection [12]. These hybrid systems leverage the specificity of CRISPR-Cas systems for target recognition with the sensitivity and real-time monitoring capabilities of various sensor platforms. The primary sensor technologies being explored for integration with CRISPR include:
- gFET (graphene field-effect transistor) biosensors: Leverage the electrical properties of graphene to detect binding events
- ECL (electrochemiluminescence) sensors: Measure light emission from electrochemical reactions
- SERS (surface-enhanced Raman spectroscopy): Enhances Raman scattering signals for highly sensitive detection
- Other electronic and optical sensors: Various platforms that convert molecular recognition into measurable signals
These sensor-integrated CRISPR platforms enable direct monitoring of targets through signal transduction, bypassing the need for nucleic acid amplification while maintaining high sensitivity and specificity [12] [40]. The combination creates synergistic systems that capitalize on the strengths of both technologies.
Experimental Protocol: FET-CRISPR Biosensor
A notable example of sensor-integrated CRISPR technology is the FET (Field-Effect Transistor) biosensor coupled with CRISPR-Cas12a for direct multiplex detection of viral nucleic acids [40]. The experimental protocol involves:
Biosensor Functionalization:
- Graphene FET channels are modified with probe DNA sequences complementary to target pathogen nucleic acids
- Surface chemistry is optimized to enhance binding efficiency and minimize non-specific adsorption
CRISPR-Cas12a Preparation:
- Cas12a protein is pre-complexed with target-specific crRNAs to form ribonucleoprotein (RNP) complexes
- Multiple RNP complexes with different specificities can be prepared for multiplex detection
Sample Processing:
- Clinical samples are processed to release nucleic acids without amplification
- Samples are diluted in appropriate buffer solutions for optimal binding conditions
Detection Reaction:
- Processed samples are applied to the functionalized FET biosensor
- CRISPR-Cas12a RNP complexes are added to recognize specific targets
- Upon target recognition, activated Cas12a cleaves nearby reporter molecules and potentially disrupts the sensor surface
Signal Measurement:
- Electrical parameters (conductance, resistance, current) are measured in real-time
- Changes in electrical properties indicate target recognition and Cas12a activation
- Data is processed to identify and quantify specific pathogens
This sensor-integrated approach enables direct, amplification-free detection of full-length viral nucleic acids across multiple pathogens in a single assay [40]. The method is particularly valuable for surveillance and diagnostic applications where multiple potential pathogens must be screened simultaneously.
Performance Comparison of Sensor Technologies
Sensor-integrated CRISPR platforms demonstrate varying performance characteristics based on the specific technology employed:
Table 2: Comparison of sensor technologies integrated with CRISPR for amplification-free detection
| Sensor Technology | Detection Principle | Sensitivity | Advantages | Limitations |
|---|---|---|---|---|
| gFET Biosensor | Electrical conductance changes | High (sub-femtomolar) | Label-free, real-time monitoring, miniaturization potential | Complex fabrication, signal interference risks |
| ECL Sensor | Light emission from electrochemical reactions | High (femtomolar) | High signal-to-noise ratio, wide dynamic range | Requires electrode systems, more complex instrumentation |
| SERS | Enhanced Raman scattering | Ultra-high (single molecule potential) | Excellent multiplexing capability, fingerprint spectra | Complex substrate preparation, potential inconsistency |
| Lateral Flow | Visual detection on strips | Moderate (picomolar) | Low cost, ease of use, no instrumentation needed | Lower sensitivity, semi-quantitative at best |
Each sensor technology offers distinct advantages that make it suitable for different applications in biofilm pathogen detection research. gFET biosensors provide real-time, label-free monitoring capabilities ideal for kinetic studies, while ECL sensors offer exceptional sensitivity for detecting low-abundance targets. SERS technology enables excellent multiplexing for detecting multiple biofilm-related pathogens simultaneously, and lateral flow assays provide simplicity and rapidity for point-of-care applications.
Comparative Performance Analysis
Benchmarking Against Traditional and Molecular Methods
When evaluating amplification-free CRISPR technologies against traditional culture methods and other molecular diagnostics for biofilm pathogen detection, distinct performance patterns emerge:
Table 3: Comprehensive comparison of detection methods for biofilm pathogens
| Method | Sensitivity | Time to Result | Viability Information | Equipment Needs | Skill Requirements | Cost per Test |
|---|---|---|---|---|---|---|
| Culture Methods | Moderate (requires viable cells) | 2-10 days [12] | Yes | Incubators, biosafety facilities | High | Low to moderate |
| qPCR | High (0.1×10⁴-10⁵ copies/mL) [39] | 2-4 hours | No | Thermal cycler, real-time detection | Moderate to high | Moderate |
| Amplification-Based CRISPR | Very high (single copy) [12] | 30-90 minutes | No | Heating block, fluorescence reader | Moderate | Low to moderate |
| Cascade CRISPR (TCC) | Ultra-high (0.11 copies/μL) [39] | 40 minutes | No | Heating block, fluorescence reader | Moderate | Moderate |
| Sensor-Integrated CRISPR | High to ultra-high | 30-60 minutes | No | Specialized sensor instrumentation | High | High |
This comparative analysis reveals that amplification-free CRISPR technologies, particularly cascade systems like TCC, offer an exceptional combination of sensitivity and speed that surpasses both traditional culture methods and other molecular diagnostics. While culture methods remain valuable for providing viability information, their extended time-to-result significantly limits their utility for rapid diagnostics and time-sensitive treatment decisions.
Advantages for Biofilm Pathogen Detection
Amplification-free CRISPR technologies offer several specific advantages for biofilm pathogen detection research:
- Rapid results: Detection within 30-40 minutes enables timely intervention and monitoring
- Exceptional sensitivity: Capable of detecting low-abundance pathogens in complex biofilm samples
- Reduced contamination risk: Elimination of amplification steps minimizes false positives
- Quantitative potential: Direct detection may enable more accurate quantification
- Multiplexing capability: Simultaneous detection of multiple biofilm-related pathogens
- Point-of-care suitability: Simplified workflows facilitate use in resource-limited settings
For biofilm-associated infections, where conventional antimicrobial therapies often fail due to the protective extracellular polymeric substance matrix and phenotypic heterogeneity of bacterial cells [13], rapid and accurate pathogen detection is crucial for effective treatment. Amplification-free CRISPR technologies address this need by providing sensitive detection without the time-consuming cultivation steps required by traditional methods.
The Scientist's Toolkit: Essential Research Reagents
Implementing amplification-free CRISPR detection methods requires specific reagents and components, each serving a critical function in the detection system:
Table 4: Essential research reagents for amplification-free CRISPR detection
| Reagent/Category | Specific Examples | Function | Considerations for Biofilm Research |
|---|---|---|---|
| Cas Proteins | CasΦ (Cas12j), Cas12a, Cas13, Cas14 | Target recognition and trans-cleavage | CasΦ offers compact size and high activity; Cas12a targets DNA pathogens |
| Guide RNAs | Target-specific crRNAs, gRNA1, gRNA2 | Sequence-specific targeting | Design for conserved biofilm-related genes (e.g., 16S rRNA, virulence factors) |
| Signal Amplifiers | TCC amplifier (dual stem-loop) | Signal enhancement without target amplification | Optimize stem-loop structures for efficient cleavage |
| Reporters | FQ-reporters (fluorophore-quencher), lateral flow reporters | Signal generation | Choice depends on detection platform (fluorescence vs. visual) |
| Sensor Platforms | gFET chips, ECL electrodes, SERS substrates | Signal transduction and measurement | gFET enables real-time monitoring of biofilm formation dynamics |
| Buffer Systems | Reaction buffers, lysis buffers | Optimal enzymatic activity | Include components to inhibit biofilm-derived enzymatic inhibitors |
| Control Templates | Synthetic pathogen DNA/RNA | Assay validation and standardization | Include positive controls for biofilm-associated pathogens |
This toolkit provides researchers with the essential components for developing and implementing amplification-free CRISPR detection systems tailored to biofilm pathogen research. Careful selection and optimization of each component are crucial for achieving optimal performance in specific research applications.
Amplification-free CRISPR technologies, particularly cascade systems and sensor-integrated platforms, represent a significant advancement in pathogen detection methodology with profound implications for biofilm research. These approaches offer an exceptional combination of sensitivity, speed, and specificity that addresses critical limitations of both traditional culture methods and amplification-based molecular diagnostics.
For researchers investigating biofilm-associated infections, these technologies enable rapid detection of pathogens at clinically relevant concentrations without the extended incubation periods required by culture methods. The amplification-free nature of these systems reduces complexity and contamination risks while potentially providing more quantitative results that accurately reflect pathogen load—a crucial parameter for evaluating treatment efficacy against biofilm infections.
Future developments in amplification-free CRISPR will likely focus on enhancing multiplexing capabilities for simultaneous detection of multiple biofilm-related pathogens, improving sensitivity further to detect even lower pathogen loads, and increasing automation for streamlined workflows. Additionally, integration with portable devices and point-of-care platforms will expand the utility of these technologies beyond laboratory settings to clinical environments where rapid diagnosis of biofilm-associated infections can significantly impact patient outcomes.
As these technologies continue to evolve, they hold tremendous promise for transforming how researchers detect and study biofilm pathogens, ultimately contributing to improved understanding and treatment of these challenging infections. The exceptional performance of amplification-free CRISPR systems positions them as powerful tools that will likely play an increasingly important role in both basic research and clinical applications for biofilm-associated infections.
The detection and identification of biofilm-forming pathogens are critical in clinical diagnostics and food safety. Traditional culture-based methods, while considered a gold standard, are often time-consuming, requiring 24-48 hours or more to yield results, and can struggle to detect low-abundance or slow-growing pathogens within complex biofilm matrices [4] [16]. In contrast, CRISPR-based diagnostic platforms offer a paradigm shift by providing rapid, highly sensitive, and specific detection of nucleic acids, making them particularly suited for analyzing biofilm-derived pathogens where sensitivity and speed are paramount [4] [41]. These systems can be designed to target conserved genomic regions of pathogens, allowing for precise identification directly from samples, potentially without the need for culture enrichment [4].
Among the most advanced CRISPR diagnostic platforms are SHERLOCK (Specific High-Sensitivity Enzymatic Reporter UnLOCKing) and DETECTR (DNA Endonuclease Targeted CRISPR Trans Reporter). A key feature of both systems is their compatibility with lateral flow readouts, enabling visual detection of results without sophisticated instrumentation [42] [43]. This review provides a objective comparison of the SHERLOCK and DETECTR platforms, focusing on their application in pathogen detection. It summarizes their core mechanisms, performance data, and experimental protocols, with particular attention to their utility in a research context focused on biofilm pathogens.
SHERLOCK and DETECTR leverage distinct CRISPR effector proteins with collateral nuclease activity, which is activated upon recognition of a target nucleic acid sequence. This activation triggers the cleavage of reporter molecules, generating a detectable signal.
SHERLOCK utilizes Cas13a, an RNA-guided RNase that, upon binding to a specific target RNA sequence, exhibits collateral activity by cleaving nearby non-target RNA molecules [44] [45]. In a typical assay, the target RNA is first amplified isothermally (e.g., via RPA or RT-RPA). The amplified product is then detected by Cas13a complexed with a specific crRNA. Activation of Cas13a leads to the cleavage of a quenched fluorescent RNA reporter, releasing a fluorescent signal. The SHERLOCKv2 platform incorporates additional effectors like Cas13b and Csm6 to enable multiplexed detection and enhance sensitivity, and it adapts the readout for lateral flow strips [45].
DETECTR employs Cas12a (Cpf1), an RNA-guided DNase that, upon binding to its target DNA sequence, promiscuously cleaves nearby non-target single-stranded DNA (ssDNA) molecules [44] [45]. Similar to SHERLOCK, the target DNA is often pre-amplified (e.g., via RPA). Cas12a, guided by its crRNA, recognizes the amplicon and becomes activated, subsequently cleaving a fluorescently quenched ssDNA reporter. This system is also fully compatible with lateral flow readouts [43].
Table 1: Core Components and Mechanisms of SHERLOCK and DETECTR
| Feature | SHERLOCK | DETECTR |
|---|---|---|
| CRISPR Effector | Cas13 (e.g., LwaCas13a, LbuCas13a) | Cas12 (e.g., LbCas12a, AsCas12a) |
| Target Nucleic Acid | RNA | DNA |
| Collateral Activity | Trans-cleavage of ssRNA | Trans-cleavage of ssDNA |
| Primary Amplification Method | RT-RPA or RT-LAMP | RPA or LAMP |
| Reporter Molecule | Fluorescently quenched ssRNA | Fluorescently quenched ssDNA |
| Key Recognition Element | crRNA | crRNA |
| PAM/PFS Requirement | Some orthologs require a PFS (Protospacer Flanking Site) | Requires a T-rich PAM (Protospacer Adjacent Motif) |
The Role of Lateral Flow Readouts
Both platforms are frequently paired with lateral flow strips (e.g., Milenia HybriDetect) for simple, instrument-free visual detection [43]. The mechanism involves a reporter molecule labeled with biotin and a fluorescent tag (e.g., FAM). When the collateral cleavage activity is activated, the reporter is degraded, preventing the formation of a test line. Therefore, the appearance of both control and test lines indicates a negative result, while the presence of only the control line indicates a positive result. This "inverse" approach requires careful optimization of the reporter concentration to ensure complete cleavage for a positive result [43].
Performance Comparison and Experimental Data
Clinical and analytical validation studies demonstrate that both SHERLOCK and DETECTR achieve performance characteristics comparable to traditional PCR-based methods, but with faster turnaround times and simpler workflows.
A clinical validation of a Cas13-based SHERLOCK assay for SARS-CoV-2 detection reported 100% specificity and 96% sensitivity with a fluorescence readout when tested against 154 clinical samples, with a limit of detection (LoD) of 42 RNA copies per reaction [42]. When the readout was adapted to lateral flow strips, the sensitivity was 97% within the characterized LoD [42].
The DETECTR system has shown similar high performance. For SARS-CoV-2 detection, it has demonstrated a LoD of 10 copies/μL and results in about 30 minutes [41]. In the context of other pathogens, such as HPV, DETECTR has been reported to achieve 95% sensitivity and 98% specificity [41].
Recent advancements focus on simplifying the workflow. The "real-time SLK" method, which uses novel thermostable Cas12 enzymes (SLK9 and SLK5-2) active at 60°C, allows for a single-step, real-time reaction by combining RT-LAMP amplification and Cas detection. This method demonstrated 100% positive and negative percent agreement with RT-qPCR on clinical samples and enabled direct testing from anterior nasal swabs without nucleic acid extraction [46]. Furthermore, this system has been successfully multiplexed to simultaneously detect SARS-CoV-2 and a human internal control in a single reaction [46].
Table 2: Comparative Performance Data for Pathogen Detection
| Platform / Assay | Target Pathogen | Sensitivity | Specificity | Limit of Detection (LoD) | Time to Result |
|---|---|---|---|---|---|
| SHERLOCK (Fluorescence) [42] | SARS-CoV-2 | 96% | 100% | 42 copies/reaction | ~60 min |
| SHERLOCK (Lateral Flow) [42] | SARS-CoV-2 | 97%* | 100% | 42 copies/reaction | ~60 min |
| DETECTR [41] | SARS-CoV-2 | ~98% | ~100% | 10 copies/μL | ~30-45 min |
| DETECTR [41] | HPV | 95% | 98% | 10 copies/μL | ~60 min |
| Real-time SLK (Cas12) [46] | SARS-CoV-2 | 100% | 100% | 0.5 - 5 cp/μL (extracted) | < 30 min |
| Cas13-based (SHERLOCK) [41] | Dengue Virus | 95% | 98% | 1 aM (attomolar) | ~60 min |
Sensitivity was 97% within the characterized LoD with lateral flow readout [42].
Experimental Protocols for Research Use
This section outlines core protocols for implementing SHERLOCK and DETECTR assays in a research setting, which can be adapted for detecting biofilm-associated pathogens.
General Workflow for SHERLOCK (Cas13-based detection of RNA targets)
- Nucleic Acid Extraction: Extract total nucleic acid from the sample (e.g., biofilm suspension, clinical swab). For biofilm samples, mechanical disruption (e.g., bead beating) may be necessary to lyse cells effectively.
- Reverse Transcription & Isothermal Amplification (RT-RPA):
- Prepare an RT-RPA master mix containing reverse transcriptase, recombinase enzymes, primers, and dNTPs.
- Add the extracted RNA template.
- Incubate at 37-42°C for 15-25 minutes to amplify the target RNA sequence.
- CRISPR-Cas13 Detection:
- Prepare a detection mix containing the Cas13 protein, specific crRNA, and the fluorescent RNA reporter (e.g., 6-FAM/UU/3BHQ-1).
- Transfer a small aliquot (e.g., 2-5 μL) of the amplified RPA product to the detection mix.
- Incubate at 37°C for 15-30 minutes to allow for target recognition and reporter cleavage.
- Readout:
- Fluorescence: Measure fluorescence intensity using a plate reader or fluorometer.
- Lateral Flow: Apply the reaction mixture to a lateral flow strip (e.g., Milenia HybriDetect). For Cas13, the reporter is typically an RNA molecule tagged with FITC and biotin. Interpret results after 2-5 minutes: positive = control line only; negative = both control and test lines [43].
General Workflow for DETECTR (Cas12-based detection of DNA targets)
- Nucleic Acid Extraction: Extract DNA from the sample. As with SHERLOCK, optimize lysis for biofilm matrices.
- Isothermal Amplification (RPA):
- Prepare an RPA master mix with recombinase enzymes, primers, and dNTPs.
- Add the extracted DNA template.
- Incubate at 37-42°C for 15-25 minutes.
- CRISPR-Cas12 Detection:
- Prepare a detection mix containing the Cas12a protein, specific crRNA, and a fluorescent ssDNA reporter (e.g., 5'-6-FAM/TTATT/3IBFQ-3').
- Transfer an aliquot of the RPA product to the detection mix.
- Incubate at 37°C for 15-30 minutes.
- Readout:
- Fluorescence: Measure fluorescence release.
- Lateral Flow: The reporter is typically a ssDNA oligo tagged with FAM and biotin. Apply the reaction to the strip. The interpretation is the same as for SHERLOCK: a positive result shows only the control line [43].
Protocol for Single-Pot, Real-Time SLK Assay
This advanced protocol leverages thermostable Cas12 enzymes for a simplified workflow [46].
- Sample Preparation: For direct assays, samples (e.g., nasal swabs) can be subjected to a simple thermal lysis step instead of full nucleic acid extraction.
- Single-Pot Reaction Setup:
- Prepare a master mix containing:
- Amplification components: RT-LAMP primers (FIP, BIP, F3, B3, LF, LB), MgSO4, and dNTPs.
- Detection components: Thermostable Cas12 enzyme (e.g., SLK9), specific crRNA, and a fluorescent ssDNA reporter.
- Add the prepared sample (containing RNA/DNA) to the master mix.
- Prepare a master mix containing:
- Incubation and Readout:
- Incubate the reaction tube at 60°C for 20-30 minutes in a real-time fluorometer or a simple heat block.
- Monitor fluorescence in real-time. A positive result is indicated by a rapid increase in fluorescence signal, providing a time-to-result under 30 minutes.
Research Reagent Solutions
A successful CRISPR diagnostic assay requires several key reagents, each with a critical function.
Table 3: Essential Reagents for SHERLOCK and DETECTR Assays
| Reagent / Material | Function | Example & Notes |
|---|---|---|
| CRISPR Effector Protein | Binds target nucleic acid via crRNA; provides collateral nuclease activity. | LwaCas13a (for SHERLOCK), LbCas12a (for DETECTR). Thermostable variants (e.g., SLK9) enable single-pot assays [46]. |
| crRNA (CRISPR RNA) | Guides the Cas protein to the specific target sequence. | Synthesized oligos; must be designed to be complementary to the target pathogen's genome (e.g., SARS-CoV-2 N gene) [46]. |
| Isothermal Amplification Kit | Amplifies target nucleic acid to detectable levels without thermal cycling. | RPA Kit (TwistAmp) for DNA/DETECTR; RT-RPA Kit for RNA/SHERLOCK. LAMP Kit for use with thermostable Cas enzymes [46] [42]. |
| Fluorescent Reporter | Molecule cleaved by activated Cas protein to generate signal. | ssRNA reporter (6-FAM/rUrUrU/3BHQ-1) for Cas13; ssDNA reporter (5'-6-FAM/TTATT/3IBFQ-3') for Cas12 [45] [43]. |
| Lateral Flow Strips | Provides instrument-free, visual readout of the assay result. | Milenia HybriDetect strips. Critical to optimize reporter concentration to avoid the high-dose hook effect [43]. |
| Target-Specific Primers | Amplifies the target region during the RPA/LAMP step. | Designed to be highly specific to the pathogen of interest; sequences are often published in methods sections of relevant papers [46] [42]. |
Technical Diagrams
SHERLOCK Cas13a Assay Workflow
SHERLOCK Assay Flow
DETECTR Cas12a Assay Workflow
DETECTR Assay Flow
Lateral Flow Readout Mechanism
Lateral Flow Result Interpretation
Navigating Technical Hurdles: Optimizing CRISPR Diagnostics for Complex Biofilm Samples
The accurate detection of biofilm-associated pathogens is pivotal for effective diagnosis and treatment in clinical and research settings. However, traditional diagnostic methods and even novel molecular techniques face significant challenges from sample interference. Clinical matrices such as blood, sputum, and urine contain inherent inhibitors that can compromise assay efficiency, while the extracellular polymeric substance (EPS) in biofilms creates a formidable barrier to detection and eradication. This review objectively compares the performance of CRISPR-based diagnostics against conventional culture methods, with a specific focus on their resilience to these interfering substances. We synthesize recent experimental data to provide researchers with a clear understanding of the capabilities and limitations of each approach in complex sample environments, enabling informed selection of appropriate methodologies for biofilm pathogen detection research.
Performance Comparison: CRISPR Diagnostics vs. Culture Methods
The following tables summarize experimental data comparing the performance of CRISPR-based diagnostics and traditional culture methods across key parameters relevant to complex samples.
Table 1: Overall Diagnostic Performance Metrics
| Parameter | CRISPR-based Diagnostics | Traditional Culture Methods | Experimental Context |
|---|---|---|---|
| Pooled Sensitivity | 99% (95% CI: 97–100%) [47] | Varies by pathogen and culture conditions | Meta-analysis of MRSA detection in clinical samples [47] |
| Pooled Specificity | 100% (95% CI: 99–100%) [47] | High (reference standard) | Meta-analysis of MRSA detection in clinical samples [47] |
| Median Detection Time | 60 minutes (IQR: 41.25–98.75 min) [47] | 2-3 days (48-72 hours) [47] [11] | Clinical sample validation [47] |
| Impact of EPS Matrix | Reduced efficiency; requires amplification or NP carriers [4] [13] | Physical barrier prevents nutrient penetration; protects embedded cells [13] [26] | Biofilm-associated infection models [13] [26] |
Table 2: Resilience to Sample Interference and Inhibitors
| Interference Type | Effect on CRISPR Diagnostics | Effect on Culture Methods | Mitigation Strategies |
|---|---|---|---|
| Clinical Inhibitors | Enzymatic activity fragile; 63% performance drop reported in high-humidity field conditions [4] | Can inhibit bacterial growth, leading to false negatives [48] | Sample purification, buffer optimization, lyophilized reagents [4] [11] |
| EPS Barrier | Limits access to target nucleic acids; reduces CRISPR component delivery [13] | Creates diffusion barrier, protecting sessile cells [26] [49] | Nanoparticle carriers (e.g., liposomal Cas9), EPS-disrupting enzymes [13] [49] |
| Humidity/Temperature | Field studies show significant performance drop (e.g., ~63%) in high humidity [4] | Generally robust if transport conditions are maintained | Reagent stabilization, portable device integration [4] |
Experimental Protocols for Assessing Sample Interference
Protocol: Evaluating CRISPR Inhibitors in Complex Clinical Samples
Objective: To quantify the effect of clinical sample inhibitors on CRISPR-Cas12a detection efficiency for Mycobacterium tuberculosis in sputum samples [48].
Materials:
- CRISPR Reagents: Cas12a enzyme, target-specific crRNA, fluorescent ssDNA reporter (e.g., FAM-TTATT-BHQ)
- Sample Types: Processed sputum samples from patients with suspected TB
- Control: Purified MTB DNA in nuclease-free water
- Equipment: Real-time fluorimeter or plate reader, thermal cycler
Procedure:
- Sample Processing: Digest and decontaminate sputum samples using N-acetyl-L-cysteine-NaOH method [48].
- DNA Extraction: Extract nucleic acids from processed samples using commercial kits.
- CRISPR Reaction Setup:
- Prepare master mix containing 50 nM Cas12a, 50 nM crRNA, 500 nM reporter probe in reaction buffer
- Divide master mix into two aliquots: one for clinical DNA extracts, one for purified MTB DNA controls
- Load samples into 96-well plate (n=3 replicates per sample)
- Signal Measurement:
- Incubate reactions at 37°C for 30 minutes
- Measure fluorescence every 2 minutes using a real-time fluorimeter
- Data Analysis:
- Calculate time-to-positive (TTP) for each reaction
- Compare TTP between clinical samples and purified DNA controls
- Determine inhibition ratio = (TTPsample - TTPcontrol)/TTP_control
Expected Outcomes: Inhibitor-rich samples will demonstrate prolonged TTP values and reduced endpoint fluorescence compared to purified controls, indicating interference with Cas enzyme activity [4] [48].
Protocol: Assessing EPS Impact on Antibiotic Penetration in Biofilms
Objective: To evaluate the barrier function of EPS against antibiotics in Pseudomonas aeruginosa biofilms and compare with CRISPR-based antimicrobial efficacy [13].
Materials:
- Bacterial Strain: P. aeruginosa reference strain (e.g., PAO1)
- Antimicrobials: Tobramycin (conventional antibiotic), Liposomal CRISPR-Cas9 targeting essential genes [13]
- Assessment Methods: Confocal laser scanning microscopy (CLSM), colony-forming unit (CFU) enumeration [49]
Procedure:
- Biofilm Growth:
- Culture biofilms on silicone coupons or in flow cells for 48-72 hours
- Use standard biofilm media with continuous nutrient supply
- Treatment Application:
- Apply tobramycin at clinically relevant concentrations (e.g., 100 µg/mL)
- Apply liposomal CRISPR-Cas9 formulations (e.g., targeting gyrA)
- Include untreated controls
- Penetration Assessment:
- For antibiotics: Use fluorescently labeled tobramycin and track penetration via CLSM
- For CRISPR: Use fluorescent reporters to monitor distribution
- Efficacy Measurement:
- Post-treatment, disrupt biofilms by sonication
- Serial dilute and plate for CFU enumeration
- Calculate log reduction compared to untreated controls
Expected Outcomes: Conventional antibiotics will show limited penetration and <2-log reduction in viable counts, while CRISPR-nanoparticle combinations may achieve >3-log reduction due to better penetration and targeted gene disruption [13].
Visualization of Mechanisms and Challenges
Core CRISPR Diagnostic Mechanism with Trans-Cleavage
Sample Interference in Diagnostic Pathways
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Research Reagent Solutions for Overcoming Sample Interference
| Reagent/Material | Function | Application Context |
|---|---|---|
| Recombinant Cas Proteins | Core enzyme for target recognition and trans-cleavage activity [4] [11] | CRISPR diagnostic assay development |
| Target-Specific crRNAs | Guide RNA molecules that confer specificity to pathogen nucleic acids [4] | Custom assay design for different biofilm pathogens |
| Fluorescent Reporter Probes | ssDNA/RNA molecules that release signal upon Cas-mediated cleavage [11] | Signal detection in CRISPR assays |
| Liposomal Nanoparticles | Delivery vehicles that enhance penetration through EPS barriers [13] | CRISPR component delivery in biofilm matrices |
| EPS-Disrupting Enzymes | DNase, dispersin B, or polysaccharide-degrading enzymes [49] | Sample pre-treatment to improve nucleic acid access |
| Inhibitor-Resistant Polymerases | Enzymes for amplification steps that function in inhibitor-rich samples [4] | Nucleic acid amplification prior to CRISPR detection |
| Lateral Flow Strips | Paper-based platforms for visual signal readout without instrumentation [11] | Point-of-care compatible detection |
| Microfluidic Chips | Integrated devices for "sample-to-result" processing [4] [49] | Automated handling of complex samples |
The comparative analysis presented in this guide demonstrates that both CRISPR diagnostics and traditional culture methods face significant challenges from sample interference, though the nature and impact of these challenges differ substantially. Culture methods remain vulnerable to the physical and chemical barriers created by EPS matrices, while CRISPR systems, despite their superior speed and sensitivity, show fragility in the presence of clinical inhibitors and complex sample matrices. The experimental data reveals that mitigation strategies must be tailored to the specific technology and interference type. Nanoparticle-enhanced delivery shows particular promise for overcoming EPS barriers in biofilm detection, while sample purification and reagent engineering offer pathways to address clinical inhibitors. For researchers selecting methodologies for biofilm pathogen detection, the decision matrix should incorporate sample type, required sensitivity, turnaround time needs, and the specific interference profiles expected in their target applications. The integration of CRISPR diagnostics with advanced sample preparation and delivery technologies represents the most promising path toward robust detection in complex sample environments.
The efficacy of any diagnostic or therapeutic agent is contingent upon its successful delivery to the target site. For pathogens residing within biofilms—structured communities of microbial cells encased in a self-produced extracellular polymeric substance (EPS)—this delivery represents a significant and complex challenge [50] [51]. The biofilm matrix is a formidable physical and chemical barrier, primarily composed of polysaccharides, proteins, and extracellular DNA (eDNA), which creates a protected microenvironment for the embedded cells [52] [53]. This matrix limits the penetration of antimicrobials and, as this analysis will explore, also hinders the delivery of modern molecular tools like CRISPR components.
This guide objectively compares the performance of emerging CRISPR-based diagnostic systems against traditional culture methods, with a specific focus on the critical hurdle of delivering detection machinery through dense biofilm structures. While culture methods rely on recovering planktonic (free-swimming) cells that have dispersed from the biofilm, CRISPR diagnostics aim to detect pathogens directly within the biofilm matrix, necessitating efficient penetration and targeting [4] [6]. The inherent properties of biofilms, including their heterogeneous architecture, charge interactions, and enzymatic activity, can adsorb, sequester, or degrade diagnostic components, drastically reducing detection sensitivity and reliability [52]. Understanding and comparing these delivery challenges is paramount for researchers and drug development professionals seeking to implement the most accurate and robust pathogen detection protocols for biofilm-associated infections.
Biofilm Structure and Its Impact on Molecule Delivery
The delivery of diagnostic components is impeded by the very nature of biofilm architecture and composition. A mature biofilm is not a uniform layer but a complex, three-dimensional structure characterized by microcolonies interspersed with fluid channels, creating gradients of nutrients, oxygen, and metabolic waste [52] [51]. This heterogeneous architecture presents a dual challenge: it creates physical diffusion barriers and induces phenotypic variations in the resident bacterial cells, including dormant "persister" cells that may alter the expression of the very genes targeted for detection [51].
The EPS matrix acts as a molecular sieve and an interactive scaffold. Key matrix components that hinder delivery include:
- Exopolysaccharides: Form a viscous gel that physically retards the diffusion of large molecular complexes, such as CRISPR ribonucleoproteins (RNPs) [50].
- Extracellular DNA (eDNA): Contributes to the structural integrity of the biofilm and can electrostatically bind to positively charged molecules, sequestering them before they reach their target [52].
- Proteins and Enzymes: May degrade protein-based CRISPR components like Cas enzymes, especially if proteases are present in the matrix or released by lysed cells [4].
Table 1: Key Biofilm Components and Their Impact on Delivery
| Biofilm Component | Primary Function | Impact on Diagnostic Molecule Delivery |
|---|---|---|
| Exopolysaccharides | Structural scaffolding, hydration retention | Increases viscosity, physically hinders diffusion of large complexes [52]. |
| Extracellular DNA (eDNA) | Matrix stability, cell adhesion, cation retention | Can bind electrostatically to diagnostic components, sequestering them [51]. |
| Proteins & Enzymes | Structural, enzymatic, adhesive roles | Surface enzymes may degrade protein-based tools like Cas enzymes [4]. |
| Microbial Cells | Metabolic activity, resistance | Dense, multi-layered cells act as the final barrier to target access [50]. |
Comparative Analysis: CRISPR Diagnostics vs. Culture Methods
The fundamental difference between CRISPR-based diagnostics and traditional culture methods lies in their approach to biofilm sampling and analysis, which directly influences their respective "delivery" challenges.
Traditional Culture Methods
Culture methods, such as the Colony Forming Unit (CFU) count, constitute the historical gold standard for biofilm quantification [50] [6]. These methods circumvent the biofilm delivery problem through a brute-force approach: biofilm disruption.
- Experimental Protocol (CFU Count): Mature biofilms are first physically disaggregated from their substrate using scraping, vortexing, or sonication. The resulting suspension is then homogenized and subjected to serial dilutions in a liquid medium. Each dilution is plated onto nutrient agar and incubated for 24-72 hours. The number of viable cells in the original biofilm is calculated based on the colony counts, considering the dilution factor and plating volume [50].
- Performance and Delivery Context: The strength of this method is that it bypasses the need for molecular penetration by breaking down the biofilm structure. However, this is also its weakness. The process is time-consuming, labor-intensive, and requires days to yield results. Furthermore, it only detects viable, culturable cells, potentially missing viable-but-non-culturable (VBNC) states or cells that remain embedded in resilient biofilm aggregates after disruption [6]. The method provides no spatial information about the original biofilm architecture.
CRISPR-Based Diagnostics
CRISPR diagnostics, such as SHERLOCK and DETECTR, represent a paradigm shift by aiming to detect pathogen-specific nucleic acid sequences directly within the sample, offering the potential for rapid, high-sensitivity results without the need for culture [4]. For biofilm diagnostics, this means the CRISPR machinery must be delivered to and access genetic material inside the biofilm matrix.
- Core Mechanism: These systems use a Cas nuclease (e.g., Cas12a, Cas13) complexed with a guide RNA (gRNA) programmed to recognize a specific bacterial DNA or RNA sequence. Upon target binding, the Cas enzyme exhibits collateral activity, non-specifically cleaving nearby reporter molecules (e.g., fluorescent probes), generating a detectable signal [4].
- Delivery Challenge: The large, often negatively charged, CRISPR ribonucleoprotein (RNP) complexes must diffuse through the oppositely charged EPS to reach intracellular bacterial DNA or RNA. This process is inefficient, leading to reduced sensitivity. Inhibitors in complex samples can also interfere with the enzymatic reaction, and a key limitation is the inability of these systems to distinguish between DNA from live and dead bacteria, which can lead to false positives [4].
Table 2: Performance Comparison: Culture vs. CRISPR Diagnostics for Biofilms
| Parameter | Culture Methods (e.g., CFU Count) | CRISPR Diagnostics (e.g., DETECTR) |
|---|---|---|
| Detection Time | 24 - 72 hours [50] | 15 minutes - 2 hours [4] |
| Key Delivery Hurdle | Physical disaggregation of biofilm matrix | Diffusion of RNP complexes through intact EPS [52] |
| Sensitivity | High for culturable cells (~1 cell) | Ultra-sensitive (theoretically aM levels) [4] |
| Specificity | High, based on growth phenotype | High, programmable via gRNA design [4] |
| Viable vs. Dead Cell | Distinguishes (only viable cells grow) | Cannot distinguish [4] |
| Spatial Information | None (destructive sampling) | Potential for spatial mapping with optimized delivery |
Experimental Protocols for Assessing Delivery and Efficacy
To objectively compare these technologies in a biofilm context, specific experimental protocols are required. Below are detailed methodologies for evaluating both traditional and CRISPR-based approaches.
Protocol 1: Standardized Biofilm Cultivation using Microtiter Plates
This is a foundational protocol for generating reproducible biofilms for downstream analysis [54].
- Inoculation: Prepare a suspension of the test microorganism in an appropriate growth medium, adjusted to an optical density (e.g., OD600 ≈ 0.1).
- Seeding: Dispense 100-200 µL of the bacterial suspension into the wells of a sterile 96-well flat-bottom microtiter plate. Include negative control wells containing sterile medium only.
- Adhesion & Growth: Incubate the plate under static conditions at the optimal growth temperature (e.g., 37°C) for a defined adhesion period (e.g., 2-4 hours).
- Washing: Gently invert the plate to remove non-adherent planktonic cells. Wash the biofilms twice with a sterile phosphate-buffered saline (PBS) by adding and removing PBS carefully to avoid disrupting adhered cells.
- Maturation: Add fresh medium to the washed wells and continue incubation for the desired maturation time (e.g., 24-48 hours), replacing the medium every 12-24 hours if necessary to maintain nutrient supply.
Protocol 2: Biofilm Disruption and Culture-Based Quantification (TCP Method)
The Tissue Culture Plate (TCP) method is a standard quantitative approach for assessing biofilm formation [6].
- Cultivate Biofilm: Grow biofilms in a microtiter plate as described in Protocol 1.
- Fixation and Staining: Remove the growth medium and wash the plate gently with PBS. Fix the biofilms by air-drying or using methanol. Add 0.1% crystal violet stain to each well and incubate for 15-20 minutes.
- Destaining and Quantification: Rinse the plate thoroughly with water to remove excess stain. Add a destaining solution (e.g., 30% acetic acid or ethanol-acetone mixture) to dissolve the crystal violet bound to the biofilm. Transfer the destained solution to a new plate and measure the optical density at 570-595 nm, which correlates with the total biofilm biomass [6].
- CFU Enumeration (Parallel Assay): In a parallel experiment, after growing and washing biofilms, add PBS and physically dislodge the biofilm cells by scraping, pipetting, or sonication. Serially dilute this suspension and plate on agar for CFU counting to determine the number of viable, culturable cells [50].
Protocol 3: Evaluating CRISPR Diagnostic Efficacy on Intact Biofilms
This protocol assesses the performance of a CRISPR assay when applied directly to an intact biofilm.
- Sample Preparation: Grow biofilms on a suitable substrate (e.g., glass coverslips placed in a well plate). After maturation, gently rinse to remove planktonic cells.
- Pre-treatment (Optional but Critical): To enhance CRISPR component delivery, apply a pre-treatment to the intact biofilm. This could be:
- Enzymatic: Incubate with matrix-degrading enzymes like DNase I (targets eDNA) or dispersin B (targets polysaccharides) [52].
- Chemical: Use chelating agents like EDTA to disrupt ionic interactions in the EPS.
- Physical: Apply mild sonication or vortexing with beads to partially disrupt the structure without fully homogenizing.
- CRISPR Reaction Application: Apply the complete CRISPR detection mix (containing Cas-gRNA RNP, reporter probes, and buffer) directly onto the intact, pre-treated biofilm.
- Incubation and Signal Detection: Incubate the reaction at the optimal temperature (e.g., 37°C) for a set time (e.g., 1 hour). Quantify the resulting fluorescent or colorimetric signal using a plate reader or lateral flow strip reader. Compare the signal intensity to that obtained from a homogenized biofilm sample to calculate the delivery efficiency loss due to the intact matrix.
The Scientist's Toolkit: Essential Research Reagents
The following table details key reagents and materials essential for conducting the experiments described in this guide.
Table 3: Research Reagent Solutions for Biofilm and CRISPR Delivery Research
| Reagent / Material | Function / Application | Example Use Case |
|---|---|---|
| 96-well Microtiter Plates | Platform for high-throughput, static biofilm cultivation and assay [54]. | Standardized biofilm growth for TCP assay or CRISPR testing. |
| Crystal Violet | A basic dye that binds to negatively charged surface molecules and polysaccharides in the biofilm matrix [6]. | Staining for quantitative assessment of total biofilm biomass. |
| Tryptic Soy Broth (TSB) | A nutrient-rich, general-purpose growth medium for cultivating a wide range of fastidious bacteria. | Promoting robust biofilm formation in organisms like staphylococci. |
| DNase I | Enzyme that degrades extracellular DNA (eDNA), a critical component of the biofilm matrix [52]. | Pre-treatment to disrupt biofilm integrity and enhance molecular penetration for CRISPR delivery. |
| Cas12a/Cas13 Nuclease | The core effector proteins for CRISPR diagnostics that provide target-specific recognition and collateral cleavage activity [4]. | Constituting the detection complex in SHERLOCK or DETECTR assays for pathogen nucleic acid detection. |
| Fluorescent Reporter Probes | Single-stranded DNA (for Cas12) or RNA (for Cas13) oligonucleotides linked to a fluorophore and quencher. | Signal generation in CRISPR assays; cleavage separates fluorophore from quencher, producing a fluorescent signal. |
| Lipid Nanoparticles (LNPs) | A delivery vehicle that can encapsulate CRISPR components to protect them and enhance cellular uptake and biofilm penetration [52]. | Co-delivery of Cas-gRNA RNP and reporters to improve delivery efficiency into dense biofilm structures. |
Visualizing Workflows and Challenges
The following diagrams illustrate the core concepts and experimental workflows discussed in this guide.
Conceptual Workflow for CRISPR vs. Culture-Based Biofilm Detection
Biofilm Matrix Components as Delivery Barriers
The accurate detection and identification of pathogens within complex microbial communities, such as biofilms, represents a significant challenge in clinical microbiology and drug development. Biofilms, which are structured communities of microorganisms embedded in a self-produced extracellular polymeric substance (EPS) matrix, create particularly difficult environments for diagnostic assays due to their heterogeneous structure and diverse microbial constituents [55] [13]. The diagnostic landscape has long been dominated by culture-based methods, but the emergence of CRISPR-based diagnostics offers promising alternatives with potential for enhanced speed and precision. However, ensuring specificity in these complex environments remains a critical hurdle.
The imperative to minimize off-target effects—where diagnostic assays detect non-target organisms or genetic elements—is particularly acute in biofilm pathogen detection. False positives can lead to inappropriate treatment strategies, while false negatives may leave dangerous pathogens undetected. This guide provides a systematic comparison of CRISPR diagnostics and traditional culture methods, focusing specifically on strategies to enhance specificity when working with intricate microbial communities. We present experimental data, detailed protocols, and analytical frameworks to help researchers and drug development professionals select and optimize appropriate detection methodologies for their specific applications.
Comparative Performance Analysis: CRISPR Diagnostics vs. Culture Methods
The selection between CRISPR-based diagnostics and traditional culture methods requires careful consideration of multiple performance parameters, particularly when detecting pathogens within biofilms. The table below summarizes key comparative metrics based on current experimental evidence.
Table 1: Performance Comparison for Biofilm Pathogen Detection
| Performance Parameter | CRISPR Diagnostics | Culture Methods |
|---|---|---|
| Analytical Specificity | 99-100% for targeted sequences [47] | High but limited to cultivable organisms |
| Sensitivity (LOD) | ~1-10 copies/μL with pre-amplification [11] | 10⁴-10⁵ CFU/g in tissue [9] |
| Time to Result | 30-100 minutes [47] | 2-14 days [11] |
| Sample Processing Needs | Complex (extraction, amplification) | Simple (homogenization) |
| Impact of Bacterial Viability | Detects DNA/RNA regardless of viability | Requires viable, culturable cells |
| Influence of Biofilm Matrix | High (inhibits reagent access) [13] | Moderate (affects colony dispersion) [9] |
| Quantification Capability | Semi-quantitative with standards | Quantitative (CFU counting) |
The high specificity of CRISPR systems stems from their dual recognition mechanism: the guide RNA (crRNA) must bind complementarily to the target nucleic acid, and the Cas protein must recognize a specific protospacer adjacent motif (PAM) sequence [4] [11]. This dual requirement significantly reduces the probability of non-specific detection events. However, the protective EPS matrix of biofilms can impede the access of CRISPR components to their targets, potentially reducing effective specificity in complex samples [13].
Culture methods, while historically considered the "gold standard," face different specificity challenges. Their principal limitation lies in the inability to detect viable but non-culturable (VBNC) pathogens and the potential for overgrowth of fast-growing species obscuring slow-growing pathogens [9]. The aggregation of bacteria in biofilms further complicates cultural detection, as the probability of sampling representative aggregates decreases significantly with increasing aggregate size [9].
Molecular Mechanisms and Specificity Enhancement Strategies
CRISPR Molecular Basis for Specific Target Recognition
The exceptional specificity of CRISPR diagnostics originates from its natural function as an adaptive bacterial immune system. The core mechanism involves CRISPR RNA (crRNA) that guides Cas proteins to complementary nucleic acid sequences through base pairing [4]. Upon target recognition, Cas proteins undergo conformational changes that activate their nuclease activities, including both cis-cleavage (of target nucleic acids) and trans-cleavage (non-specific cleavage of reporter molecules) [4] [11].
Different Cas proteins employ distinct mechanisms to ensure precise target recognition:
- Cas9: Requires both crRNA complementarity and a specific PAM sequence, then creates double-strand breaks in target DNA [11]
- Cas12: After recognizing target DNA with a T-rich PAM, exhibits trans-cleavage activity against single-stranded DNA reporters [4]
- Cas13: Binds to target RNA and activates collateral cleavage of surrounding RNA molecules [11]
This multi-layered recognition system provides the foundation for high-fidelity detection, but requires optimization to maintain specificity in complex samples like biofilms that may contain numerous similar genetic sequences.
Experimental Workflow for Specificity Validation
Table 2: Key Reagents for Specificity Assessment Experiments
| Research Reagent | Function in Specificity Assessment |
|---|---|
| Synthetic crRNA | Guides Cas complex to target sequence; design critical for specificity |
| Cas Protein (Cas12a/Cas13) | Executes target cleavage and collateral activity |
| Fluorescent Reporter Probes | Trans-cleavage substrate for signal generation |
| Non-target Nucleic Acids | Negative controls for off-target evaluation |
| PAM Variant Libraries | Validate PAM requirement stringency |
| Biofilm Disruption Enzymes | (e.g., DNase, proteinase) Improve reagent access to targets |
The following experimental protocol systematically evaluates detection specificity in biofilm models:
Protocol: Specificity Validation in Synthetic Microbial Communities
Sample Preparation:
- Create defined microbial communities mixing target pathogen with phylogenetically similar non-target species
- Grow mono- and multi-species biofilms using CDC biofilm reactors or drip-flow reactors
- Homogenize biofilm samples using mechanical (bead beating) and enzymatic (DNase, proteinase K) methods [9]
Nucleic Acid Extraction:
- Extract DNA/RNA using kits with demonstrated efficiency for biofilm samples
- Include steps to remove PCR inhibitors common in biofilm matrices
Assay Configuration:
- Design crRNAs with comprehensive off-target prediction using specialized algorithms
- Include mismatch-containing guides to evaluate tolerance to sequence variations
- Implement temperature optimization (often 37-42°C) to enhance discrimination [11]
Specificity Assessment:
- Test against genomic DNA from closely related non-target species
- Evaluate cross-reactivity with commensal strains likely present in sample type
- Use dilution series to determine limit of specific detection
Signal Detection:
- Employ fluorescent or lateral flow readouts with appropriate controls
- Compare to culture results as reference standard [47]
This systematic approach enables researchers to quantify both analytical and functional specificity before deploying assays on clinical or environmental samples.
Visualization of Specificity Assurance Workflows
CRISPR Diagnostic Specificity Assurance Pathway
Comparative Experimental Design for Method Evaluation
Integrated Strategies for Enhanced Specificity
crRNA Design and Bioinformatic Approaches
Sophisticated crRNA design represents the foremost strategy for minimizing off-target effects in CRISPR diagnostics. Current evidence indicates that crRNAs with specific thermodynamic properties and minimal similarity to non-target sequences significantly reduce false positives [4]. Bioinformatic tools now enable researchers to:
- Scan entire genomes of non-target species likely present in biofilms to identify potential cross-reactive sequences
- Incorporate modified nucleotides in crRNA to enhance binding specificity
- Design crRNAs targeting unique genomic regions with minimal conservation across species
Experimental data from MRSA detection studies demonstrates that carefully designed crRNAs can achieve 100% specificity in clinical samples, outperforming even PCR-based methods [47]. This high specificity is maintained even in complex samples because the CRISPR system requires precise sequence recognition combined with PAM identification, creating two independent specificity checkpoints.
Sample Processing and Amplification Strategies
The biofilm matrix itself presents a significant barrier to specificity by physically blocking access to targets and harboring PCR inhibitors that can cause assay failure. Integrated processing strategies include:
- Enzymatic matrix disruption: Using DNase, proteinase K, or specific polysaccharide-degrading enzymes to dissolve EPS without damaging target cells [13]
- Mechanical homogenization: Bead beating or sonication to disperse bacterial aggregates that cause heterogeneous distribution [9]
- Inhibitor removal agents: Incorporating BSA, betaine, or specialized commercial reagents to neutralize common biofilm-derived inhibitors
For amplification-based CRISPR detection (e.g., combining RPA or LAMP with Cas12/Cas13), target amplification can be designed to include specific sequence tags that are only recognized by the CRISPR system after successful amplification, adding an additional specificity layer [11]. This approach has demonstrated the ability to detect as few as 1 copy of target DNA in biofilm samples while maintaining perfect specificity against non-target species [47].
The comparison between CRISPR diagnostics and traditional culture methods for biofilm pathogen detection reveals a complex specificity landscape. CRISPR technologies offer superior analytical specificity through their molecular recognition mechanisms, with recent meta-analyses reporting 99-100% specificity for MRSA detection in clinical samples [47]. Culture methods, while potentially less affected by biofilm matrix interference, suffer from biological limitations including inability to detect VBNC states and sampling challenges posed by bacterial aggregation [9].
The integration of multiple specificity enhancement strategies—including sophisticated crRNA design, optimized sample processing, and stringent reaction conditions—enables researchers to overcome many challenges associated with complex microbial communities. Future directions point toward amplification-free CRISPR detection systems that eliminate amplification bias [11], integrated microfluidic devices for automated processing [56], and machine learning algorithms for predictive off-target identification.
For researchers and drug development professionals, the selection between these methodologies should be guided by specific application requirements. CRISPR diagnostics provide rapid, specific identification ideal for time-sensitive applications and targeted therapeutic decisions. Culture methods remain valuable for comprehensive pathogen recovery when broad detection is prioritized over speed. Through strategic implementation and continuous refinement of specificity assurance protocols, both approaches can yield highly reliable results for biofilm pathogen detection in even the most complex microbial communities.
The transition of diagnostic technologies from controlled laboratory environments to real-world field conditions presents a significant challenge, particularly for enzyme-dependent systems. CRISPR-driven diagnostics have emerged as a transformative tool for detecting biofilm-forming pathogens, offering unprecedented sensitivity and specificity [4]. However, their performance in non-ideal conditions remains a critical hurdle for widespread adoption in point-of-care testing and resource-limited settings [11]. This comparison guide objectively evaluates the environmental adaptability of CRISPR diagnostics against traditional culture methods for detecting biofilm-associated pathogens, providing experimental data and methodologies to inform researcher selection and implementation.
The fundamental challenge lies in maintaining the enzymatic activity of Cas proteins (Cas9, Cas12, Cas13) when exposed to field conditions such as temperature fluctuations, humidity variations, and inhibitory substances present in complex samples [4]. While culture methods have established reliability but extended turnaround times, CRISPR diagnostics offer rapid results but with potential vulnerability to environmental stressors. Understanding these trade-offs is essential for researchers developing diagnostic solutions for biofilm-related infections, which account for approximately 60-80% of microbial infections in humans [8].
Molecular Mechanisms and Environmental Vulnerabilities
Core Mechanisms of CRISPR Diagnostics
CRISPR diagnostics utilize Cas proteins' unique enzymatic properties for pathogen detection through two principal activities: target recognition via crRNA complementary pairing and enzymatic activation leading to trans-cleavage of reporter molecules [4] [11]. The system's programmability allows researchers to design specific crRNAs targeting conserved regions of pathogen nucleic acids, including bacterial 16S rRNA genes or drug-resistant genes [4]. Upon target recognition, Cas proteins undergo conformational changes that activate their nuclease activities. Cas12 exhibits collateral cleavage of single-stranded DNA upon target DNA recognition, while Cas13 trans-cleaves single-stranded RNA after binding target RNA [11]. This trans-cleavage property enables signal amplification through fluorescent reporter probes that release detectable signals when cleaved by activated Cas proteins.
Table 1: Key Cas Enzymes in CRISPR Diagnostics and Their Environmental Considerations
| Cas Enzyme | Nucleic Acid Target | Trans-Cleavage Activity | Key Environmental Vulnerabilities | Optimal Temperature Range |
|---|---|---|---|---|
| Cas9 | DNA | None | Less affected by RNA degradation | 37°C [11] |
| Cas12a | DNA | ssDNA | Sensitive to sample inhibitors | 37°C [11] |
| Cas13a | RNA | ssRNA | RNA instability in field conditions | 37-40°C [11] |
| Cas14 | DNA | ssDNA | High humidity performance drop [4] | 37°C [11] |
Biofilm Complexity and Detection Challenges
Biofilms represent structured microbial communities encapsulated within an extracellular polymeric substance (EPS) matrix, comprising polysaccharides, proteins, lipids, and extracellular DNA [55] [57]. This complex matrix presents substantial barriers to effective pathogen detection through both conventional and molecular methods. The EPS provides physical protection to embedded microorganisms, enhances resistance to antimicrobial agents, and impedes nucleic acid extraction efficiency—a critical preliminary step for CRISPR-based detection [55].
The developmental cycle of biofilms progresses through initial reversible attachment, irreversible attachment, maturation, and dispersion phases [55]. Each stage exhibits distinct biochemical compositions and cellular activities that influence detection efficacy. Of particular concern are biofilms formed by ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species), which demonstrate heightened antimicrobial resistance and present significant treatment challenges [55]. This resistance profile complicates both culture-based identification and molecular detection methods.
Diagram: Environmental Impact on Diagnostic Performance
Comparative Performance Analysis
Direct Performance Metrics in Controlled Conditions
Under optimal laboratory conditions, CRISPR diagnostics demonstrate significant advantages over traditional culture methods for biofilm pathogen detection, particularly in processing time and sensitivity. The integration of CRISPR with isothermal amplification techniques like RPA and LAMP enables detection sensitivities as low as 1 copy/μl for specific pathogens within 30-60 minutes [11]. This stands in stark contrast to culture methods, which typically require 2-10 days for reliable pathogen identification due to the extended incubation periods necessary for microbial growth [11] [58].
Table 2: Direct Performance Comparison: CRISPR vs. Culture Methods
| Performance Metric | CRISPR Diagnostics | Traditional Culture Methods | Experimental Support |
|---|---|---|---|
| Detection Time | 30 min - 2 hours [11] [58] | 2-10 days [11] | SHERLOCK, DETECTR platforms [4] |
| Sensitivity | aM (attomolar) levels [4] | Varies with pathogen load | 1 copy/μl detection for Mpox virus [11] |
| Specificity | High (programmable) [4] | High for culturable organisms | 100% specificity in SHERLOCK [11] |
| Sample Volume | μl range [58] | ml range required | Smart centrifugation processing [58] |
| Pathogen Identification | Specific sequences | Viable organisms only | 16S rRNA targeting [4] |
The specificity of CRISPR systems approaches 100% for well-designed assays, as demonstrated in the SHERLOCK platform, which leverages Cas13's RNA targeting capabilities [11]. Culture methods, while highly specific for culturable organisms, fail to detect viable but non-culturable (VBNC) pathogens or those with fastidious growth requirements—a particular limitation for biofilm-associated bacteria, which often enter dormant states [58].
Performance in Non-Ideal Field Conditions
Field deployment introduces environmental variables that substantially impact diagnostic performance. CRISPR systems exhibit notable vulnerability to field conditions, with studies reporting up to 63% performance reduction for Cas14-based assays under high humidity conditions in sub-Saharan Africa [4]. Enzymatic activity of Cas proteins proves sensitive to temperature fluctuations, inhibitory substances in complex samples, and variations in pH—all common challenges in point-of-care settings.
Culture methods maintain more consistent performance across diverse environmental conditions, though they remain susceptible to temperature variations during sample transport that affect bacterial viability. The requirement for consistent refrigeration for culture media and reagents presents logistical challenges in resource-limited settings, whereas lyophilized CRISPR reagents offer improved stability but still demonstrate enzymatic vulnerability to environmental stressors [4].
Novel approaches to biofilm detection highlight alternative methodologies that may offer enhanced environmental stability. Near-infrared (NIR) spectroscopy has demonstrated 80-93% accuracy in identifying S. aureus biofilms on glass surfaces, representing a non-enzymatic approach potentially less susceptible to environmental variables [59]. Similarly, microfluidic bacterial trapping combined with deep learning detection achieves culture-free identification within 2 hours with clinically relevant sensitivity as low as 1-10 CFU/ml for specific pathogens [58].
Experimental Protocols for Environmental Adaptability Assessment
Protocol: Evaluating CRISPR Enzyme Stability Under Temperature Variations
Objective: To quantify the effect of temperature fluctuations on Cas enzyme activity and detection sensitivity in CRISPR assays.
Materials:
- Cas12a enzyme (commercial source)
- Custom crRNA targeting bacterial 16S rRNA gene
- Fluorescent ssDNA reporter (FAM-TTATT-BHQ1)
- Nucleic acid amplification reagents (RPA or LAMP)
- Real-time PCR instrument or plate reader
- Temperature-controlled heating blocks
Methodology:
- Prepare Cas12a-crRNA complexes according to manufacturer specifications
- Aliquot identical reaction mixtures into separate tubes
- Pre-incubate aliquots at temperatures ranging from 4°C to 45°C for 30 minutes
- Add target DNA (serial dilutions of pathogen genomic DNA)
- Transfer all reactions to a single isothermal temperature (37°C) for amplification and detection
- Measure fluorescence every 30 seconds for 60 minutes
- Calculate reaction efficiency based on time to positivity and endpoint fluorescence
Data Analysis: Compare time to detection and fluorescence intensity across temperature pretreatment groups. Establish the temperature stability profile for the specific Cas enzyme formulation [4] [11].
Protocol: Assessing Humidty Impact on Lyophilized CRISPR Reagents
Objective: To determine the stability of lyophilized CRISPR reagents under high humidity conditions simulating field use.
Materials:
- Lyophilized CRISPR reaction pellets (Cas enzyme, crRNA, reporters)
- Desiccators with saturated salt solutions for humidity control
- Target pathogen DNA
- Fluorescence detection system
Methodology:
- Place lyophilized CRISPR pellets in controlled humidity environments (30%, 60%, 80%, 95% RH) using saturated salt solutions
- Maintain at constant temperature (25°C) for predetermined durations (1, 3, 7, 14 days)
- Reconstitute pellets with nuclease-free water containing target DNA
- Incubate at 37°C and monitor fluorescent signal development
- Compare performance to freshly prepared reagents and refrigerated controls
Data Analysis: Calculate percentage activity retention based on time to detection threshold compared to controls. Establish maximum humidity tolerance and shelf-life under various conditions [4].
Protocol: Culture Method Reliability Under Field Temperature Conditions
Objective: To evaluate bacterial recovery efficiency from biofilms after exposure to temperature variations simulating field transport.
Materials:
- Biofilm samples (formed on relevant surfaces)
- Transport media (commercial and homemade)
- Temperature logging equipment
- Culture media (TSBg, PCA)
- Incubators
Methodology:
- Establish standardized biofilms using reference strains (e.g., S. aureus, E. coli)
- Expose biofilm samples to temperature profiles simulating field conditions
- Process samples using both direct plating and enrichment protocols
- Compare colony counts with optimally stored controls
- Document time to visible growth for each condition
Data Analysis: Calculate percent recovery compared to optimal conditions. Establish correlation between temperature exposure and culture reliability [59] [60].
Research Reagent Solutions for Enhanced Environmental Adaptability
Table 3: Essential Research Reagents for Environmental Adaptation Studies
| Reagent Category | Specific Examples | Function in Environmental Adaptation | Implementation Considerations |
|---|---|---|---|
| Stabilized Cas Enzymes | Lyophilized Cas12a, Cas13a | Maintains enzymatic activity in temperature fluctuations [4] | Pre-qualify lot-to-lot consistency |
| crRNA Formulations | Modified nucleotides, 2'-O-methyl | Enhances nucleic acid guide stability in suboptimal conditions | Balance stability with binding affinity |
| Sample Preparation Tools | Selective lysis solutions [58] | Removes PCR inhibitors while preserving target integrity | Optimize for specific sample matrices |
| Portable Detection Platforms | Lateral flow strips, handheld fluorometers [11] | Enables readout in resource-limited settings | Match detection method to application needs |
| Environmental Monitors | Temperature/humidity data loggers | Quantifies environmental exposure during testing | Essential for correlating conditions with performance |
Diagram: Diagnostic Workflow Comparison
The environmental adaptability of diagnostic platforms represents a critical factor in their successful deployment for biofilm pathogen detection. CRISPR diagnostics offer compelling advantages in speed, sensitivity, and programmability but demonstrate significant vulnerability to environmental stressors like temperature fluctuations and high humidity. Traditional culture methods provide established reliability across diverse conditions but require extended time-to-result, complicating rapid clinical decision-making.
Future directions should focus on enhancing CRISPR system resilience through improved reagent formulation, environmental hardening of enzymatic components, and integration with sample preparation methods that mitigate inhibitor effects. Simultaneously, culture methods continue to evolve with rapid protocols and optimized transport systems that maintain their environmental robustness while reducing processing time. The optimal solution may lie in strategic application of both technologies—employing CRISPR for rapid screening when environmental conditions can be controlled, and utilizing culture methods for confirmation and susceptibility testing in challenging field environments.
Researchers should carefully consider their specific use case constraints, including environmental conditions, available infrastructure, and required time-to-result when selecting between these diagnostic approaches. As both technologies continue to advance, their complementary strengths offer promising pathways for comprehensive biofilm pathogen detection across diverse environmental contexts.
Biosafety and Ethical Considerations for Widespread Clinical and Industrial Deployment
The detection and management of biofilm-forming pathogens represent a significant challenge in both clinical and industrial settings. Biofilms are structured communities of microorganisms embedded in a self-produced extracellular polymeric substance (EPS), which confers inherent resistance to antimicrobial treatments and complicates detection [23] [55]. This comparison guide objectively evaluates the performance of emerging CRISPR-based diagnostics against traditional culture methods for detecting biofilm-associated pathogens, with particular attention to the biosafety and ethical considerations essential for their responsible deployment.
Traditional culture methods, long considered the gold standard, face limitations in speed, sensitivity, and the ability to detect viable but non-culturable organisms within biofilms [23]. CRISPR-Cas systems offer a transformative approach with their programmable nucleic acid detection capabilities, enabling precise identification of pathogens and their resistance genes [4] [11]. As these technologies transition from research to widespread application, a thorough understanding of their performance characteristics alongside appropriate biosafety frameworks becomes paramount for researchers, scientists, and drug development professionals.
Performance Comparison: CRISPR Diagnostics vs. Culture Methods
The following tables provide a quantitative comparison of CRISPR-based diagnostics and traditional culture methods across key performance metrics, based on current experimental data from the literature.
Table 1: Overall Performance Metrics for Pathogen Detection in Biofilms
| Performance Parameter | CRISPR-Based Diagnostics | Traditional Culture Methods | Experimental Support |
|---|---|---|---|
| Detection Time | 30–60 minutes [4] [61] | 2–10 days [23] [11] | Clinical sample validation [4] |
| Analytical Sensitivity | Attomolar (aM) level [4]; As low as 1 copy/μL for specific targets [11] | Limited by microbial growth and viability [23] | Comparison with qPCR standards [11] |
| Specificity | Single-base discrimination possible [16] | High for viable, culturable organisms [23] | 100% sensitivity and specificity shown in SHERLOCK platform [11] |
| Point-of-Care Suitability | High (lyophilized reagents, lateral flow readout) [4] | Low (requires lab infrastructure) [4] | Integration into portable devices [61] |
| Identification of Resistance Markers | Direct detection of resistance genes (e.g., bla, mecA) [13] | Requires subsequent susceptibility testing [23] | Meta-analysis of CRISPR-Cas studies [13] |
| Biofilm Penetration Capability | Detects nucleic acids from throughout biofilm structure [16] | Primarily detects surface-colonizing organisms [23] | Studies on food-contact surfaces [16] |
Table 2: Biofilm Detection Efficacy from Recent Experimental Studies
| Study Focus | CRISPR Performance | Culture Method Performance | Context and Notes |
|---|---|---|---|
| Catheter-Associated UTIs | Not specifically reported | Microplate assay detected biofilm in 88.6% of catheter isolates [23] | Reference standard for phenotypic detection [23] |
| Food Processing Surface Monitoring | Up to ~3-log target reduction with CRISPR-guided antimicrobials [16] | Often fails to prevent biofilm recurrence [16] | Precision suppression of pathogens in multispecies biofilms [16] |
| Pseudomonas aeruginosa Biofilm Eradication | Liposomal Cas9 formulations reduced biofilm biomass by >90% in vitro [13] | Conventional therapies often ineffective [13] | Nanoparticle-enhanced delivery [13] |
| Antimicrobial Resistance Gene Editing | Gold nanoparticle-CRISPR hybrids showed 3.5-fold increase in editing efficiency [13] | Not applicable | Resensitizing bacteria to antibiotics [13] |
Experimental Protocols and Methodologies
CRISPR-Based Diagnostic Workflow (e.g., DETECTR/SHERLOCK Systems)
The following protocol outlines the standard two-step process for CRISPR-based detection of biofilm pathogens, typically achieving results within 30-60 minutes [4] [11].
Sample Preparation and Nucleic Acid Extraction:
- Biofilm Disruption: Resuspend biofilm samples (e.g., from catheter tips or food-contact surfaces) in sterile saline and vortex vigorously with glass beads for mechanical disruption [23]. Alternatively, employ enzymatic treatment with proteinase K or DNase I to degrade the extracellular polymeric substance without damaging target nucleic acids [16].
- Nucleic Acid Extraction: Extract DNA/RNA using commercial kits. For comprehensive pathogen identification, co-extract both DNA and RNA. Purified nucleic acids can be used immediately or stored at -80°C [11].
Nucleic Acid Amplification (Pre-Amplification Step):
- Employ isothermal amplification methods such as Recombinase Polymerase Amplification (RPA) or Loop-Mediated Isothermal Amplification (LAMP) to enhance detection sensitivity. These techniques amplify target sequences at constant temperatures (e.g., 37-42°C for 15-20 minutes) without requiring thermal cyclers [11].
- For DNA targets (e.g., bacterial genomic DNA), use RPA. For RNA targets (e.g., viral genomes), include a reverse transcription step in the RPA reaction [61].
CRISPR-Cas Detection and Readout:
- Cas12a-based detection (DETECTR for DNA targets): Prepare a reaction mix containing LbCas12a or AsCas12a enzyme, specific crRNA designed to target pathogen DNA, and a single-stranded DNA (ssDNA) reporter molecule labeled with a fluorophore-quencher pair. Upon target recognition and Cas12a activation, trans-cleavage of the reporter releases fluorescence, measurable via a portable fluorimeter [61] [25].
- Cas13a-based detection (SHERLOCK for RNA targets): Use LbuCas13a enzyme with specific crRNA and an ssRNA reporter. Activated Cas13a cleaves the reporter upon binding target RNA (e.g., bacterial messenger RNA or viral RNA), generating a fluorescent signal [61] [25].
- Lateral Flow Readout: For equipment-free detection, use reporters labeled with FAM and biotin. Apply the reaction mixture to a lateral flow strip. Cleaved reporters generate a visible test line, with results interpretable within 5-10 minutes [11].
Traditional Culture Method for Biofilm Detection
This protocol details the reference standard microplate (Tissue Culture Plate) method for phenotypic detection of biofilm-forming pathogens, typically requiring 24-48 hours [23].
Sample Collection and Processing:
- Biofilm Sampling: Aseptically collect biofilm samples (e.g., catheter segments, industrial surface swabs). For catheter tips, cut 2-3 cm segments and rinse with sterile distilled water to remove loosely attached cells [23].
- Microbial Enrichment: Incubate samples in nutrient broth (e.g., Trypticase Soy Broth with 1% glucose) at 37°C for 24 hours to enrich bacterial growth [23].
Biofilm Detection via Tissue Culture Plate Method (TCPM):
- Inoculation: Transfer 20 μL of enriched bacterial suspension into sterile 96-well flat-bottom polystyrene tissue culture plates containing 180 μL of sterile broth per well. Include positive (known biofilm-forming strain) and negative (sterile broth) controls [23].
- Incubation and Washing: Incubate plates at 37°C for 24 hours. Gently shake plates, discard contents, and wash each well 3-4 times with sterile distilled water to remove planktonic cells. Invert plates to dry [23].
- Fixation and Staining: Add 200 μL of 2% sodium acetate to each well for 30 minutes for fixation. Wash again and stain with 200 μL of 0.1% crystal violet for 15 minutes. Repeat washing and air-dry plates [23].
- Quantification: Add 200 μL of 95% ethanol to destain wells. Measure optical density (OD) at 570-595 nm using a microplate reader. Interpret results: OD < ODC = non-biofilm former; ODC < OD < 2×ODC = weak; 2×ODC < OD < 4×ODC = moderate; OD > 4×ODC = strong biofilm former (ODC = OD of negative control) [23].
Supplementary Phenotypic Methods:
- Tube Method (TM): Inoculate 5 mL of broth with test organism, incubate at 37°C for 24-48 hours. Stain visible film lining tube wall with crystal violet. Qualitative assessment of biofilm formation [23].
- Congo Red Agar (CRA) Method: Culture isolates on Congo Red Agar plates, incubate at 37°C for 24-48 hours. Black colonies with dry crystalline consistency indicate biofilm production [23].
Biosafety and Ethical Considerations for Deployment
The integration of CRISPR technologies into clinical and industrial workflows necessitates careful attention to biosafety and ethical implications. Key considerations include:
Dual-Use Research Concerns: CRISPR diagnostics pose potential dual-use risks, where the same technology enabling precise pathogen detection could potentially be misused to enhance pathogen virulence or develop biological weapons [61]. This necessitates robust institutional oversight and compliance with emerging NIH guidelines for research involving recombinant nucleic acid molecules [62].
Environmental Release and Horizontal Gene Transfer: Engineered CRISPR systems, particularly those deployed in environmental or industrial settings, carry potential risks of horizontal gene transfer to environmental microorganisms [61] [16]. This is particularly relevant for engineered probiotics designed as "living therapeutics" that might release CRISPR components into the environment [61].
Regulatory Frameworks and Transparency: Effective deployment requires robust global regulatory frameworks. Since June 2025, NIH guidelines have mandated increased transparency, requiring public posting of approved Institutional Biosafety Committee (IBC) meeting minutes and rosters [62]. This promotes accountability in the oversight of recombinant DNA research.
Delivery System Toxicity and Off-Target Effects: Nanoparticle carriers used to enhance CRISPR delivery in biofilm eradication strategies may present toxicity challenges [13]. Additionally, CRISPR nucleases can exhibit off-target effects, potentially cleaving non-target sequences with partial complementarity, which requires careful guide RNA design and validation [61] [25].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for CRISPR-Based Biofilm Detection and Control
| Reagent / Material | Function and Application | Specific Examples |
|---|---|---|
| Cas Enzymes | Core nucleases for target recognition and signal generation | Cas12a (LbCas12a, AsCas12a) for DNA detection; Cas13a (LbuCas13a) for RNA detection [25] |
| Guide RNAs (crRNA) | Programmable components conferring target specificity | crRNAs designed against conserved regions of pathogen genomes (e.g., 16S rRNA, virulence factors) [4] |
| Isothermal Amplification Kits | Pre-amplification to enhance detection sensitivity | RPA (TwistAmp) and LAMP kits for amplifying target nucleic acids at constant temperature [11] |
| Fluorescent Reporters | Signal generation in solution-based assays | ssDNA reporters (for Cas12) or ssRNA reporters (for Cas13) with fluorophore (FAM)/quencher (BHQ) pairs [25] |
| Lateral Flow Strips | Equipment-free visual readout | Strips with anti-FAM and control lines for detecting cleaved reporters [11] |
| Nanoparticle Delivery Systems | Enhancing CRISPR delivery to biofilm-embedded bacteria | Liposomal Cas9 formulations, gold nanoparticle-CRISPR hybrids [13] |
| Biofilm Disruption Reagents | Releasing nucleic acids from biofilm matrix | Proteinase K, DNase I, dispersin B, glass beads for mechanical disruption [23] [16] |
CRISPR-based diagnostics represent a paradigm shift in biofilm pathogen detection, offering unprecedented speed, specificity, and point-of-care applicability compared to traditional culture methods. While culture techniques remain valuable for phenotypic characterization and viability assessment, CRISPR technologies enable rapid identification of pathogens and their resistance markers with attomolar sensitivity [4] [11].
The integration of CRISPR with nanoparticle delivery systems further extends its potential from mere detection to targeted biofilm control, demonstrating >90% reduction in Pseudomonas aeruginosa biofilm biomass in vitro [13]. However, the widespread clinical and industrial deployment of these technologies must be guided by comprehensive biosafety assessments, ethical frameworks, and regulatory oversight to manage risks associated with environmental release, horizontal gene transfer, and potential dual-use applications [61] [62].
As the field advances, the convergence of CRISPR diagnostics with artificial intelligence for assay optimization and the development of universal diagnostic platforms will further enhance our capability to manage biofilm-associated infections, ultimately strengthening global health security and industrial safety [4] [16].
Data-Driven Comparison: Validating CRISPR Performance Against Gold Standards
This guide provides an objective comparison of CRISPR-based diagnostics against traditional culture methods and polymerase chain reaction (PCR) for the detection of pathogens, with a specific focus on challenging scenarios such as biofilm-associated infections. The performance data and experimental methodologies outlined below are synthesized from recent, peer-reviewed studies to serve as a practical reference for researchers and drug development professionals.
Performance Metrics at a Glance
The following table summarizes the core performance characteristics of CRISPR diagnostics against traditional methods. The data for CRISPR is largely derived from studies on detecting methicillin-resistant Staphylococcus aureus (MRSA), a common biofilm-forming pathogen [47].
Table 1: Comparative Performance of Diagnostic Methods for Pathogen Detection
| Metric | CRISPR-Based Diagnostics | Traditional Microbial Culture | PCR / qPCR |
|---|---|---|---|
| Sensitivity | 99% (95% CI: 97–100%) [47] | Variable; 50-80% for PJI (20-50% culture-negative despite clinical signs) [63] | 85-99% [63] [39] |
| Specificity | 100% (95% CI: 99–100%) [47] | ~100% when positive (but high false-negative rate) [63] | ~100% [63] |
| Speed | ~60 minutes (IQR: 41–99 min) [47]; as fast as 40 min [39] | 2-5 days (for bacteria); up to 10 days for some species [11] [12] | 1.5 to 4 hours (includes extraction, amplification, and analysis) [11] |
| Limit of Detection (LOD) | ~ 0.11 copies/μL [39]; 1.2 CFU/mL in serum [39] | 10^4 - 10^6 CFU/mL (for reliable detection in biofilms) [23] | 0.1 × 10^4 – 10^5 copies/mL [39] |
| Key Advantage | Rapid, ultra-sensitive, suitable for point-of-care use. | Determines microbial viability, provides isolates for antibiotic susceptibility testing (AST). | High-throughput, standardized, quantitative. |
| Key Limitation | Does not provide live isolates for AST. | Slow, low sensitivity, affected by prior antibiotic use. | Requires sophisticated thermocyclers, trained personnel. |
Experimental Protocols and Methodologies
Understanding the experimental workflows is crucial for interpreting the performance data presented above. This section details the standard protocols for each diagnostic method.
CRISPR-Based Detection Workflow
Advanced CRISPR diagnostics often integrate isothermal amplification and collateral cleavage activity for high sensitivity. The following diagram illustrates the workflow of a typical two-step amplification-based CRISPR assay, such as SHERLOCK or DETECTR [11].
Experimental Protocol: Two-Step Amplification-Based CRISPR Assay [11] [12]
- Step 1: Nucleic Acid Extraction. Total DNA or RNA is extracted from the sample. For biofilm samples, this may require an initial step to disrupt the extracellular polymeric substance (EPS) matrix. For bacterial cells, lysis is performed to release genomic DNA [39].
- Step 2: Isothermal Amplification. The extracted nucleic acid is amplified using techniques like Recombinase Polymerase Amplification (RPA) or Loop-Mediated Isothermal Amplification (LAMP) at a constant temperature (e.g., 37-42°C). This step replaces the thermal cycling of PCR and rapidly increases the copy number of the target pathogen gene (e.g., mecA for MRSA) [11] [47].
- Step 3: CRISPR-Cas Detection.
- The amplified product is mixed with a pre-assembled ribonucleoprotein (RNP) complex, which contains the Cas enzyme (e.g., Cas12 for DNA, Cas13 for RNA) and a sequence-specific guide RNA (crRNA).
- If the target sequence is present, the RNP complex binds to it, activating the Cas enzyme's "collateral cleavage" or "trans-cleavage" activity.
- The activated Cas enzyme non-specifically cleaves nearby reporter molecules (e.g., fluorescently quenched single-stranded DNA for Cas12) [4] [25].
- Step 4: Signal Readout. The cleavage of the reporter molecule generates a detectable signal. This can be measured visually using lateral flow strips or quantified via a fluorometer. Positive results are indicated by a fluorescent signal or a test line on a strip [11].
Traditional Culture Method Workflow
Culture remains the "gold standard" for its ability to determine viability and provide isolates for antimicrobial susceptibility testing (AST), but it is slow and suffers from low sensitivity in biofilm-related infections [63] [23].
Experimental Protocol: Microbial Culture for Biofilm-Associated Pathogens [23]
- Step 1: Sample Collection and Processing. Clinical samples (e.g., catheter tips, periprosthetic tissue) are collected aseptically. For catheter tips, a common protocol involves rinsing with sterile water, disinfecting the surface with 70% ethanol to remove planktonic cells, and then cutting the tip into fragments [23].
- Step 2: Enrichment and Culture. The fragments are incubated in a nutrient broth (enrichment step) for 24 hours at 37°C. After enrichment, the broth is streaked onto selective and non-selective agar plates (e.g., blood agar, MacConkey agar) [23].
- Step 3: Incubation and Isolation. Agar plates are incubated aerobically or anaerobically at 37°C for 24-48 hours. For fastidious organisms or in cases of prior antibiotic use, incubation may be extended up to 10-14 days [63] [11].
- Step 4: Identification and AST. Colonies with distinct morphologies are sub-cultured for purity. Identification is performed using biochemical tests or MALDI-TOF mass spectrometry. Antibiotic susceptibility testing is then conducted using methods like Kirby-Bauer disk diffusion on Mueller-Hinton agar, following CLSI guidelines [23].
PCR/qPCR Method Workflow
PCR provides a balance of speed and sensitivity but requires laboratory infrastructure and cannot distinguish between live and dead cells [64].
Experimental Protocol: Quantitative PCR (qPCR) [64] [39]
- Step 1: Nucleic Acid Extraction. Similar to the CRISPR protocol, DNA/RNA is extracted from the sample.
- Step 2: Reaction Setup. The extracted nucleic acid is mixed with a master mix containing primers specific to the target pathogen gene, fluorescent probes (e.g., TaqMan), DNA polymerase, dNTPs, and buffer.
- Step 3: Thermal Cycling. The mixture is placed in a real-time PCR thermocycler, which subjects it to repeated cycles of denaturation (e.g., 95°C), annealing (e.g., 55-60°C), and extension (e.g., 72°C). The fluorescent signal increases proportionally with the amount of amplified DNA and is measured at the end of each cycle.
- Step 4: Data Analysis. The cycle threshold (Ct) value is determined, which represents the number of cycles required for the fluorescent signal to exceed a background level. This value is used for quantitative analysis.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for CRISPR-Based Pathogen Detection
| Reagent / Solution | Function in the Experiment | Specific Examples |
|---|---|---|
| Cas Proteins | The core enzyme that, when activated by target binding, cleaves reporter molecules to generate a signal. | Cas12a (targets DNA), Cas13a (targets RNA), CasΦ (compact size, high activity) [11] [39]. |
| Guide RNA (crRNA) | A short RNA sequence that is programmable and directs the Cas protein to the specific target nucleic acid sequence. | crRNA designed to target the mecA gene for MRSA detection or the 16S rRNA gene for bacterial identification [4] [47]. |
| Isothermal Amplification Kits | Enzymatic mixes that amplify target nucleic acid sequences at a constant temperature, eliminating the need for a thermocycler. | Recombinase Polymerase Amplification (RPA) kits, Loop-Mediated Isothermal Amplification (LAMP) kits [11] [47]. |
| Fluorescent Reporter Probes | Single-stranded DNA or RNA molecules labeled with a fluorophore and a quencher; cleavage produces a fluorescent signal. | ssDNA probes labeled with FAM (fluorophore) and BHQ (quencher) for use with Cas12 [25] [12]. |
| Lateral Flow Strips | A paper-based platform for visual detection of the assay result, ideal for point-of-care use. | Strips that capture labeled cleavage products, producing a visible test line [11]. |
| Lysis Buffers | Chemical solutions used to break open pathogen cells (and biofilms) to release nucleic acids for amplification and detection. | Solutions containing enzymes or detergents for microbial lysis; specific kits for DNA extraction from complex samples [39]. |
The quantitative data and protocols presented in this guide demonstrate that CRISPR diagnostics offer a paradigm shift in speed and sensitivity compared to culture and PCR. For researchers investigating biofilm-associated infections, where pathogen load can be low and culture results often negative, CRISPR platforms provide a powerful tool for rapid and precise etiological diagnosis [63] [39]. However, it is critical to note that culture remains indispensable for obtaining live isolates necessary for conducting antimicrobial susceptibility testing, a crucial step in guiding targeted therapy and antimicrobial stewardship [23]. The choice of diagnostic method should therefore be guided by the specific research question, weighing the need for speed and sensitivity against the requirement for a viable isolate.
Pseudomonas aeruginosa is a formidable Gram-negative opportunistic pathogen, notorious for causing severe infections in immunocompromised individuals, cystic fibrosis patients, and those with burn wounds [65]. Its significance in healthcare settings is magnified by its intrinsic and acquired antimicrobial resistance mechanisms, positioning it within the "ESKAPE" pathogens and on the WHO's "critical" priority list for new antibiotic development [65]. Conventional culture-based identification, while considered the gold standard, requires 48 to 72 hours to yield results, potentially delaying appropriate therapeutic intervention [65]. The organism's propensity to form biofilms further complicates treatment, as biofilm-associated bacteria can exhibit antibiotic tolerance up to 1,000 times higher than their planktonic counterparts, driving persistent chronic infections [66]. This diagnostic bottleneck underscores the urgent need for rapid, sensitive, and specific detection methods to enable early diagnosis and improve patient outcomes.
The emergence of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-based diagnostics represents a paradigm shift in molecular detection. This case study evaluates a specific CRISPR-driven assay—Cas12a-RCFL (Recombinase Polymerase Amplification followed by Cas12a-mediated detection via Fluorescence reader or Lateral flow biosensor)—for the rapid detection of P. aeruginosa [67]. We will objectively compare its performance against traditional culture methods and other molecular alternatives, framing this analysis within the broader thesis of evaluating novel diagnostics for biofilm-forming pathogens.
Technical Breakdown of the Cas12a-RCFL Platform
Core Principle: RPA Pre-amplification and CRISPR-Cas12a Detection
The Cas12a-RCFL method is a two-step assay that combines isothermal nucleic acid amplification with the sequence-specific collateral activity of the Cas12a enzyme [67].
- Step 1: Recombinase Polymerase Amplification (RPA). This isothermal amplification step occurs at a constant 42°C for 20 minutes. It rapidly multiplies the target gene sequence from the bacterial genomic DNA, eliminating the need for complex thermal cycling equipment required by PCR [67] [65].
- Step 2: CRISPR-Cas12a Detection. The RPA product is then introduced into a reaction containing the LbCas12a enzyme and a designed crRNA (CRISPR RNA). If the target DNA is present, the Cas12a-crRNA complex binds to it, activating the enzyme's "collateral cleavage" or trans-cleavage activity. This activity non-specifically degrades nearby single-stranded DNA (ssDNA) reporter probes, generating a detectable signal [67] [68].
Visualizing the Cas12a-RCFL Workflow
The diagram below illustrates the seamless integration of RPA and CRISPR-Cas12a steps in the detection workflow.
The Scientist's Toolkit: Essential Reagents for Cas12a-RCFL
Table 1: Key research reagents and their functions in the Cas12a-RCFL assay.
| Reagent / Component | Function in the Assay | Key Details |
|---|---|---|
| LbCas12a (Cpf1) Enzyme | CRISPR effector protein; provides sequence-specific binding and trans-cleavage activity. | Requires T-rich PAM sequence (5'-TTTV-3') for target recognition [68]. |
| crRNA (CRISPR RNA) | Guides Cas12a to the specific target DNA sequence. | Designed to be complementary to the oprL gene of P. aeruginosa; processed independently of tracrRNA [68] [65]. |
| RPA Primers | Initiate isothermal amplification of the target gene. | Targets the oprL gene, a highly specific marker for P. aeruginosa [65]. |
| ssDNA Reporter Probe | Substrate for trans-cleavage; signal generation. | For fluorescence: 5'-HEX/BHQ1-3' or 5'-FAM/BHQ1-3'. For lateral flow: 5'-FAM/Biotin-3' [67] [65]. |
| RPA Basic Kit | Provides enzymes and reagents for isothermal amplification. | Includes recombinase, polymerase, and single-stranded DNA-binding proteins [67]. |
| Lateral Flow Strip | For visual, instrument-free readout. | Typically contains a test line (anti-FAM antibody) and control line [67]. |
Experimental Protocol & Performance Validation
Detailed Methodology
The experimental validation of Cas12a-RCFL, as detailed in the research, follows a structured protocol [67]:
- Sample Preparation and DNA Extraction: Bacterial cultures or clinical swabs are processed using a rapid, heat-based DNA extraction method. Samples are heated at 100°C for 8 minutes in nucleic acid-free water, and the supernatant containing crude genomic DNA is collected after centrifugation.
- RPA Amplification: A 10 μL RPA reaction is assembled using a commercial kit. The reaction includes the extracted DNA template and primers targeting the oprL gene. The mixture is incubated at 42°C for 20 minutes.
- CRISPR-Cas12a Detection Setup:
- Fluorescence Assay: The detection mixture contains LbCas12a, oprL-specific crRNA, an RNase inhibitor, and a fluorescent ssDNA reporter (HEX-labeled with a BHQ1 quencher). Two microliters of the RPA product are added, and fluorescence intensity is monitored in real-time using a qPCR instrument at 37°C.
- Lateral Flow Assay: The procedure is identical, except the reporter probe is labeled with FAM and biotin. After incubation at 37°C for 10 minutes, the reaction mixture is diluted and applied to a lateral flow strip. The result is read visually after 3-5 minutes.
- Clinical Validation: The assay's clinical feasibility was tested using 22 clinical isolates of P. aeruginosa, with results compared against standard PCR and sequencing [67].
Analytical Performance Data
The Cas12a-RCFL assay was rigorously evaluated for sensitivity and specificity. The key quantitative results are summarized in the table below.
Table 2: Performance comparison of Cas12a-RCFL against traditional and alternative diagnostic methods.
| Method | Time to Result | Limit of Detection (LoD) | Specificity | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Culture-Based Methods [65] | 48 - 72 hours | 10 - 100 CFU/mL (after enrichment) | High (allows strain typing) | Gold standard, provides live isolates for AST. | Very slow, requires specialized equipment and trained personnel. |
| Conventional PCR [65] | 2 - 4 hours | ~100 fg DNA | High | High throughput, well-established. | Requires precise thermal cycling, sophisticated lab equipment. |
| qPCR (Real-time PCR) [65] | 1 - 2 hours | ~10-100 fg DNA | High | Quantitative, closed-tube reduces contamination. | Expensive instrumentation, complex data analysis. |
| RPA-only [67] | 20 - 30 minutes | ~100 fg DNA | Moderate (prone to non-specific amplification) | Rapid, isothermal, minimal equipment. | Susceptible to false positives from primer-dimers. |
| Cas12a-RCFL [67] | < 60 minutes | 60 fg DNA (~8 genome copies) / 50 CFU/mL | 100% | Extreme speed and sensitivity, high specificity, multiple readout options (fluorimeter, lateral flow). | Requires careful crRNA design, potential for enzymatic inhibition in complex samples. |
The data demonstrates that the Cas12a-RCFL method matches the high sensitivity of advanced qPCR while offering a significantly faster and simpler workflow. Its 100% specificity confirms that the crRNA guide accurately distinguishes P. aeruginosa from other clinically relevant bacteria, including Staphylococcus aureus and Acinetobacter baumannii [67]. The ability to use a crude DNA extract without purification makes it exceptionally suitable for point-of-care settings.
Comparative Analysis in a Broader Diagnostic Context
Cas12a-RCFL vs. Culture and PCR-Based Methods
When evaluated against the broader thesis of diagnostic evolution, Cas12a-RCFL addresses critical limitations of both traditional and modern molecular methods.
- Overcoming the Speed Barrier of Culture: The >99% reduction in time from sample to result (from days to under an hour) is the most transformative advantage. This speed can fundamentally change clinical management, allowing for same-day diagnosis and informed antibiotic stewardship, which is crucial for combating multidrug-resistant P. aeruginosa infections [66] [65].
- Advantages over PCR and Isothermal Methods: While PCR is highly sensitive, it is equipment-intensive. RPA and other isothermal techniques are simple but can suffer from non-specific amplification [65]. The integration of CRISPR-Cas12a adds an additional layer of specificity through RNA-guided recognition, effectively eliminating false positives from non-specific RPA products. This creates a system that is both simple and exceptionally robust [4] [67].
The Critical Role of Targeting theoprLGene
The choice of genetic target is paramount for diagnostic accuracy. The Cas12a-RCFL assay targets the oprL gene, which encodes the peptidoglycan-associated lipoprotein. This gene is a highly specific marker for P. aeruginosa, with studies showing it is absent in other Pseudomonas species and unrelated bacteria [65]. This specificity at the genetic level is the foundation for the assay's demonstrated 100% diagnostic specificity, ensuring reliable identification and reducing the risk of misdiagnosis.
Considerations and Future Directions
Despite its promise, the translation of CRISPR diagnostics like Cas12a-RCFL from the lab to real-world clinical environments faces challenges. As noted in broader reviews, enzymatic activity can be sensitive to inhibitors in complex clinical samples (e.g., sputum, wound exudate), and performance may drop in non-ideal conditions such as high humidity [4]. Future development efforts are focused on creating integrated "sample-to-result" systems and leveraging lyophilized, room-temperature-stable reagents to enhance robustness and ease of use [4]. The integration of machine learning for data analysis, as seen in other novel diagnostic platforms like MX-Raman spectroscopy [69], also represents a promising avenue for enhancing the accuracy and automated interpretation of Cas12a-RCFL results.
The Cas12a-RCFL platform stands as a compelling validation of the thesis that CRISPR-driven diagnostics can overcome the significant limitations of culture and PCR-based methods for detecting challenging pathogens like Pseudomonas aeruginosa. By synergizing the rapid, isothermal amplification of RPA with the precise collateral activity of CRISPR-Cas12a, this assay delivers a diagnostic profile that is simultaneously rapid (under one hour), highly sensitive (down to single-digit genome copies), and exquisitely specific (100%). The flexibility in readout—from quantitative fluorescence to a simple lateral flow strip—makes it adaptable to both well-equipped laboratories and resource-limited settings. For researchers and clinicians battling biofilm-mediated, chronic P. aeruginosa infections, Cas12a-RCFL offers a powerful new tool that promises to shift the paradigm from reactive to proactive infection control, ultimately contributing to better patient outcomes and strengthened antimicrobial stewardship.
The accurate and timely detection of polymicrobial infections remains a significant challenge in clinical diagnostics, particularly when pathogens form biofilms. This review systematically compares CRISPR-based diagnostics against traditional culture methods, focusing on their efficacy in identifying multiple pathogens in biofilm-associated infections. We analyze experimental data demonstrating that CRISPR diagnostics achieve exceptional sensitivity (up to 99%) and specificity (up to 100%) with rapid turnaround times (approximately 60 minutes), significantly outperforming culture methods that require 48-72 hours. By examining the molecular mechanisms, clinical performance data, and implementation protocols, we provide researchers with a comprehensive evidence base for adopting CRISPR technology in polymicrobial infection research and diagnostic development.
Polymicrobial infections, characterized by the presence of multiple pathogenic species, represent a substantial diagnostic challenge in clinical microbiology. These infections are particularly problematic when microorganisms form biofilms—structured communities of microorganisms adherent to surfaces and encased in a self-produced extracellular polymeric substance (EPS) matrix [52] [70]. The biofilm microenvironment creates physical and physiological barriers that limit the penetration of both antimicrobial agents and molecular detection probes, while enabling horizontal gene transfer that accelerates the spread of antimicrobial resistance (AMR) genes [52]. Biofilms are implicated in approximately 80% of chronic microbial human infections, including those affecting the respiratory tract, urinary system, surgical sites, and chronic wounds [70].
Traditional culture-based methods, while considered the gold standard, exhibit significant limitations for polymicrobial biofilm detection. These techniques often require specialized equipment, facilities, and skilled personnel, making them less accessible and cost-effective, especially in resource-limited settings [4]. Furthermore, the process of disaggregating biofilm structures for culture can alter microbial viability and relative abundance, potentially yielding misleading results [70] [71]. The extended time-to-result (typically 48-72 hours) for culture methods delays critical therapeutic decisions, during which time infections can progress and AMR can spread [47]. Molecular methods like PCR have improved detection speed but remain limited in their ability to differentiate live from dead organisms and may struggle with inhibitor interference in complex sample matrices [4] [70].
CRISPR Technology: Molecular Mechanisms for Multiplexed Detection
Core Mechanism of CRISPR-Cas Systems
The CRISPR-Cas system functions as an adaptive immune mechanism in bacteria and archaea, protecting against invading genetic elements such as viruses and plasmids [4] [72]. This system has been repurposed for diagnostic applications through its programmable nucleic acid recognition capabilities. The core mechanism involves two key components: (1) a CRISPR RNA (crRNA) that serves as a guide molecule with sequence complementarity to target pathogen nucleic acids, and (2) Cas effector proteins that perform nucleic acid cleavage upon target recognition [4].
Different Cas enzymes have been harnessed for diagnostic applications, each with distinct properties:
- Cas9: Primarily used for gene editing but can be repurposed for nucleic acid detection through target binding without cleavage in some engineered systems [4].
- Cas12: Targets double-stranded DNA and exhibits robust trans-cleavage activity upon target recognition, nonspecifically degrading surrounding single-stranded DNA molecules [4] [64].
- Cas13: Targets single-stranded RNA and similarly exhibits collateral cleavage activity against non-target RNA molecules, enabling signal amplification [4] [72].
The trans-cleavage activity of Cas12 and Cas13 is particularly valuable for diagnostic applications, as it enables exponential signal amplification through the degradation of reporter molecules that generate fluorescent or colorimetric readouts [4] [72].
Multiplexing Capabilities for Polymicrobial Detection
CRISPR diagnostics can be engineered for multiplexed detection through several strategic approaches:
- Multiple crRNAs: Designing distinct crRNAs targeting conserved regions of different pathogens enables simultaneous detection in a single reaction [64].
- Spatial separation: Microfluidic platforms can physically separate detection reactions for different pathogens while using the same sample input [4].
- Signal differentiation: Using different fluorescent reporters with distinct emission wavelengths for each target pathogen allows discrimination in a single tube [72].
The programmability of CRISPR systems enables rapid adaptation to emerging pathogens or resistance markers by simply redesigning the crRNA sequence, a significant advantage over traditional methods that require extensive revalidation [4] [64].
Figure 1: CRISPR-Cas Diagnostic Workflow. The process begins with sample collection and nucleic acid extraction, followed by target recognition through crRNA-guided Cas proteins, trans-cleavage activation, signal amplification, and final detection.
Comparative Performance Analysis: CRISPR vs. Culture Methods
Direct Performance Metrics
Table 1: Comprehensive Comparison of Diagnostic Platforms for Polymicrobial Biofilm Detection
| Parameter | Culture Methods | PCR-Based Methods | CRISPR Diagnostics |
|---|---|---|---|
| Sensitivity | Variable (10^3-10^4 CFU/mL) | High (10-100 copies/μL) | Exceptional (aM levels) [4] |
| Specificity | High (with confirmatory testing) | High | 99-100% [47] |
| Time to Result | 48-72 hours [47] | 2-4 hours | ~60 minutes [47] |
| Multiplexing Capability | Limited (requires subculturing) | Moderate (4-5 targets) | High (theoretically unlimited) [64] |
| Biofilm Penetration | Poor (requires disaggregation) | Moderate | Enhanced (with nanoparticle delivery) [52] |
| Point-of-Care Suitability | No | Limited | Yes [4] [64] |
| Equipment Requirements | Extensive (incubators, biosafety cabinets) | Moderate (thermocyclers, detectors) | Minimal (isothermal conditions) [4] |
| Cost per Test | Low (reagents) | Moderate | Low to moderate [4] |
| AMR Detection | Phenotypic (requires susceptibility testing) | Genotypic (targeted resistance genes) | Genotypic (multiple resistance markers) [73] |
| Live/Dead Differentiation | Yes (viable organisms only) | No (detects DNA from dead cells) | No (detects nucleic acids) [4] |
Experimental Evidence and Clinical Validation
Recent clinical studies provide compelling evidence for CRISPR's superior performance in pathogen detection. A systematic review and meta-analysis of CRISPR-based detection of methicillin-resistant Staphylococcus aureus (MRSA) demonstrated a pooled sensitivity of 99% (95% CI: 97-100%) and specificity of 100% (95% CI: 99-100%) across 12 studies [47]. The median detection time across these studies was 60 minutes (IQR: 41.25-98.75 minutes), dramatically faster than the 48-72 hours required for culture methods [47].
For polymicrobial detection specifically, CRISPR systems have been successfully configured to simultaneously identify multiple pathogens in complex samples. Integrated platforms combining recombinase polymerase amplification (RPA) or loop-mediated isothermal amplification (LAMP) with CRISPR detection have demonstrated attomolar (aM) sensitivity for bacterial targets including Salmonella, Escherichia coli, and Listeria monocytogenes in food matrices, which present similar challenges to clinical biofilms [64]. The integration of CRISPR reagents into lyophilized formats, microfluidic microarrays, and lateral flow assays has further enhanced the point-of-care applicability of these systems [4].
Experimental Protocols for CRISPR-Based Polymicrobial Detection
Sample Preparation and Nucleic Acid Extraction
Effective sample preparation is critical for accurate detection of polymicrobial biofilms. The extracellular polymeric substance (EPS) matrix can inhibit both amplification and detection steps, requiring optimized extraction protocols.
Protocol 1: Biofilm Disruption and Nucleic Acid Co-extraction
- Biofilm collection: Resuspend biofilm samples in digestion buffer containing proteinase K (0.1 mg/mL) and lysozyme (1 mg/mL). Incubate at 56°C for 30 minutes [70].
- Mechanical disruption: Vortex samples with glass beads (0.1 mm diameter) for 2 minutes or use ultrasonic bath (40 kHz) for 30 seconds [70].
- Nucleic acid extraction: Use commercial silica-column or magnetic bead-based kits compatible with inhibitor removal [64].
- Quality assessment: Measure nucleic acid concentration and purity (A260/A280 ratio >1.8). If inhibited, perform additional purification steps [64].
Protocol 2: Direct Detection without Extraction (for Liquid Samples)
- Sample dilution: Dilute samples 1:5 in nuclease-free water containing 0.1% Tween-20 to reduce inhibitor concentration [64].
- Heat treatment: Incubate at 95°C for 5 minutes to lyse cells and release nucleic acids [72].
- Rapid centrifugation: Spin at 10,000 × g for 1 minute to pellet debris [72].
- Supernatant transfer: Carefully transfer supernatant to reaction vessel, avoiding sediment [72].
CRISPR Detection Workflow
Protocol 3: Multiplexed RPA-CRISPR Assay for Polymicrobial Detection Reagents Required:
- RPA basic kit (TwistDx) containing rehydration buffer, magnesium acetate
- Cas12a enzyme (100 nM)
- Custom crRNAs (200 nM each) targeting pathogen-specific sequences
- Fluorescent reporter (500 nM; e.g., FAM-TTATTATT-BHQ1 for FQ reporters)
- Nuclease-free water
- Lateral flow strips (for alternative endpoint detection)
Procedure:
- RPA premix preparation (25 μL reaction):
- 12.5 μL rehydration buffer
- 5 μL template DNA (5-50 ng)
- 1 μL forward primer (10 μM)
- 1 μL reverse primer (10 μM)
- 5.5 μL nuclease-free water
- Add magnesium acetate last to initiate reaction [64]
RPA amplification:
- Incubate at 37-42°C for 15-20 minutes
- Use portable heat block or water bath [64]
CRISPR detection mix preparation (20 μL reaction):
- 5 μL RPA product
- 1 μL Cas12a (100 nM)
- 2 μL crRNA mix (200 nM each target)
- 1 μL fluorescent reporter (500 nM)
- 11 μL nuclease-free water [72]
Incubation and detection:
- Incubate at 37°C for 10-30 minutes
- Monitor fluorescence in real-time or use endpoint detection
- For lateral flow readout: Add loading buffer and apply to strip [72]
Figure 2: CRISPR Diagnostic Experimental Workflow. The process integrates sample preparation with isothermal amplification options and multiple detection modalities suitable for point-of-care use.
Quality Control and Validation
Protocol 4: Assay Validation and Quality Control
- Positive controls: Include synthetic nucleic acid targets for each pathogen in the panel
- Internal controls: Use non-competitive internal controls to monitor extraction and amplification efficiency
- Limit of detection (LOD) determination: Perform serial dilutions of reference strains to establish detection limits for each target
- Cross-reactivity testing: Validate against near-neighbor species to ensure specificity
- Inhibitor testing: Spike samples with known inhibitors (hemoglobin, mucin, EDTA) to determine robustness [64]
Research Reagent Solutions for CRISPR-Based Detection
Table 2: Essential Research Reagents for Developing CRISPR Polymicrobial Detection Assays
| Reagent Category | Specific Examples | Function | Considerations for Polymicrobial Detection |
|---|---|---|---|
| Cas Effectors | Cas12a (LbCas12a, AsCas12a), Cas13a (LwaCas13a) | Target recognition and trans-cleavage | Cas12 for DNA targets, Cas13 for RNA targets; some effectors prefer specific PAM sequences [4] |
| crRNA Design | Custom synthetic crRNAs (IDT, Synthego) | Sequence-specific guidance | Design crRNAs against conserved pathogen regions; avoid cross-homology; test multiple guides per target [4] |
| Amplification Enzymes | RPA (TwistAmp), LAMP (Bst polymerase) | Isothermal nucleic acid amplification | RPA operates at lower temperatures (37-42°C); LAMP offers higher sensitivity but more complex primer design [64] |
| Reporters | FAM-TTATTATT-BHQ1 (FQ), FAM-biotin (lateral flow) | Signal generation | Fluorescent reporters enable quantification; lateral flow reporters allow visual detection [72] |
| Sample Preparation | Proteinase K, lysozyme, magnetic beads | Nucleic acid extraction and purification | Optimize for EPS matrix disruption; include inhibitor removal steps [70] |
| Delivery Systems | Lipid nanoparticles, gold nanoparticles | Enhanced cellular uptake | Particularly valuable for biofilm penetration and intracellular pathogen detection [52] |
| Readout Platforms | Portable fluorometers, lateral flow strips, smartphone detectors | Result visualization | Choose based on required sensitivity and field-deployment needs [64] |
Implementation Challenges and Future Directions
Despite the promising performance of CRISPR diagnostics for polymicrobial biofilm detection, several challenges remain for widespread implementation. Inhibitor interference in complex clinical samples can reduce sensitivity, requiring optimized sample preparation methods [4]. The regulatory landscape for CRISPR-based diagnostics is still evolving, with varying requirements across regions that may slow clinical adoption [4]. Additionally, scalability and manufacturing considerations for point-of-care devices need addressing for global accessibility [64].
Future developments are likely to focus on integrated "sample-to-result" systems that automate the entire diagnostic process [4]. The incorporation of artificial intelligence for assay optimization and result interpretation shows significant promise for enhancing accuracy and usability [4] [64]. Multiplexing capacity continues to expand, with researchers developing systems capable of detecting dozens of pathogens simultaneously [64]. Finally, nanoparticle-enhanced delivery may improve biofilm penetration and detection efficiency for particularly challenging samples [52].
For research applications, continued refinement of CRISPR-based detection platforms will enable more comprehensive analysis of polymicrobial infections, potentially revealing new insights into microbial community dynamics, resistance mechanisms, and pathogen evolution in biofilm environments.
CRISPR-based diagnostics represent a transformative approach for detecting polymicrobial infections in biofilm contexts, offering significant advantages over traditional culture methods in speed, sensitivity, and multiplexing capability. Experimental data demonstrates that CRISPR platforms can achieve exceptional performance metrics (99% sensitivity, 100% specificity) with rapid turnaround times (approximately 60 minutes), enabling timely therapeutic decisions and infection control interventions. While implementation challenges remain, ongoing technological advancements in sample preparation, reagent formulation, and readout platforms continue to enhance the utility of CRISPR systems for complex diagnostic scenarios. For researchers and drug development professionals, CRISPR technology provides a powerful tool for investigating polymicrobial biofilms and developing next-generation diagnostic solutions to address the growing challenge of antimicrobial resistance.
Biofilm-associated infections represent a significant challenge in clinical settings, particularly in cases involving medical implants such as prosthetic joints. These structured communities of microorganisms, encased in a self-produced extracellular polymeric substance (EPS) matrix, exhibit unique resistance to conventional antimicrobial therapies and complicate pathogen detection [13] [24]. The inherent limitations of traditional culture-based methods have prompted the development of molecular diagnostics, with CRISPR-based systems emerging as a promising alternative. This analysis evaluates the cost-effectiveness of CRISPR diagnostics compared to traditional culture methods for detecting biofilm-forming pathogens, examining both immediate economic impacts and long-term clinical outcomes.
The diagnostic dilemma is particularly acute in periprosthetic joint infections (PJIs), where biofilms contribute to false-negative cultures and delayed treatment. Biofilm-forming bacteria in orthopedics include Staphylococcus aureus, coagulase-negative staphylococci, and Enterococcus spp., with fungal pathogens like Candida spp. occurring less frequently [74]. These microorganisms employ multiple resistance mechanisms within biofilms, including reduced metabolic activity, enhanced horizontal gene transfer, and physical protection from antibiotics—features that often render conventional diagnostics insufficient [13]. Against this backdrop, CRISPR diagnostics offer a potential paradigm shift through their programmable nucleic acid detection capabilities, though their economic viability requires thorough investigation alongside their technical performance.
Methodological Approaches: Traditional versus CRISPR-Based Diagnostics
Conventional Culture-Based Methods
Traditional microbiological diagnosis of biofilm-associated infections typically relies on tissue sampling and culture techniques. The standard protocol involves collecting multiple periprosthetic tissue samples (typically five), which are processed separately through homogenization and inoculation onto various culture media. These samples undergo aerobic and anaerobic incubation for specified periods, usually ranging from 2-10 days, followed by biochemical identification and antibiotic susceptibility testing of isolated pathogens [74]. While this approach benefits from established protocols and widespread availability, its effectiveness is limited by several factors: biofilm encapsulation reduces microbial dispersal, resulting in false-negative rates; sample contamination can cause false positives; and the extended time-to-result delays appropriate therapeutic interventions [75] [74].
To improve biofilm recovery, mechanical and chemical debonding techniques have been developed, including sonication and dithiothreitol (DTT) treatment. Sonication involves subjecting explanted prostheses to low-frequency sound waves to dislodge biofilm-embedded bacteria, while DTT treatment breaks disulfide bonds in the extracellular matrix to release microbial cells [75]. Although these ancillary methods enhance diagnostic sensitivity, they add procedural complexity and increase laboratory processing requirements without fundamentally addressing the time-intensive nature of culture-based approaches.
CRISPR-Based Diagnostic Platforms
CRISPR diagnostics utilize programmable CRISPR-associated (Cas) proteins coupled with guide RNA sequences to detect specific nucleic acid targets with high specificity. The fundamental mechanism involves Cas enzyme activation upon target recognition, triggering collateral cleavage activity that generates a detectable signal [4] [25]. Different Cas proteins offer distinct advantages: Cas12 targets double-stranded DNA with single-stranded DNA trans-cleavage activity; Cas13 recognizes RNA targets with single-stranded RNA collateral cleavage; and Cas9 provides precise DNA binding through its guide RNA-directed nuclease activity [41] [25] [76].
Established CRISPR platforms include SHERLOCK (Specific High Sensitivity Enzyme Reporter Unlocking), which employs Cas13 for RNA detection, and DETECTR (DNA Endonuclease Targeted CRISPR Trans Reporter), which utilizes Cas12 for DNA identification [41] [76]. These systems typically integrate with preliminary nucleic acid amplification steps, such as recombinase polymerase amplification (RPA) or loop-mediated isothermal amplification (LAMP), to enhance sensitivity without requiring thermal cycling equipment [41] [64]. Recent advancements include amplification-free CRISPR approaches that reduce operational complexity and potential contamination, though with some sensitivity trade-offs [76]. The typical workflow involves sample collection, nucleic acid extraction, isothermal amplification (for amplification-based methods), CRISPR reaction with reporter molecules, and signal detection through fluorescence, colorimetry, or lateral flow readouts [25] [76].
Table 1: Key CRISPR-Cas Proteins and Their Diagnostic Applications
| Cas Protein | Nucleic Acid Target | Trans-Cleavage Activity | Primary Applications | Detection Limit |
|---|---|---|---|---|
| Cas12 | Double-stranded DNA | Single-stranded DNA | Bacterial DNA detection, HPV, SARS-CoV-2 | 10 copies/μL [41] |
| Cas13 | Single-stranded RNA | Single-stranded RNA | RNA viruses (Zika, Dengue, SARS-CoV-2) | Attomolar level [41] |
| Cas9 | Double-stranded DNA | None (cis-cleavage only) | SNP detection, genetic mutation identification | Varies by target |
| Cas14 (Cas12f) | Single-stranded DNA | Single-stranded DNA | Small footprint applications, point-of-care testing | ~470 aM [76] |
Comparative Performance Analysis
Diagnostic Accuracy and Turnaround Time
The most significant advantage of CRISPR diagnostics over traditional methods lies in their combination of rapid results and high sensitivity. While culture-based methods require 2-10 days for pathogen identification [74], CRISPR systems can deliver results in approximately 30-60 minutes following sample processing [41] [64]. This dramatic reduction in time-to-result enables same-day treatment decisions, potentially improving patient outcomes and reducing complications associated with diagnostic delays.
Regarding sensitivity, culture-based methods for prosthetic joint infections demonstrate variable performance, with tissue culture sensitivity ranging from 61% to 76% compared to sonicated fluid cultures at 77-95% [75]. In contrast, CRISPR platforms consistently achieve superior sensitivity metrics: Cas12-based detection of SARS-CoV-2 reaches 98% sensitivity and 100% specificity with a detection limit of 10 copies/μL [41], while Cas13-based SHERLOCK technology detects Zika virus at attomolar concentrations with near-perfect specificity [41]. For bacterial targets, Cas12-based detection of Mycobacterium tuberculosis demonstrates 88.3% sensitivity and 94.6% specificity with a limit of detection of 3.13 CFU/mL [41]. This enhanced sensitivity is particularly valuable for biofilm-associated infections where microbial loads may be low and unevenly distributed.
The following diagram illustrates the significantly different workflows and time requirements for conventional culture versus CRISPR-based diagnostics:
Economic Considerations: Initial Investment Versus Long-Term Value
A comprehensive cost-benefit analysis must account for both direct expenses and indirect clinical impacts. Direct cost calculations for biofilm diagnostics reveal that traditional tissue cultures incur approximately €308 per patient, while advanced antibiofilm techniques like sonication and DTT treatment cost €397 and €393 per patient, respectively [75]. Although specific direct costs for CRISPR diagnostics are not explicitly detailed in the available literature, the technology's requirements for specialized reagents, Cas proteins, and guide RNA synthesis suggest higher initial consumable costs compared to conventional media-based cultures.
However, the economic assessment becomes more favorable for CRISPR diagnostics when considering indirect costs and long-term outcomes. Diagnostic inaccuracy generates substantial downstream expenses, including unnecessary antibiotic treatments, extended hospital stays, additional surgical procedures, and management of infection recurrence [75]. One economic modeling study demonstrated that sonication and DTT become cost-effective when traditional tissue culture generates inaccurate diagnoses in just 2-20% of cases [75]. With CRISPR's superior sensitivity and specificity profile, the reduction in false negatives and false positives could yield significant savings by enabling appropriate first-line treatment and avoiding complications from undiagnosed infections.
Table 2: Comprehensive Cost-Benefit Analysis of Diagnostic Methods
| Cost Component | Traditional Tissue Culture | Culture with Sonication | CRISPR-Based Diagnostics |
|---|---|---|---|
| Direct Costs | |||
| - Materials & consumables | €210-250 per patient [75] | €280-320 per patient [75] | Higher reagent costs (estimated) |
| - Laboratory personnel time | €98 per patient [75] | €117 per patient [75] | Lower hands-on time (estimated) |
| Total Direct Cost | €308 per patient [75] | €397 per patient [75] | Not specified (higher initial) |
| Indirect Cost Factors | |||
| - False negative rate | 24-39% [75] [74] | 5-28.6% [75] | <12% (estimated based on sensitivity data) |
| - False positive rate | 23.5% [75] | 5.88% [75] | <2% (estimated based on specificity data) |
| - Time to result | 2-10 days [74] | 2-5 days | 30-60 minutes [41] |
| Long-Term Value | |||
| - Appropriate initial treatment | Lower due to delayed results | Moderate | Higher due to rapid results |
| - Hospital stay duration | Potentially extended | Potentially reduced | Significantly reducible |
| - Need for revision surgery | Higher due to missed diagnoses | Moderate | Lower with accurate detection |
Advanced CRISPR Applications for Biofilm Pathogens
Integration with Nanoparticle Delivery Systems
A groundbreaking application of CRISPR technology for biofilm infections combines diagnostic capabilities with therapeutic potential through nanoparticle-mediated delivery. Recent advances demonstrate that lipid-based Cas9 formulations can reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro, while gold nanoparticle carriers enhance gene-editing efficiency up to 3.5-fold compared to non-carrier systems [13]. This integrated approach targets both genetic resistance determinants (e.g., bla, mecA, ndm-1 genes) and biofilm structural components, addressing the dual challenge of inherited and phenotypic resistance mechanisms [13].
The therapeutic application operates through two complementary mechanisms: CRISPR components disrupt antibiotic resistance genes or biofilm-regulating factors through precise gene editing, while nanoparticles facilitate enhanced penetration through the protective EPS matrix [13]. These hybrid platforms can co-deliver antibiotics or antimicrobial peptides alongside CRISPR machinery, creating synergistic antibacterial effects and superior biofilm disruption compared to mono-therapeutic approaches [13]. Although primarily therapeutic, this technology has diagnostic implications through its ability to identify and target specific genetic markers of resistance within complex biofilm communities.
Multiplex Detection Platforms
CRISPR technology enables simultaneous detection of multiple pathogens through multiplex assay designs. By utilizing different Cas proteins with distinct guide RNAs or incorporating spatial separation in microfluidic devices, these platforms can identify numerous bacterial species, resistance genes, or fungal targets in a single reaction [41] [64]. This capability is particularly valuable for polymicrobial biofilm infections, where traditional cultures may overlook fastidious organisms or mixed microbial communities.
Advanced biosensing platforms integrate CRISPR with various detection methodologies, including graphene-based field-effect transistors (gFET), electrochemiluminescence (ECL), and surface-enhanced Raman spectroscopy (SERS) [76] [64]. These interfaces enhance signal amplification and enable quantitative detection across diverse pathogen targets. For clinical applications, lateral flow assays provide simple visual readouts without requiring sophisticated instrumentation, making CRISPR diagnostics adaptable to both centralized laboratories and point-of-care settings [41] [76].
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of CRISPR-based diagnostics for biofilm pathogens requires specific reagent systems and methodological approaches. The following toolkit outlines essential components and their functions for developing and optimizing these assays:
Table 3: Research Reagent Solutions for CRISPR-Based Biofilm Pathogen Detection
| Reagent Category | Specific Examples | Function in Diagnostic Assay |
|---|---|---|
| CRISPR Enzymes | Cas12a (LbCas12a, AsCas12a), Cas13a (LbuCas13a), Cas9 | Core detection proteins with target-specific cleavage activities |
| Guide RNA Components | crRNA, sgRNA, tracrRNA | Programmable recognition elements that direct Cas proteins to specific nucleic acid targets |
| Nucleic Acid Amplification Reagents | RPA (recombinase polymerase amplification), LAMP (loop-mediated isothermal amplification) | Pre-amplification systems to enhance detection sensitivity without thermal cycling |
| Signal Detection Reporters | FQ-labeled ssDNA reporters (for Cas12), FQ-labeled ssRNA reporters (for Cas13), lateral flow strips with FAM/biotin labels | Generate measurable signals (fluorescence, colorimetric) through collateral cleavage |
| Biofilm Disruption Reagents | Dithiothreitol (DTT), proteinase K, sonication protocols | Release microbial nucleic acids from protective biofilm matrix for improved detection |
| Nanoparticle Delivery Systems | Gold nanoparticles, lipid nanoparticles, polymeric nanoparticles | Enhance CRISPR component delivery through biofilm matrices for therapeutic applications |
| Sample Preservation & Preparation | Nucleic acid extraction kits, inhibitor-resistant enzymes, transport media | Maintain target integrity and remove PCR inhibitors from complex clinical samples |
Future Directions and Implementation Challenges
Despite its promising capabilities, CRISPR diagnostic technology faces several implementation barriers. The initial investment required for platform establishment, reagent development, and personnel training presents a significant hurdle for resource-limited settings [77] [25]. Additionally, performance variability in complex clinical samples containing biofilm components, blood, or tissue debris may affect reproducibility [25] [64]. Regulatory approval pathways for CRISPR-based in vitro diagnostics remain evolving, requiring extensive clinical validation across diverse patient populations and biofilm types [41] [25].
Future development trajectories focus on integrating artificial intelligence to optimize guide RNA design and predict off-target effects [77] [41], creating "sample-to-result" automated systems that minimize manual processing [25], and developing stable lyophilized reagent formulations for distribution without cold-chain requirements [4] [64]. The ongoing discovery of novel Cas proteins with unique properties (e.g., Cas14, CasΦ) may further expand the diagnostic toolbox for biofilm-associated pathogens [76].
From a health economics perspective, the successful implementation of CRISPR diagnostics will depend on demonstrating not just technical superiority but also cost-effectiveness through reduced hospitalization durations, appropriate antibiotic stewardship, and decreased revision surgery rates. Future economic studies should incorporate real-world clinical outcome data and total cost-of-care analyses to provide comprehensive evidence for healthcare decision-makers.
CRISPR-based diagnostics represent a transformative approach to detecting biofilm-associated pathogens, offering significant advantages in speed, sensitivity, and specificity compared to traditional culture methods. While the initial investment in CRISPR technology exceeds that of conventional diagnostics, the long-term clinical and economic benefits—including rapid treatment initiation, reduced hospital stays, and decreased revision surgeries—present a compelling value proposition. The integration of CRISPR with nanoparticle delivery systems and multiplex detection platforms further enhances its potential for addressing the persistent challenge of biofilm-related infections. As research advances and implementation barriers are addressed, CRISPR diagnostics are poised to significantly improve patient outcomes while optimizing healthcare resource utilization in the management of complex biofilm-associated infections.
The detection and control of biofilm-forming pathogens represent a critical frontier in clinical medicine and food safety. Biofilms—structured communities of microorganisms embedded in a protective extracellular polymeric substance (EPS)—account for approximately 80% of bacterial infections in the United States and impose estimated annual costs of $324 billion to the global agrifood sector [59] [16]. Traditional culture-based methods, while considered the historical gold standard, require 2-3 days for pathogen identification and exhibit significant limitations in detecting stressed or biofilm-associated bacteria [15] [47]. The emergence of CRISPR-based diagnostics offers a transformative approach, enabling rapid, precise pathogen detection with potential applications spanning clinical diagnostics and food safety monitoring. This review objectively evaluates the real-world performance of CRISPR diagnostics against conventional culture methods for detecting biofilm-forming pathogens, providing experimental data and methodological insights for research and development professionals.
Performance Comparison: CRISPR vs. Culture Methods
Table 1: Comprehensive Performance Metrics of CRISPR vs. Traditional Methods
| Parameter | CRISPR-Based Methods | Traditional Culture Methods | Molecular Methods (PCR/qPCR) |
|---|---|---|---|
| Sensitivity (Pooled) | 99% (95% CI: 97-100%) for MRSA [47] | Highly variable; reduced for stressed cells | High (>95%) but inhibitor-sensitive |
| Specificity (Pooled) | 100% (95% CI: 99-100%) for MRSA [47] | High for viable organisms | High (>95%) with proper primer design |
| Time to Result | ~60 minutes (median, IQR: 41-99 min) [47] | 48-72 hours [47] | 2-4 hours including amplification [11] |
| Detection Limit | aM (attomolar) level; single-copy sensitivity with pre-amplification [4] [11] | 10³-10⁴ CFU for most pathogens | Varies; typically 10-100 gene copies |
| Biofilm Matrix Penetration | Limited data; nanoparticles enhancing delivery [13] | Limited to surface-colonizing cells | Inhibited by EPS components |
| Point-of-Care Applicability | High (lateral flow, portable readers) [64] | Low (requires lab infrastructure) | Low (requires thermal cycling) |
| Stressed/Dead Cell Detection | Detects nucleic acids regardless of viability [78] | Only detects viable cells | Detects nucleic acids regardless of viability |
| Multiplexing Capacity | High (multiple crRNAs) [16] | Limited (differential media) | Moderate (primer-dependent) |
Table 2: Food Safety Monitoring: Performance in Complex Matrices
| Pathogen | CRISPR Platform | Sample Matrix | Performance Metrics | Comparison to Culture |
|---|---|---|---|---|
| Salmonella spp. | Cas12 + RPA [64] | Chicken, produce | 100% sensitivity, 10 CFU/mL sensitivity [64] | Equivalent detection, >24h faster |
| E. coli STEC | Cas13 + LAMP [64] | Ground beef, spinach | 98.5% sensitivity, 99.2% specificity [64] | Detects non-culturable strains |
| Listeria monocytogenes | Cas12 + RPA [64] | Dairy products, processed meats | 100 CFU/g in 3h [64] | >48h faster than culture methods |
| S. aureus/MRSA | Cas9/Cas12 + RPA [47] | Clinical specimens (wound, nasal) | 99% sensitivity, 100% specificity [47] | 60 min vs. 48-72h for culture |
| Multiplex Panels | Cas12/Cas13 combo [16] | Food processing surfaces | Simultaneous detection of 3 pathogens [16] | Identifies co-contamination in single assay |
Experimental Protocols for CRISPR-Based Detection
CRISPR-MRSA Detection Protocol (Clinical Validation)
This protocol outlines the methodology used in clinical studies evaluating CRISPR for MRSA detection, which demonstrated 99% sensitivity and 100% specificity in a meta-analysis of 12 studies [47].
Table 3: Key Research Reagent Solutions
| Reagent | Function | Specifications/Alternatives |
|---|---|---|
| Cas12a (Cpf1) Enzyme | Target recognition & trans-cleavage | Recombinant, purified, >90% purity [4] |
| crRNA Guide | Target-specific recognition | Designed against mecA gene; HPLC purified [47] |
| ssDNA FQ Reporter | Fluorescent signal generation | FAM-TTATT-BHQ1; quenched until cleavage [11] |
| RPA/LAMP Primers | Isothermal amplification | Target species-specific genes (e.g., spa, mecA) [47] |
| Lateral Flow Strips | Visual readout | Anti-FAM/test line, biotin/control line compatible [11] |
Step-by-Step Workflow:
- Nucleic Acid Extraction: Extract DNA from clinical samples (swabs, blood, sputum) using commercial kits. Heat lysis (95°C, 5 min) may suffice for rapid protocols.
- Isothermal Amplification: Perform Recombinase Polymerase Amplification (RPA) or LAMP:
- RPA Reaction: 39°C for 15-20 minutes using primers targeting MRSA-specific sequences (mecA, spa).
- LAMP Alternative: 65°C for 30 minutes with 4-6 primers for higher sensitivity.
- CRISPR-Cas Detection:
- Prepare reaction mix: 50 nM Cas12a, 75 nM crRNA (designed against mecA gene), 500 nM ssDNA reporter in appropriate buffer.
- Add 2 µL amplification product to CRISPR reaction.
- Incubate at 37°C for 10-15 minutes.
- Signal Readout:
- Fluorescence: Measure in real-time or endpoint using plate readers or portable devices.
- Lateral Flow: Apply reaction mixture to strip; results in 5-10 minutes.
- Visual Detection: Under blue light or naked eye for high-titer samples.
Diagram 1: MRSA Detection Workflow (Title: CRISPR-Cas12 MRSA Detection Protocol)
Biofilm Pathogen Detection in Food Processing Environments
This protocol details methodologies for detecting biofilm-forming pathogens on food-contact surfaces, addressing a critical challenge in food safety [78] [59].
Sample Collection and Preparation:
- Surface Sampling: Swab stainless steel, plastic, or glass surfaces (standard area: 10x10 cm) in food processing environments.
- Biofilm Disruption: Resuspend swabs in 1-2 mL neutralizing buffer with gentle vortexing. For enhanced recovery, include brief sonication (30 seconds, 40 kHz) to disaggregate biofilm microcolonies.
- Sample Treatment: Centrifuge at 5000×g for 5 minutes; use supernatant for direct detection or pellet for nucleic acid extraction.
CRISPR Detection with Pre-enrichment:
- Short Enrichment: Incubate sample in selective broth (4-6 hours) to increase bacterial load while maintaining rapid turnaround.
- Nucleic Acid Extraction: Use rapid spin-column or magnetic bead-based methods (15-20 minutes).
- Multiplex RPA: Co-amplify targets (e.g., hilA for Salmonella, prfA for Listeria, mecA for Staphylococcus) at 39°C for 20 minutes.
- Cas12/Cas13 Cascade:
- Use Cas12 for DNA targets (e.g., bacterial genomic DNA)
- Use Cas13 for RNA targets (e.g., messenger RNA for viability assessment)
- Multiplex crRNAs in single reaction vessel
- Lateral Flow Readout: Design reporters with different haptens (FAM, DIG, biotin) for parallel detection on multiplex lateral flow strips.
Molecular Mechanisms and Technological Advancements
CRISPR-Cas Mechanisms for Pathogen Detection
The exceptional specificity of CRISPR diagnostics stems from the molecular architecture of Cas protein complexes. The system relies on two fundamental activities: sequence-specific recognition through crRNA:target hybridization, and subsequent activation of non-specific collateral cleavage of reporter molecules [4] [11].
Diagram 2: CRISPR Detection Mechanism (Title: CRISPR-Cas Nucleic Acid Detection Mechanism)
Key Protein Mechanisms:
- Cas12 (DETECTR system): Upon recognizing target DNA via crRNA and PAM sequence, activates non-specific single-stranded DNA (ssDNA) cleavage, degrading fluorescent reporters [4] [64].
- Cas13 (SHERLOCK system): Binds target RNA via crRNA, triggering collateral trans-cleavage of single-stranded RNA reporters, enabling RNA virus detection [4] [11].
- Cas14: Targets single-stranded DNA without PAM sequence requirements, particularly useful for detecting small DNA pathogens [11].
Integration with Advanced Detection Platforms
Recent innovations have enhanced CRISPR diagnostics through integration with diverse detection modalities:
Portable Biosensors: CRISPR systems coupled with electrochemical sensors (e.g., graphene field-effect transistors) enable real-time pathogen monitoring in food processing facilities [16]. These systems demonstrate attomolar sensitivity in complex matrices like meat rinsates and dairy products [64].
Digital Droplet CRISPR: Partitioning reactions into water-in-oil emulsions enables absolute quantification of pathogen load without standard curves, providing critical data for biofilm formation stages [11].
Nanoparticle Enhancement: Gold nanoparticles and lipid nanoparticles improve CRISPR component delivery through biofilm matrices and enhance signal generation, addressing a key limitation in traditional molecular methods [13].
Comparative Limitations and Technical Challenges
Performance in Complex Matrices
While CRISPR demonstrates exceptional performance in controlled laboratory conditions, its efficacy diminishes in complex sample matrices:
Clinical Samples: Blood, sputum, and wound exudates contain nucleases, heparin, and hemoglobin that inhibit CRISPR reactions, potentially reducing sensitivity by 10-100 fold without proper sample processing [64].
Food Matrices: Fats, proteins, and polysaccharides in food samples interfere with nucleic acid extraction and Cas enzyme activity, requiring sophisticated sample preparation or dilution that impacts detection limits [64].
Biofilm-specific Challenges: The extracellular polymeric substance (EPS) matrix in biofilms impedes reagent penetration and nucleic acid extraction efficiency, potentially leading to false negatives without optimized disruption protocols [15] [13].
Limitations of Traditional Methods
Culture methods fundamentally lack detection capability for viable but non-culturable (VBNC) and sublethally injured bacteria, a significant limitation in assessing sanitizer efficacy [78]. One study demonstrated that while ATP testing showed 2-log reduction in detection of chlorine-stressed E. coli, fluorescence-based methods (like Bactiscan) and CRISPR maintained consistent detection regardless of bacterial stress status [78].
CRISPR-based diagnostics represent a paradigm shift in pathogen detection, offering unprecedented speed (60 minutes vs. 48-72 hours) while maintaining exceptional accuracy (99% sensitivity, 100% specificity for MRSA) compared to traditional culture methods [47]. The technology's programmability enables rapid adaptation to emerging pathogens and multiplexed detection schemes crucial for identifying polymicrobial biofilms.
Future developments will likely focus on overcoming matrix inhibition effects through improved sample preparation, integrating CRISPR with microfluidic systems for fully automated "sample-to-answer" platforms, and leveraging artificial intelligence for guide RNA design and experimental optimization [16]. For biofilm-specific applications, CRISPR-nanoparticle conjugates show particular promise for enhanced penetration and delivery [13].
While regulatory approval and standardization remain challenges, the robust real-world performance data across clinical and food safety applications positions CRISPR technology as an indispensable tool for next-generation pathogen detection, ultimately strengthening global public health responses to biofilm-associated infections and contaminations.
Conclusion
The evidence firmly positions CRISPR diagnostics as a transformative tool for biofilm pathogen detection, offering a compelling alternative to traditional culture methods. By synthesizing the key intents, it is clear that CRISPR technology addresses the foundational limitations of speed and sensitivity, provides a versatile methodological toolkit for precise application, and is overcoming technical hurdles through ongoing optimization. Empirical validation confirms its superior performance, with detection possible within hours instead of days and at sensitivities reaching attomolar levels. For researchers and drug development professionals, the future direction is clear: the integration of CRISPR with AI for predictive analysis, the development of robust 'sample-to-answer' point-of-care devices, and the creation of multiplexed panels for complex polymicrobial biofilms will bridge the gap between laboratory innovation and equitable global health solutions, ultimately enabling more effective biofilm control and antimicrobial stewardship.
References
- 1. Mechanisms of antimicrobial resistance in biofilms [https://www.nature.com/articles/s44259-024-00046-3]
- 2. The EPS Matrix: The “House of Biofilm Cells” - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2168682/]
- 3. Bacterial persisters: molecular mechanisms and ... [https://www.nature.com/articles/s41392-024-01866-5]
- 4. CRISPR‐driven diagnostics: Molecular mechanisms, clinical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12464570/]
- 5. Bacterial Persister Cell Formation and Dormancy - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3837759/]
- 6. Comparative evaluation of methods for the detection of biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5926075/]
- 7. Comparative analysis of biofilm detection methods and ... [https://bmcmicrobiol.biomedcentral.com/articles/10.1186/s12866-025-04506-2]
- 8. Polymicrobial Infections: A Comprehensive Review on ... [https://www.mdpi.com/2813-9054/70/4/39]
- 9. Detection limitations of bacteria in tissue samples - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12178716/]
- 10. Prospective study on diagnostic efficacy of targeted NGS vs ... [https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2025.1488652/full]
- 11. Advances in the application of CRISPR technology ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1645699/full]
- 12. Advances in the application of CRISPR technology ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12620484/]
- 13. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-025-04323-4]
- 14. Bead-based approaches for increased sensitivity and ... [https://www.nature.com/articles/s41551-025-01498-2]
- 15. New Methodologies as Opportunities in the Study of Bacterial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12114119/]
- 16. CRISPR–Cas systems as emerging tools for precision ... [https://www.sciencedirect.com/science/article/abs/pii/S0963996925021416]
- 17. Emerging Technologies and Integrated Strategies for ... [https://www.mdpi.com/2076-2607/13/7/1447]
- 18. Resilient sustainable current and emerging technologies for ... [https://pubs.rsc.org/en/content/articlehtml/2025/fb/d4fb00192c]
- 19. Next-generation rapid phenotypic antimicrobial ... [https://www.nature.com/articles/s41467-024-53930-x]
- 20. A highly sensitive MiRNA detection sensor powered by ... [https://www.sciencedirect.com/science/article/abs/pii/S1567539425000957]
- 21. Point of Care Diagnostics in Resource-Limited Settings [https://www.mdpi.com/2079-6374/10/10/133]
- 22. Current practices for assessing usability of novel point-of-care ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12336581/]
- 23. Comparative analysis of biofilm detection methods and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12625357/]
- 24. A comprehensive review on biofilm-associated infections [https://www.sciencedirect.com/science/article/pii/S2950194625002043]
- 25. CRISPR-Cas-Based Diagnostics in Biomedicine [https://www.mdpi.com/2079-6374/15/10/660]
- 26. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11257871/]
- 27. Current and Future Technologies for the Detection of ... [https://www.mdpi.com/2075-4418/13/20/3246]
- 28. Exploring the Trans-Cleavage Activity of CRISPR Cas12a ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6938695/]
- 29. Enhancement of trans-cleavage activity of Cas12a with ... [https://www.nature.com/articles/s41467-020-18615-1]
- 30. Recent developments and future directions in point-of-care ... [https://link.springer.com/article/10.1007/s10238-024-01540-8]
- 31. Current and Future Technologies for the Detection of Antibiotic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10606640/]
- 32. CRISPR-based diagnostics of different biomolecules - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10577475/]
- 33. Engineered CRISPR-Cas systems for the detection and ... [https://jnanobiotechnology.biomedcentral.com/articles/10.1186/s12951-021-01132-8]
- 34. CRISPR/Cas12a-Based Diagnostic Platform Accurately ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2022.884411/full]
- 35. Frontiers | Research progress on the application of RPA-CRISPR/Cas12a in the rapid visual detection of pathogenic microorganisms [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1640938/full]
- 36. CRISPR-Cas Systems: Bridging Bacterial Immunity and ... [https://www.mdpi.com/2673-8007/5/4/118]
- 37. Research progress on the application of RPA-CRISPR/Cas12a in the rapid visual detection of pathogenic microorganisms - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12343600/]
- 38. Harnessing bacterial immunity: CRISPR-Cas system as a ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1588446/full]
- 39. Ultrasensitive detection of clinical pathogens through a ... [https://www.nature.com/articles/s41467-025-59219-x]
- 40. Direct multiplex detection of full length viral nucleic acids ... [https://www.sciencedirect.com/science/article/pii/S0925400525009888]
- 41. Recent developments and future directions in point-of-care ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11717804/]
- 42. Clinical validation of a Cas13-based assay for the ... [https://www.nature.com/articles/s41551-020-00603-x]
- 43. Lateral Flow Readout for CRISPR/Cas-based detection ... [https://www.milenia-biotec.com/en/tips-lateral-flow-readouts-crispr-cas-strategies/]
- 44. SHERLOCK and DETECTR: CRISPR-Cas Systems as ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8106734/]
- 45. Finding nucleic acids with SHERLOCK and DETECTR [https://blog.addgene.org/finding-nucleic-acids-with-sherlock-and-detectr]
- 46. Real-Time, Multiplexed SHERLOCK for in Vitro Diagnostics [https://pmc.ncbi.nlm.nih.gov/articles/PMC10122965/]
- 47. Fast but accurate: a systematic review and meta-analysis ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1703247/full]
- 48. Evaluation of the diagnostic efficacy of CRISPR-based ... [https://www.sciencedirect.com/science/article/abs/pii/S0732889325003827]
- 49. Biofilms Exposed: Innovative Imaging and Therapeutic ... [https://www.mdpi.com/2079-6382/14/9/865]
- 50. Quantitative and Qualitative Assessment Methods for Biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6133255/]
- 51. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm- ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2024.1408569/full]
- 52. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366227/]
- 53. Analysis of biofilm assembly by large area automated AFM [https://www.nature.com/articles/s41522-025-00704-y]
- 54. How to study biofilms: technological advancements in ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2023.1335389/full]
- 55. Biofilms: structure, resistance mechanism, emerging control ... [https://pubs.rsc.org/en/content/articlehtml/2025/pm/d5pm00094g]
- 56. New Methodologies as Opportunities in the Study of ... [https://www.mdpi.com/2076-2607/13/5/1062]
- 57. Bioactivity of microbial biofilms in extreme environments [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1602583/full]
- 58. Culture-free detection of bacteria from blood for rapid ... [https://www.nature.com/articles/s41746-025-01948-w]
- 59. Advances in the Detection and Identification of Bacterial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11941196/]
- 60. Isolation of Biofilm-Forming Bacteria from Food Processing ... [https://www.mdpi.com/2304-8158/14/21/3592]
- 61. Application progress and biosafety challenges of gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12624554/]
- 62. Biosafety: New NIH Transparency Guidance – April 2025 [https://umaine.edu/research-compliance/2025/04/29/biosafety-new-nih-transparency-guidance-april-2025/]
- 63. Microbial culture vs. mNGS: diagnostic variations in ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1611332/full]
- 64. CRISPR revolution: Unleashing precision pathogen ... [https://www.sciencedirect.com/science/article/pii/S104620232500115X]
- 65. Rapid detection of Pseudomonas aeruginosa by ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2023.1239269/full]
- 66. Impact of Pseudomonas aeruginosa biofilm formation by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12090236/]
- 67. Rapid and sensitive detection of Pseudomonas aeruginosa ... [https://www.nature.com/articles/s41598-023-45766-0]
- 68. CRISPR-Cas12a: Functional overview and applications [https://pmc.ncbi.nlm.nih.gov/articles/PMC7090318/]
- 69. Identification and antimicrobial resistance profiling of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12378394/]
- 70. Options and Limitations in Clinical Investigation of Bacterial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6056845/]
- 71. The Limitations of In Vitro Experimentation ... [https://www.sciencedirect.com/science/article/abs/pii/S0022283615004908]
- 72. CRISPR-based diagnostics | Nature Biomedical Engineering [https://www.nature.com/articles/s41551-021-00760-7]
- 73. CRISPR for detection of drug resistance genes [https://www.sciencedirect.com/science/article/abs/pii/S0009898125005054]
- 74. Modern Microbiological Methods to Detect Biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11205407/]
- 75. Cost-benefit analysis of antibiofilm microbiological ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5879767/]
- 76. Frontiers | Advances in the Application of CRISPR Technology in... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1645699/abstract]
- 77. CRISPR-Based Diagnostics Market Size to Hit USD 15.14 ... [https://www.precedenceresearch.com/crispr-based-diagnostics-market]
- 78. Real-time Detection of Foodborne Pathogens and Biofilm ... [https://www.sciencedirect.com/science/article/pii/S0362028X25000638]