CRISPR-Cas Systems and Quorum Sensing: Decoding the Regulatory Network Controlling Bacterial Biofilm Formation and Its Therapeutic Potential
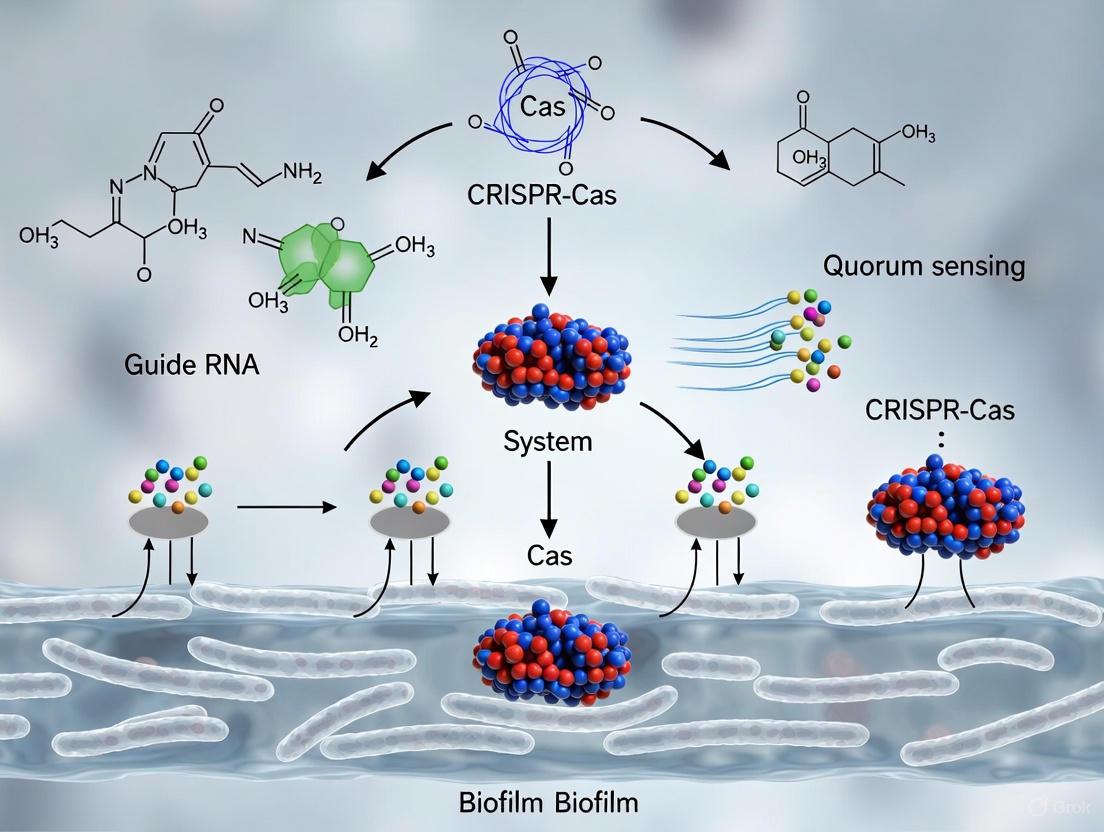
This article synthesizes current research on the intricate bidirectional relationship between CRISPR-Cas systems and quorum sensing (QS) in regulating bacterial biofilm formation, a key driver of antimicrobial resistance.
CRISPR-Cas Systems and Quorum Sensing: Decoding the Regulatory Network Controlling Bacterial Biofilm Formation and Its Therapeutic Potential
Abstract
This article synthesizes current research on the intricate bidirectional relationship between CRISPR-Cas systems and quorum sensing (QS) in regulating bacterial biofilm formation, a key driver of antimicrobial resistance. We explore the foundational biology, where QS signaling upregulates CRISPR-Cas expression to enhance bacterial immunity at high cell densities, and conversely, where specific Cas proteins directly modulate biofilm architecture and virulence. The review critically assesses innovative methodological approaches, including CRISPR-Cas9 and CRISPRi for precision targeting of QS and biofilm genes, and the synergistic use of nanoparticle delivery systems to enhance efficacy. We further troubleshoot challenges such as off-target effects and delivery optimization, and provide a comparative validation of emerging technologies against traditional methods. Aimed at researchers and drug development professionals, this analysis highlights the transformative potential of targeting the CRISPR-QS axis to develop next-generation, resistance-proof antimicrobial therapies against persistent biofilm-associated infections.
The Molecular Dialogue: How Quorum Sensing Regulates CRISPR and CRISPR Influences Biofilms
Quorum Sensing as a Master Regulator of CRISPR-Cas Immune Defenses
In nature, bacteria frequently exist in high-density populations such as biofilms and microcolonies, making them particularly vulnerable to viral predation and horizontal gene transfer (HGT). To manage this increased risk, bacteria have evolved a sophisticated regulatory mechanism wherein quorum sensing (QS)—a cell-cell communication system—controls the expression and activity of CRISPR-Cas adaptive immune systems. This regulatory integration allows bacterial populations to coordinate their defensive strategies based on cell density, enhancing immunity when the threat of infection is highest while minimizing the fitness costs of constitutive defense system expression. The molecular dialogue between QS and CRISPR-Cas represents a sophisticated level of population-level regulation in prokaryotes, demonstrating that bacteria temporally control their immune investments according to perceived risk [1] [2].
This technical guide examines the mechanistic basis of QS-mediated CRISPR-Cas regulation, detailing the molecular pathways, experimental evidence, and physiological significance of this relationship. Understanding this crosstalk is crucial for researchers investigating bacterial pathogenesis, microbial ecology, and novel antimicrobial strategies that exploit these regulatory networks [3].
Molecular Mechanisms of Quorum Sensing
Fundamental QS Principles Across Bacterial Species
Quorum sensing is a cell-density dependent communication system that enables bacteria to coordinate gene expression collectively. The process follows three fundamental principles: (1) production of extracellular signaling molecules called autoinducers (AIs), (2) accumulation and detection of these AIs when a critical threshold concentration is reached at high cell density, and (3) activation of response regulators that reprogram gene expression to control collective behaviors [2].
Table 1: Major Quorum Sensing Systems in Model Organisms
| Bacterial Species | QS System | Signaling Molecule | Regulatory Components | Key Controlled Functions |
|---|---|---|---|---|
| Serratia sp. ATCC39006 | LuxI/LuxR-type | C4-HSL (N-butanoyl-L-homoserine lactone) | SmaI (synthase), SmaR (receptor) | CRISPR-Cas expression, secondary metabolites, motility |
| Pseudomonas aeruginosa | LasI/LasR | 3O-C12-HSL | LasI (synthase), LasR (receptor) | Virulence factors, secretion systems, biofilm formation |
| Pseudomonas aeruginosa | RhlI/RhlR | C4-HSL | RhlI (synthase), RhlR (receptor) | Virulence factors, rhamnolipids, secondary metabolites |
| Staphylococcus aureus | Agr | Autoinducing peptide (AIP) | AgrD (pro-AIP), AgrB (processing/export), AgrC (receptor), AgrA (response regulator) | Virulence factors, toxin production |
| Vibrio cholerae | Two-component | AI-2 | LuxS, LuxPQ, LuxU, LuxO | Virulence factors, biofilm formation |
Gram-negative bacteria typically use acyl-homoserine lactones (AHLs) as signaling molecules, which are produced by LuxI-family synthases and detected by cytoplasmic LuxR-type transcription factors. In Serratia sp. ATCC39006, the LuxI/LuxR homologs SmaI and SmaR control QS-dependent behaviors, with SmaI producing predominantly C4-HSL [1]. In contrast, Gram-positive bacteria generally use processed oligopeptides (autoinducing peptides, AIPs) as signals that are detected by membrane-bound two-component histidine kinase receptors [2].
QS Regulatory Dynamics
The Serratia QS circuit operates primarily through a de-repression mechanism. In the absence of sufficient AHL signals at low cell density, the transcriptional regulator SmaR acts as a repressor of QS-controlled genes. As cell density increases and AHLs accumulate, these signaling molecules bind to SmaR, inhibiting its DNA-binding activity and thereby derepressing gene expression. This mechanism ensures that energy-intensive processes like adaptive immunity are only activated when population density justifies collective action [1].
Figure 1: Quorum Sensing Regulation Mechanism in Serratia. At low cell density (left), SmaR repressor binds DNA and inhibits CRISPR-Cas expression. At high cell density (right), accumulated AHL molecules bind SmaR, preventing DNA binding and derepressing CRISPR-Cas systems.
Experimental Evidence for QS Regulation of CRISPR-Cas
Transcriptional Control of CRISPR-Cas Systems by QS
Groundbreaking research in Serratia sp. ATCC39006 demonstrated that QS directly regulates the expression of multiple CRISPR-Cas systems. This bacterium possesses three functional CRISPR-Cas systems (types I-E, I-F, and III-A), each with associated CRISPR arrays [1].
Key Experimental Findings:
- Expression analysis throughout growth revealed that cas operon expression for all three CRISPR-Cas systems, along with CRISPR1 (type I-E) and CRISPR2 (type I-F) arrays, was significantly reduced in an smaI mutant unable to produce AHL signals
- Complementation studies restored wild-type expression levels by adding synthetic C4-HSL to smaI mutant cultures, confirming the specific role of AHL signaling
- Repressor function was demonstrated through smaR deletion, which restored CRISPR-Cas expression in the smaI mutant background, and by plasmid-based SmaR expression, which reduced promoter activity from QS-regulated CRISPR and cas promoters
- The type III-A system-associated arrays (CRISPR3 and CRISPR4) showed low expression in wild-type cells and were not significantly regulated by QS [1]
Table 2: QS Regulation of CRISPR-Cas Components in Serratia
| CRISPR-Cas System | Component | Fold Reduction in smaI mutant | Complementation by C4-HSL | SmaR Repression |
|---|---|---|---|---|
| Type I-E | cas operon | Significant reduction | Yes | Confirmed |
| Type I-E | CRISPR1 array | Significant reduction | Yes | Confirmed |
| Type I-F | cas operon | Significant reduction | Yes | Confirmed |
| Type I-F | CRISPR2 array | Significant reduction | Yes | Confirmed |
| Type III-A | cas operon | Significant reduction | Yes | Confirmed |
| Type III-A | CRISPR3/4 arrays | No significant change | No | Not applicable |
Functional Impact on Interference and Adaptation
Beyond transcriptional control, QS signaling significantly affects the functional capabilities of CRISPR-Cas systems, including both interference (target destruction) and adaptation (spacer acquisition).
Interference Assays: Researchers measured interference efficiency by exposing Serratia to conjugative plasmids containing sequences targeted by spacers in each CRISPR-Cas system. Wild-type and QS-deficient strains were compared for their ability to prevent plasmid acquisition [1].
Table 3: Impact of QS on CRISPR-Cas Interference Efficiency
| CRISPR-Cas System | Interference Target | Reduction in smaI mutant | Rescue by Exogenous AHL |
|---|---|---|---|
| Type I-E | Plasmid with cognate protospacer | ~20-fold | Yes |
| Type I-F | Plasmid with cognate protospacer | ~500-fold | Yes |
| Type III-A | Plasmid with cognate protospacer | ~240-fold | Yes |
The variation in QS-dependence among systems suggests different regulatory thresholds or additional regulatory bottlenecks. For instance, the type I-E system showed the weakest QS effect on interference despite strong QS regulation of its cas8e promoter, possibly due to limiting amounts of other components like Cas3 [1].
Adaptation Assays: Spacer acquisition—the process of incorporating new sequences from invaders into CRISPR arrays—was also impaired in QS-deficient strains for the type I-E and I-F systems. This demonstrates that QS controls not only the expression of interference components but also the ability to develop new immunities [1].
Detailed Experimental Protocols
Protocol 1: Assessing QS Regulation of CRISPR-Cas Expression
Objective: Measure transcriptional activity of cas genes and CRISPR arrays in wild-type versus QS-deficient mutants throughout growth.
Materials:
- Bacterial strains: Wild-type Serratia sp. ATCC39006, isogenic smaI mutant, smaR mutant, and smaI smaR double mutant
- Growth medium: Appropriate rich or defined medium
- AHL solution: 100 μM synthetic C4-HSL in acidified ethyl acetate
- Reporter constructs: Transcriptional fusions of cas/CRISPR promoters to reporter genes
- RNA extraction kit, DNase I, reverse transcription reagents, qPCR reagents
Method:
- Inoculate parallel cultures of each strain and grow with shaking at appropriate temperature
- For complementation tests, add C4-HSL to smaI mutant cultures at inoculation
- Monitor growth by OD600 and collect samples at multiple time points (early exponential, mid-exponential, late exponential, early stationary, stationary)
- Extract total RNA, treat with DNase I, and verify RNA quality
- Perform reverse transcription and quantitative PCR using primers specific for:
- cas operon mRNAs (I-E, I-F, III-A)
- CRISPR array primary transcripts
- Reference housekeeping genes
- For promoter activity assays, measure reporter gene activity throughout growth
Analysis:
- Calculate relative expression using the 2^(-ΔΔCt) method
- Compare expression kinetics across strains and growth phases
- Statistical analysis (ANOVA) of expression differences between wild-type and mutants at equivalent growth phases [1]
Protocol 2: Interference Efficiency Assays
Objective: Quantify the ability of CRISPR-Cas systems to prevent acquisition of targeted plasmids in QS-proficient versus QS-deficient backgrounds.
Materials:
- Donor strain: Conjugation-proficient strain carrying target plasmid with appropriate protospacer and PAM sequence
- Recipient strains: Wild-type and QS-mutant Serratia with functional CRISPR-Cas systems
- Control plasmids: Non-targeted plasmids with different selection markers
- Conjugation filters, selective media, AHL solutions
Method:
- Grow donor and recipient strains to appropriate density
- Mix donor and recipient cells at optimized ratios, concentrate by filtration, and place filters on solid medium
- Incubate for conjugation (typically 4-8 hours)
- Resuspend cells, plate serial dilutions on selective media to count transconjugants
- Include controls with non-targeted plasmids to establish baseline conjugation efficiency
- For AHL rescue experiments, add C4-HSL to both growth and conjugation steps for QS-deficient mutants
Analysis:
- Calculate conjugation efficiency as transconjugants per recipient
- Determine interference efficiency as the reduction in conjugation for targeted versus non-targeted plasmids
- Compare interference efficiency between wild-type and QS-mutant strains [1]
Research Reagent Solutions
Table 4: Essential Research Reagents for QS-CRISPR Studies
| Reagent Category | Specific Examples | Function/Application |
|---|---|---|
| Bacterial Strains | Serratia sp. ATCC39006 wild-type and isogenic QS mutants (smaI, smaR, smaI smaR) | Principal model system for studying QS-CRISPR interactions |
| Signaling Molecules | Synthetic C4-HSL, 3O-C12-HSL, other AHLs | QS complementation studies, signal response profiling |
| QS Inhibitors | Baicalein, other quorum quenching enzymes | Inhibit QS signaling to validate regulatory relationships |
| Target Plasmids | Conjugative plasmids with protospacer sequences matching CRISPR spacers | Interference efficiency assays, requires appropriate PAM |
| Reporter Systems | Transcriptional fusions of cas promoters to luciferase or fluorescent proteins | Quantify promoter activity in different QS backgrounds |
| CRISPR Components | Cas protein antibodies, crRNA detection probes | Validate expression at protein and RNA levels |
Complex Regulatory Networks in Pathogens
The relationship between QS and CRISPR-Cas extends beyond Serratia to other bacterial pathogens, though with species-specific variations. In Pseudomonas aeruginosa, which possesses the LasI/LasR and RhlI/RhlR QS systems that produce 3O-C12-HSL and C4-HSL respectively, QS regulates multiple virulence factors and secretion systems [2] [4].
Interestingly, studies using the QS inhibitor baicalein in P. aeruginosa revealed unexpected complexity. Contrary to initial predictions that QS inhibition would decrease CRISPR-mediated immunity, treatment actually increased bacterial resistance to phage infection in some contexts. This suggests that QS inhibition can indirectly affect CRISPR immunity by altering other bacterial processes, such as type IV pilus expression, which subsequently impacts phage adsorption and CRISPR system evolution [3].
This paradoxical effect highlights the complexity of bacterial regulatory networks, where QS connects CRISPR-Cas systems with other cellular processes including:
- Biofilm formation: QS controls matrix production that protects bacterial communities
- Secretion systems: Multiple type I-VI secretion systems are QS-regulated
- Metabolic adaptation: QS optimizes resource allocation in dense populations
- Stress response: QS coordinates population-level stress tolerance [2] [4]
Figure 2: Integrated Regulatory Network of QS and CRISPR-Cas. Quorum sensing centrally regulates multiple CRISPR-Cas systems alongside other virulence and fitness functions, creating a complex network that responds to threats like phage infection and horizontal gene transfer.
The regulatory connection between quorum sensing and CRISPR-Cas systems represents a sophisticated bacterial strategy for optimizing immune function according to population density and infection risk. By activating CRISPR-Cas defenses primarily at high cell densities, bacteria balance the substantial fitness costs of maintaining these systems against their protective benefits [1].
This relationship has important implications for both basic microbiology and applied research:
- Microbial Ecology: Understanding how bacterial communities regulate defenses informs models of phage-bacteria coevolution
- Antimicrobial Development: QS inhibitors could potentially modulate CRISPR-Cas activity as part of combination therapies
- Biotechnology: Harnessing QS regulation could improve CRISPR-based biocontainment in industrial applications
- Medical Microbiology: Understanding these pathways may reveal new targets for combating antibiotic-resistant pathogens
Future research should focus on elucidating the complete regulatory networks connecting QS to CRISPR-Cas across diverse bacterial species, examining how these systems integrate with other defense mechanisms, and exploring translational applications that exploit this knowledge for antimicrobial development and biotechnology.
In the broader context of bacterial pathogenesis, the interplay between CRISPR-Cas systems and quorum sensing (QS) represents a sophisticated regulatory node through which bacteria coordinate defensive immunity with population density. This technical guide elucidates the molecular mechanism by which the SmaR repressor protein and acyl-homoserine lactone (AHL) autoinducers govern the expression of CRISPR-Cas operons in the model bacterium Serratia sp. ATCC39006. We detail how SmaR-mediated repression is alleviated at high cell density, leading to the de-repression of type I-E, I-F, and III-A CRISPR-Cas systems, thereby enhancing bacterial defense against mobile genetic elements. This overview, supported by quantitative data, experimental protocols, and pathway visualizations, provides researchers and drug development professionals with a framework for understanding this critical regulatory intersection, which influences both antiviral immunity and community behaviors such as biofilm formation.
Quorum Sensing (QS) is a widespread cell-cell communication mechanism that allows bacteria to collectively modify gene expression in response to population density, typically through the accumulation of small diffusible signal molecules like AHLs [5]. This system regulates diverse processes including virulence factor production, biofilm formation, and antibiotic resistance [5] [6]. Concurrently, CRISPR-Cas systems provide prokaryotes with adaptive, sequence-specific immunity against invading mobile genetic elements such as bacteriophages and plasmids [1] [7].
The convergence of these pathways—whereby bacteria use chemical communication to modulate their immune defenses—represents a sophisticated adaptation to balance the fitness costs of constitutive CRISPR-Cas expression against the increased risk of infection at high cell densities [1]. In Serratia, this integration involves the LuxIR-type QS system (comprising smaI and smaR) directly controlling the transcriptional activity of CRISPR-Cas operons and arrays [1].
Molecular Mechanism of SmaR Repression and AHL-Mediated De-repression
The Core Regulatory Components
The QS circuitry in Serratia centers on two key elements: SmaI, a LuxI-family AHL synthase that produces primarily N-butanoyl-L-homoserine lactone (C4-HSL), and SmaR, a LuxR-type transcriptional regulator that functions as a DNA-binding repressor [1]. In the absence of sufficient AHL signal, SmaR binds to promoter regions of target genes, including those encoding CRISPR-Cas systems, and represses their transcription [1] [8].
The De-repression Switch
As bacterial cell density increases, AHLs produced by SmaI accumulate in the environment. Upon reaching a critical threshold concentration, these signaling molecules bind directly to SmaR, inducing a conformational change that inhibits its DNA-binding activity [1]. This AHL-mediated inactivation of SmaR results in de-repression of the CRISPR-Cas loci, allowing elevated expression of cas genes and CRISPR arrays when the population is most vulnerable to viral predation and horizontal gene transfer [1]. This mechanism is summarized in the diagram below.
Diagram Title: SmaR Repression and AHL-Mediated De-repression of CRISPR-Cas
Genetic Evidence for the Repression Mechanism
Genetic analyses demonstrate that mutation of smaI (AHL-deficient) significantly reduces expression of cas operons and CRISPR arrays, whereas mutation of smaR alone has little effect [1]. Crucially, a double smaI smaR mutant restores CRISPR-Cas expression to wild-type levels, confirming that SmaR acts as a repressor whose activity requires the absence of AHL signaling [1]. Furthermore, heterologous expression of SmaR in trans significantly reduces promoter activity from QS-regulated CRISPR and cas promoters, but not from non-QS regulated control promoters [1].
Quantitative Analysis of QS-Dependent CRISPR-Cas Regulation
Transcriptional Regulation of cas Operons and CRISPR Arrays
Quantitative assessments of transcript levels throughout bacterial growth reveal that QS regulation differentially affects the three CRISPR-Cas systems in Serratia. The table below summarizes the expression changes observed in a signaling-deficient smaI mutant compared to the wild-type strain.
Table 1: Expression of CRISPR-Cas Components in smaI Mutant vs. Wild-Type Serratia
| CRISPR-Cas System | Component | Expression in smaI mutant | QS Regulation |
|---|---|---|---|
| Type I-E | cas operon | Significantly reduced [1] | Yes |
| CRISPR1 array | Significantly reduced [1] | Yes | |
| Type I-F | cas operon | Significantly reduced [1] | Yes |
| CRISPR2 array | Significantly reduced [1] | Yes | |
| Type III-A | cas operon | Significantly reduced [1] | Yes |
| CRISPR3 array | Low expression, no further reduction [1] | No | |
| CRISPR4 array | Low expression, no further reduction [1] | No |
Functional Consequences for Immunity
The transcriptional regulation by QS directly impacts CRISPR-Cas immune function, affecting both interference and adaptation capabilities. The defense impairment in QS-deficient mutants can be rescued by adding exogenous synthetic C4-HSL [1]. The table below quantifies the reduction in interference efficiency against targeted plasmids in the smaI mutant.
Table 2: Interference Efficiency Against Targeted Plasmids in smaI Mutant
| CRISPR-Cas System | Target | Reduction in Interference (smaI vs WT) | Rescue with Exogenous C4-HSL |
|---|---|---|---|
| Type I-E | Plasmid with CRISPR1 protospacer | ~20-fold [1] | Yes [1] |
| Type I-F | Plasmid with CRISPR2 protospacer | ~500-fold [1] | Yes [1] |
| Type III-A | Plasmid with CRISPR3 protospacer | ~240-fold [1] | Yes [1] |
Additionally, the acquisition of new spacers (adaptation) by the type I-E and I-F systems is impaired in the absence of QS signaling, further demonstrating that QS modulates both the establishment and execution of CRISPR immunity [1].
Experimental Protocols for Investigating SmaR-CRISPR Regulation
Assessing CRISPR-Cas Expression Dynamics
Protocol 1: Transcriptional Analysis of cas Operons and CRISPR Arrays
- Objective: Quantify expression of CRISPR-Cas components throughout bacterial growth in wild-type, smaI, smaR, and smaI smaR double mutant strains.
- Methodology:
- Culture Conditions: Grow bacterial strains in appropriate liquid medium, sampling at regular intervals corresponding to different growth phases (early exponential, late exponential, stationary) [1].
- RNA Extraction: Isolve total RNA from samples, ensuring removal of genomic DNA.
- Reverse Transcription Quantitative PCR (RT-qPCR): Design primers specific to cas operons (e.g., cas8e for type I-E) and CRISPR arrays (CRISPR1, CRISPR2) [1]. Include control genes not regulated by QS.
- Data Analysis: Normalize expression to control genes and compare transcript levels between strains and growth phases.
- Key Controls:
Functional Interference Assays
Protocol 2: Conjugation-Based Interference Assay
- Objective: Measure the functional capacity of QS-regulated CRISPR-Cas systems to defend against invading DNA.
- Methodology:
- Donor Strain Preparation: Use donor bacteria (e.g., E. coli) carrying a conjugative plasmid with a protospacer sequence matching a spacer in the Serratia CRISPR array (e.g., first spacer of CRISPR1 for type I-E) and including the appropriate Protospacer Adjacent Motif (PAM) [1].
- Recipient Strains: Grow wild-type and QS mutant Serratia strains to high cell density.
- Conjugation: Co-culture donor and recipient strains for a fixed period to allow plasmid transfer.
- Selection and Quantification: Plate conjugations on selective media to count recipient Serratia cells that have received either the targeted plasmid or a non-targeted control plasmid.
- Interference Calculation: Interference efficiency is calculated as the reduction in conjugation frequency for the targeted plasmid compared to the control plasmid [1].
- Key Parameters:
- Use multiple biological replicates.
- Confirm the presence of the correct PAM sequence for the relevant CRISPR-Cas type [1].
The workflow for these key experiments is visualized below.
Diagram Title: Key Experimental Workflows for Studying QS-CRISPR Regulation
The Scientist's Toolkit: Essential Research Reagents
The following table catalogs crucial reagents and genetic tools for investigating the SmaR-AHL-CRISPR regulatory axis.
Table 3: Essential Research Reagents for SmaR-AHL-CRISPR Studies
| Reagent / Tool | Function and Application | Specific Example / Source |
|---|---|---|
| smaI Mutant Strain | AHL-signaling deficient mutant; used to assess QS-dependence of CRISPR-Cas expression and function [1]. | Serratia sp. ATCC39006 smaI::KnR [1]. |
| smaR Mutant Strain | Lacks the SmaR repressor; used to confirm repressor function, often in combination with smaI mutation [1]. | Serratia sp. ATCC39006 smaR::CmR [1]. |
| Synthetic C4-HSL | Pure AHL autoinducer (N-butanoyl-L-homoserine lactone); used for complementation experiments to restore signaling in smaI mutants [1]. | Commercially available from various chemical suppliers (e.g., Sigma-Aldrich). |
| Conjugative Plasmid with Protospacer | Plasmid carrying a sequence matching a Serratia CRISPR spacer and the correct PAM; serves as target for interference assays [1]. | e.g., Plasmid with protospacer matching first spacer of CRISPR1 (type I-E) [1]. |
| Heterologous SmaR Expression Plasmid | Plasmid for in trans expression of SmaR; used to demonstrate direct repression of CRISPR-Cas promoters [1]. | e.g., pACYC-derived plasmid with inducible smaR [1]. |
| AHL Biosensor Strains | Reporter strains that produce a detectable output (e.g., violacein pigment) in response to AHLs; used to quantify AHL production [6]. | Chromobacterium violaceum CV026 (responds to short-chain AHLs like C4-HSL) [6]. |
The precise mechanistic understanding of SmaR repression and AHL-mediated de-repression illustrates a sophisticated evolutionary adaptation where bacteria optimize resource allocation by activating costly CRISPR-Cas defenses only when population density indicates a heightened risk of infection. This regulatory paradigm extends beyond Serratia, with QS systems in pathogens like Pseudomonas aeruginosa also modulating CRISPR-Cas activity [9] [8].
From a therapeutic perspective, this intersection offers novel targets. For instance, QS inhibitors could potentially be used to modulate bacterial immunity [9] [6], while engineered CRISPR systems could themselves be designed to disrupt QS pathways controlling virulence and biofilm formation [10] [11]. A deep understanding of the SmaR-AHL-CRISPR mechanism thus not only advances fundamental microbial ecology but also paves the way for innovative anti-infective strategies that exploit the intricate regulatory networks governing bacterial sociality and defense.
While CRISPR-Cas systems are recognized as prokaryotic adaptive immune systems, emerging research reveals their significant non-canonical functions in regulating bacterial pathogenesis. This technical review examines the role of Cas3, the signature enzyme of Type I CRISPR-Cas systems, as a central modulator of bacterial virulence and biofilm formation. We synthesize recent findings demonstrating that Cas3 in Acinetobacter baumannii and other pathogens directly influences the expression of virulence factors, biofilm-related genes, and metabolic pathways. Furthermore, we explore the intricate regulatory networks connecting CRISPR-Cas systems with quorum sensing (QS) mechanisms, revealing complex bidirectional relationships that shape bacterial behavior. The evidence presented establishes Cas3 as a critical factor balancing immune defense and pathogenicity, offering novel perspectives for therapeutic intervention against multidrug-resistant bacterial infections.
CRISPR-Cas systems represent adaptive immune mechanisms widespread in bacteria and archaea, providing sequence-specific protection against invasive genetic elements [12]. These systems are categorized into two classes and six primary types based on their effector complexes and mechanisms of action [13]. Type I systems, the most prevalent in bacteria, are characterized by the multifunctional Cas3 protein which possesses both helicase and DNase activities [14].
Beyond their established role in immunity, CRISPR-Cas systems are increasingly recognized for their non-canonical functions in regulating fundamental bacterial processes [12] [15]. Cas3 has emerged as a particularly significant regulator of bacterial pathogenicity, influencing biofilm formation, virulence factor expression, and host-pathogen interactions. In Acinetobacter baumannii, a notorious multidrug-resistant pathogen, Cas3 directly modulates bacterial virulence and pathogenicity, positioning it as a crucial factor in disease progression [14] [16].
This review examines the molecular mechanisms through which Cas3 influences bacterial virulence and biofilm formation, explores its integration with QS networks, and discusses the therapeutic implications of these findings for combating resistant bacterial infections.
Molecular Mechanisms of Cas3 in Virulence Regulation
Cas3-Dependent Modulation of Virulence Factors
Research using A. baumannii ATCC19606 has demonstrated that deletion of the cas3 gene (type I-Fa system) significantly impairs bacterial virulence through multiple mechanisms [14] [16]. The molecular pathways affected include:
- Downregulation of outer membrane protein A (OmpA): An essential virulence factor involved in immune evasion and host cell adhesion [14]
- Alteration of carbon metabolism and oxidative phosphorylation pathways: Impacting bacterial energy production and stress adaptation [14] [16]
- Reduced expression of biofilm-associated factors: Including poly-N-acetylglucosamine (PNAG) and pilus assembly components [17]
Mechanistically, transcriptome sequencing analysis revealed that cas3 deletion leads to significant differential expression of virulence-associated genes, while paradoxically upregulating other cas genes within the CRISPR-Cas system except for cas6 [16].
Hierarchical Regulatory Axis Controlling Cas3 Expression
The expression and activity of Cas3 in A. baumannii I-Fb systems are governed by a sophisticated regulatory network involving key transcriptional regulators:
Table 1: Regulatory Proteins Governing Cas3 Expression in A. baumannii
| Regulator | Function | Effect on Cas3 | Mechanism |
|---|---|---|---|
| H-NS | Histone-like nucleoid structuring protein | Repression | Directly binds AT-rich regions in cas3 promoter [17] |
| BaeR | Response regulator of two-component system | Indirect repression | Positively regulates H-NS expression [17] |
| LeuO/StpA | Transcriptional antagonists | Activation | Counteracts H-NS-mediated silencing [17] |
DNA pull-down and electrophoretic mobility shift assays (EMSA) have confirmed direct binding of H-NS to the cas3 promoter region, establishing a repressive effect on both interference activity and adaptive immunity functions of the I-Fb CRISPR-Cas system [17]. This hierarchical regulation (BaeR→H-NS→Cas3) represents a sophisticated control mechanism that fine-tunes Cas3 expression in response to environmental conditions.
Experimental Evidence: Methodologies and Key Findings
Genetic Manipulation and Phenotypic Characterization
To elucidate Cas3 functions, researchers have employed comprehensive genetic and phenotypic approaches:
Strain Construction and Validation:
- Creation of isogenic mutant strains: 19606Δcas3 (cas3 deletion) and 19606Δcas3/pcas3 (complemented strain) in A. baumannii ATCC19606 [14]
- Verification via PCR and sequencing to confirm genetic manipulations [14]
- Construction of double-knockout strains (Δh-ns-cas3 and ΔbaeR-cas3) to dissect regulatory networks [17]
Growth and Viability Assessment:
- Standard bacterial growth curves monitored via OD600 and viable cell counts over 24 hours
- Confirmation that cas3 deletion does not significantly affect basal growth kinetics [14] [16]
Quantitative Analysis of Virulence-Associated Phenotypes
Table 2: Experimental Assessment of Cas3-Mediated Virulence Modulation
| Experimental Approach | Key Findings | Significance |
|---|---|---|
| Biofilm Quantification (Crystal Violet Staining) | Significant reduction in biofilm formation in Δcas3 mutants; complementation restored biofilm formation [14] [16] | Demonstrates Cas3 requirement for robust biofilm development |
| Confocal Laser Scanning Microscopy | Reduced fluorescence intensity and thickness in Δcas3 biofilms; EPS labeled with Alexa Fluor 647-dextran, cells with SYTO9 [14] | Visualizes structural defects in biofilm architecture |
| Host Cell Adhesion/Invasion Assays | Significant reduction in adhesion and invasion rates of Δcas3 mutants in A549 human alveolar epithelial cells (MOI=100) [14] | Indicates impaired host-pathogen interaction capabilities |
| Galleria mellonella Infection Model | 90% mortality with WT at 12h vs. 20% with Δcas3; 50% survival with Δcas3 at 96h vs. 0% with WT [14] [16] | Demonstrates attenuated virulence in vivo |
| Murine Systemic Infection Model | Significant reduction in organ bacterial loads, lung inflammation, and serum cytokines (IL-1β, IL-6, TNF-α) in Δcas3-infected mice [14] [16] | Confirms virulence attenuation in mammalian model |
The consistency of findings across multiple infection models and phenotypic assays strengthens the conclusion that Cas3 functions as a positive regulator of virulence in A. baumannii.
Intersection with Quorum Sensing Systems
The relationship between CRISPR-Cas systems and QS represents a complex bidirectional regulatory network that coordinates bacterial behavior with population density.
Contrasting Regulatory Paradigms in Gram-Positive and Gram-Negative Bacteria
Table 3: Quorum Sensing and CRISPR-Cas System Interactions
| Bacterial Species | QS System | Effect on CRISPR-Cas | Molecular Mechanism |
|---|---|---|---|
| Staphylococcus aureus (Type III-A) | Agr system | Repression at high cell density [18] [19] | AgrA binds sarA and arcR promoters, repressing positive regulators of cas gene transcription |
| Serratia & Pseudomonas aeruginosa | LuxI/R-type | Activation [18] | Enhanced CRISPR adaptation and interference activity |
| Burkholderia glumae | LuxI/R-type | Activation [18] | Increased cas gene expression |
In S. aureus, the QS-mediated repression of CRISPR-Cas activity occurs through a precisely defined mechanism: the QS regulator AgrA directly binds to promoters of two transcriptional regulators (sarA and arcR), inhibiting their expression [18] [19]. Since both SarA and ArcR function as positive regulators that promote transcription of cas genes by binding to a novel promoter Pcas, their QS-mediated repression results in decreased cas gene expression and reduced CRISPR-Cas activity at high cell density.
Biological Implications of QS-CRISPR Interplay
This density-dependent regulation may serve important biological functions:
- Facilitation of horizontal gene transfer: At high cell density, repressed CRISPR-Cas activity may allow increased acquisition of antibiotic resistance or virulence genes [18]
- Resource allocation: Redirecting cellular resources from immune defense to virulence factor production during infection [15]
- Population-level coordination: Enabling synchronized behavioral shifts across bacterial communities
The contrasting effects observed in different bacterial species highlight the context-dependent nature of CRISPR-Cas regulation and its integration with species-specific regulatory networks.
Visualization of Key Regulatory Networks
BaeR-H-NS-Cas3 Regulatory Axis in A. baumannii
Diagram Title: BaeR-H-NS-Cas3 Regulatory Hierarchy
Quorum Sensing Inhibition of CRISPR-Cas in S. aureus
Diagram Title: QS Inhibition of Type III-A CRISPR in S. aureus
The Scientist's Toolkit: Essential Research Reagents and Methodologies
Table 4: Key Experimental Resources for Cas3-Virulence Research
| Reagent/Technique | Application | Specific Example/Protocol |
|---|---|---|
| Isogenic Mutant Strains | Genetic dissection of Cas3 function | A. baumannii 19606Δcas3 and complemented strain 19606Δcas3/pcas3 [14] |
| A549 Human Alveolar Epithelial Cells | Adhesion and invasion assays | Infection at MOI=100; quantification of adherent/invaded bacteria [14] |
| Galleria mellonella Larvae | Intermediate virulence model | Injection with 1.0×10^6 CFU; survival monitoring over 96 hours [14] [16] |
| Murine Systemic Infection Model | Comprehensive virulence assessment | Intraperitoneal injection; bacterial load quantification in organs, cytokine measurement (IL-1β, IL-6, TNF-α) [14] [16] |
| Confocal Laser Scanning Microscopy with Fluorescent Staining | Biofilm architecture analysis | SYTO9 (bacterial cells) and Alexa Fluor 647-dextran (EPS matrix) [14] |
| DNA Pull-Down & EMSA | Protein-DNA interaction studies | H-NS binding to AT-rich regions of cas3 promoter [17] |
| RNA Sequencing | Transcriptome analysis | Identification of differentially expressed genes in Δcas3 mutants [16] |
Discussion and Therapeutic Implications
The emerging role of Cas3 as a virulence modulator presents intriguing possibilities for novel antibacterial strategies. The BaeR-H-NS-Cas3 regulatory axis offers multiple potential intervention points:
- Small molecule inhibitors targeting BaeR-H-NS interaction to enhance endogenous CRISPR-Cas activity against resistance genes
- QS interferents that manipulate the density-dependent regulation of CRISPR-Cas systems
- Cas3-targeted approaches that exploit its dual function in immunity and virulence regulation
The contrasting effects observed between different CRISPR-Cas types (I-Fa vs. I-Fb) and bacterial species highlight the importance of context-specific understanding when developing therapeutic interventions [14] [17]. Furthermore, the inverse relationship between CRISPR-Cas activity and biofilm formation in some pathogens suggests potential approaches for sensitizing bacteria to conventional antibiotics.
Future research should focus on elucidating the precise molecular mechanisms through which Cas3 influences virulence gene expression, particularly the potential involvement of endogenous gene targeting or indirect regulatory networks. Additionally, comprehensive studies across diverse bacterial pathogens will establish whether Cas3's role as a virulence modulator represents a conserved function or species-specific adaptation.
Cas3 has definitively emerged as a multifunctional protein that extends far beyond its canonical role in CRISPR-Cas immunity. Through direct regulation of virulence factors, biofilm components, and integration with QS networks, Cas3 represents a critical node in the pathogenicity programs of important human pathogens like A. baumannii and S. aureus. The sophisticated regulatory mechanisms governing Cas3 expression and activity, particularly the BaeR-H-NS axis, reveal how bacteria balance immune defense with virulence expression. These findings not only expand our fundamental understanding of bacterial pathogenesis but also unveil new therapeutic targets for combating multidrug-resistant infections through manipulation of endogenous bacterial defense and virulence systems.
The classical understanding of CRISPR-Cas systems as simple adaptive immune mechanisms in prokaryotes has been fundamentally transformed by the discovery of their deep integration with bacterial social behavior. This whitepaper synthesizes recent evidence establishing that a bidirectional crosstalk exists between CRISPR-Cas systems and environmental sensing mechanisms, particularly quorum sensing (QS) and biofilm development. We demonstrate that CRISPR-Cas systems not only provide defense but also actively regulate virulence and group behaviors by modulating QS pathways. Conversely, population density signals can precondition bacterial cells to alter CRISPR-Cas efficiency. This regulatory circuit has profound implications for bacterial pathogenesis, antimicrobial tolerance, and the development of novel therapeutic strategies that target this interplay. For drug development professionals, understanding this relationship opens avenues for anti-virulence therapies and biofilm-disrupting agents that could resensitize resistant infections to conventional antibiotics.
CRISPR-Cas systems, comprising clustered regularly interspaced short palindromic repeats and their associated proteins, function as adaptive immune systems in approximately 40% of bacteria and 80% of archaea [20]. These systems provide sequence-specific protection against invading mobile genetic elements like bacteriophages and plasmids through three distinct stages: adaptation, crRNA biogenesis, and interference [21]. However, beyond this canonical defense role, CRISPR-Cas systems are increasingly recognized as sophisticated regulatory nodes that integrate environmental information with population-level responses.
The conceptual framework of "Bidirectional Crosstalk" posits that CRISPR-Cas systems both influence and are influenced by community-level behaviors, primarily through interaction with quorum sensing networks. This whitepaper examines the molecular mechanisms underpinning this reciprocity, presents quantitative data establishing these relationships, details experimental methodologies for their investigation, and discusses therapeutic applications targeting this interplay. Within the broader thesis context of CRISPR's role in bacterial social behavior, this review establishes that CRISPR-Cas systems function as central processors in a complex network that connects defense, virulence, and community organization.
Molecular Mechanisms of CRISPR-QS Bidirectional Regulation
CRISPR-Mediated Regulation of Quorum Sensing and Biofilm Pathways
Evidence from multiple bacterial pathogens demonstrates that CRISPR-Cas components, particularly Cas proteins, can directly modulate the expression of genes central to quorum sensing and biofilm formation:
Direct Transcriptional Control: In Salmonella enterica serovar Enteritidis, Cas3 of the type I-E system regulates the lsr (luxS regulated) operon, a central component of the AI-2 quorum sensing pathway. Deletion of cas3 upregulates lsrFGBE genes while downregulating biofilm-forming genes and Salmonella pathogenicity island 1 (SPI-1) genes [22]. This establishes a direct molecular link between the CRISPR machinery and virulence regulation.
Virulence Factor Modulation: In Acinetobacter baumannii, deletion of cas3 (type I-Fa) significantly reduces biofilm formation and downregulates virulence factors including outer membrane protein A (OmpA) [14]. The Cas3-deficient strain exhibited reduced adhesion and invasion rates in A549 human alveolar epithelial cells, demonstrating the functional consequences of this regulatory relationship.
mRNA Targeting: In Pseudomonas aeruginosa, Cas3 targets the mRNA of the QS regulator LasR, dampening host recognition via Toll-like receptor 4 (TLR4) and consequently diminishing host defense and pro-inflammatory responses [22]. This mechanism represents a direct interface between CRISPR-mediated regulation and host-pathogen interactions.
Table 1: CRISPR-Mediated Regulation of Bacterial Social Behaviors
| CRISPR Component | Bacterial Species | Target Pathway/Genes | Functional Outcome |
|---|---|---|---|
| Cas3 (Type I-E) | Salmonella enterica | lsr operon (AI-2 QS), SPI-1 genes | Altered biofilm formation, virulence attenuation [22] |
| Cas3 (Type I-Fa) | Acinetobacter baumannii | OmpA, biofilm formation factors | Reduced biofilm, adhesion, and invasion [14] |
| Cas3 (Type I-F) | Pseudomonas aeruginosa | LasR mRNA (QS regulator) | Diminished host inflammatory response [22] |
| CRISPR-Cas System | Diverse bacteria | Carbon metabolism, oxidative phosphorylation | Enhanced survival under nutrient limitation [14] |
Environmental Preconditioning of CRISPR Efficiency
The bidirectional nature of this relationship is evidenced by how bacterial physiological states and environmental cues modulate CRISPR-Cas activity:
Metabolic Influence: In Acinetobacter baumannii, Cas3 participates in regulating carbon metabolism and oxidative phosphorylation pathways [14], suggesting that cellular metabolic status can feed back onto CRISPR-Cas function.
Epigenetic Modulation: The "CRISPR-Epigenetics Regulatory Circuit" model demonstrates that epigenetic landscapes substantially influence CRISPR editing efficiency. DNA methylation can impair Cas9 binding, particularly in highly methylated CpG islands, while histone modifications modulate chromatin accessibility [23].
Population Density Effects: Quorum sensing signals may precondition bacterial populations to optimize CRISPR-Cas defense when community density reaches critical thresholds, though the molecular mechanisms of this regulation require further elucidation.
The following diagram illustrates the core concept of bidirectional regulation between CRISPR-Cas systems and quorum sensing:
Quantitative Experimental Evidence
Phenotypic Changes in CRISPR-Modified Strains
Robust quantitative evidence supports the functional significance of CRISPR-QS crosstalk. The table below summarizes key phenotypic changes observed in Cas3-deficient strains across multiple bacterial species:
Table 2: Quantitative Phenotypic Changes in Cas3-Deficient Strains
| Bacterial Species | Biofilm Reduction | Adhesion/Invasion Defect | In Vivo Virulence Attenuation | Experimental Model |
|---|---|---|---|---|
| Acinetobacter baumannii | Significant reduction (p<0.05) [14] | Adhesion rate significantly reduced [14] | 50% survival in Galleria mellonella vs. 0% with WT [14] | Galleria mellonella, murine infection |
| Salmonella enterica | Downregulated biofilm-forming genes [22] | Reduced invasive and intracellular capacity [22] | Increased survival of infected chickens [22] | Chicken infection model |
| Pseudomonas aeruginosa | Not quantified | Not applicable | Diminished pro-inflammatory response [22] | Mouse models, cell culture |
Transcriptomic Alterations
RNA-Seq analysis of Salmonella Δcas3 mutant revealed comprehensive transcriptomic alterations, affecting:
- Upregulation of lsrFGBE genes in the lsr operon related to AI-2 quorum sensing
- Downregulation of biofilm-forming-related genes
- Suppression of Salmonella Pathogenicity Island 1 (SPI-1) genes encoding type III secretion system components [22]
In Acinetobacter baumannii, deletion of cas3 led to downregulation of virulence factors including outer membrane protein A (OmpA) and altered expression of genes in carbon metabolism and oxidative phosphorylation pathways [14].
Experimental Methodologies
Core Protocol: Establishing CRISPR-QS Relationships
To investigate bidirectional crosstalk between CRISPR-Cas systems and quorum sensing, researchers employ the following multidisciplinary approach:
Genetic Manipulation of CRISPR Components
- Strain Construction: Create isogenic mutant strains with deletions in key cas genes (e.g., cas3) using homologous recombination with counterselectable markers [14] [22].
- Complementation: Generate complemented strains by introducing wild-type cas genes on plasmids with inducible promoters to confirm phenotype specificity [14].
- Validation: Verify mutant and complemented strains through PCR amplification and sequencing of the modified genomic regions [14].
Phenotypic Characterization
- Biofilm Quantification: Assess biofilm formation using crystal violet staining to measure total biomass and confocal laser scanning microscopy (CLSM) with fluorescent staining (e.g., SYTO9 for cells, Alexa Fluor 647-dextran for EPS) to analyze 3D architecture and thickness [14].
- Virulence Assays:
- In vitro infection models using epithelial cell lines (e.g., A549 alveolar epithelial cells) to quantify adhesion and invasion rates at specific MOI [14].
- In vivo models including Galleria mellonella larvae and murine infection systems to assess survival rates, organ bacterial loads, and inflammatory responses [14] [22].
Molecular Analyses
- Transcriptomic Profiling: Conduct RNA-Seq analysis to identify differentially expressed genes following CRISPR component deletion [22].
- Pathway Analysis: Utilize bioinformatic tools to map altered gene expression to specific pathways, particularly quorum sensing, biofilm formation, and virulence networks [22].
The following workflow diagram outlines the key experimental steps:
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Investigating CRISPR-QS Crosstalk
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Genetic Tools | pBAD33-CM-cas3 complementation vector [14] | Functional complementation of mutant strains |
| Homologous recombination constructs | Targeted gene deletion | |
| Biofilm Assay Reagents | Crystal violet solution | Biofilm biomass quantification |
| SYTO9 green fluorescent nucleic acid stain | Bacterial cell staining in CLSM [14] | |
| Alexa Fluor 647-conjugated dextran | EPS matrix visualization in CLSM [14] | |
| Cell Culture Models | A549 human alveolar epithelial cells | Bacterial adhesion and invasion assays [14] |
| In Vivo Models | Galleria mellonella larvae | Initial virulence screening [14] |
| Murine infection models | Comprehensive pathogenicity assessment [14] | |
| Nanoparticle Systems | Liposomal Cas9 formulations [24] | Enhanced biofilm penetration and delivery |
| Gold nanoparticle-CRISPR hybrids [24] | Improved editing efficiency (3.5× increase) |
Therapeutic Applications and Future Directions
CRISPR-Nanoparticle Hybrid Systems
The bidirectional relationship between CRISPR and QS presents novel therapeutic opportunities. Integrating CRISPR-Cas with nanoparticle delivery systems enhances antibacterial efficacy through multiple mechanisms:
- Enhanced Delivery: Liposomal CRISPR-Cas9 formulations reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro [24].
- Improved Efficiency: Gold nanoparticle carriers enhance CRISPR editing efficiency up to 3.5-fold compared to non-carrier systems [24].
- Synergistic Effects: These platforms enable co-delivery with antibiotics or antimicrobial peptides, producing superior biofilm disruption [24].
Anti-Virulence Strategies
Rather than traditional bactericidal approaches, targeting the CRISPR-QS axis enables anti-virulence strategies that:
- Disrupt biofilm formation without applying direct selective pressure
- Resensitize resistant bacteria to conventional antibiotics
- Modulate host inflammatory responses to infection
Diagnostic Integration
CRISPR-based diagnostics (e.g., SHERLOCK using Cas13, DETECTR using Cas12) enable rapid, on-site detection of biofilm-forming pathogens with attomolar sensitivity and single-base specificity [25]. These can be integrated with therapeutic approaches for theranostic applications.
The bidirectional crosstalk between CRISPR-Cas systems and environmental sensing mechanisms represents a paradigm shift in our understanding of bacterial defense and social organization. CRISPR-Cas systems function not merely as immune mechanisms but as integrated regulatory nodes that process environmental information to optimize population-level responses. The experimental evidence from multiple bacterial pathogens establishes that Cas proteins directly regulate quorum sensing pathways and biofilm formation, while environmental cues reciprocally influence CRISPR efficiency.
For researchers and drug development professionals, this understanding opens promising avenues for novel antimicrobial strategies that target this regulatory interface. By disrupting the circuitry that coordinates virulence and defense, rather than directly killing pathogens, these approaches may slow the development of resistance while effectively mitigating infections. Future research should focus on elucidating the precise molecular mechanisms of this bidirectional communication and translating this knowledge into targeted therapeutic applications with enhanced specificity and reduced collateral impacts on commensal microbiota.
CRISPR-Cas systems provide prokaryotes with adaptive immunity against mobile genetic elements, but this defense comes with substantial fitness costs that necessitate sophisticated regulatory integration with bacterial physiology. Emerging research reveals that bacteria have evolved intricate mechanisms to balance CRISPR-mediated immunity with virulence requirements, primarily through connections with quorum sensing (QS) pathways and biofilm regulation. This whitepaper synthesizes current understanding of how pathogenic bacteria integrate CRISPR-Cas systems within broader regulatory networks to optimize fitness in host environments. We examine the molecular mechanisms governing these trade-offs, present quantitative data on immunity-virulence relationships, and provide detailed methodologies for investigating these connections. The findings have significant implications for developing novel antimicrobial strategies that exploit these natural fitness trade-offs to combat resistant infections.
CRISPR-Cas systems represent a remarkable evolutionary innovation, providing sequence-specific adaptive immunity against bacteriophages and plasmids across bacteria and archaea. However, the energetic costs of maintaining these complex systems—including basal expression, spacer acquisition, and immune execution—create substantial fitness burdens that must be balanced against their protective benefits [1] [26]. This balance becomes particularly critical during host infection, where resources are limited and selective pressures are high. Bacteria resolve this paradox through sophisticated regulatory mechanisms that integrate CRISPR-Cas activity with population-density signaling and virulence factor expression, enabling precise control of immune deployment based on environmental conditions and infection requirements [27] [14].
The integration of CRISPR-Cas systems with quorum sensing and virulence networks allows pathogens to dynamically adjust their investment in immunity relative to other fitness-critical processes. Understanding these regulatory connections provides not only fundamental insights into bacterial pathogenesis but also reveals potential vulnerabilities that could be exploited for therapeutic intervention. This technical guide examines the molecular basis of these fitness trade-offs, quantitative measurements of their impacts, and experimental approaches for their investigation.
Molecular Mechanisms Integrating CRISPR-Cas with Virulence Regulation
Quorum Sensing Control of CRISPR-Cas Systems
Quorum sensing (QS) enables bacterial populations to coordinate gene expression based on cell density, providing an ideal regulatory framework for CRISPR-Cas systems that must balance community-level protection against individual fitness costs. In Serratia sp. ATCC39006, QS directly regulates the expression of type I-E, I-F, and III-A CRISPR-Cas systems [1]. The LuxIR-type QS system in Serratia, comprising smaI (signal synthase) and smaR (transcriptional regulator), controls cas operon and CRISPR expression through an N-butanoyl-L-homoserine lactone (C4-HSL)-mediated de-repression mechanism:
- Low cell density: SmaR binds to promoter regions, repressing cas gene and CRISPR expression
- High cell density: C4-HSL accumulates and binds SmaR, inhibiting its DNA-binding activity and derepressing CRISPR-Cas transcription [1]
This regulatory connection has functional consequences for bacterial immunity. Strains lacking QS signaling capability showed ~20 to ~500-fold reductions in CRISPR interference efficiency against targeted plasmids, demonstrating that density-dependent signaling directly modulates defensive capacity [1]. Similarly, in Pseudomonas aeruginosa, QS controls both CRISPR-Cas expression and the Type IV pili that serve as phage receptors, creating a coordinated response that influences resistance evolution [28].
Table 1: Quorum Sensing Regulation of CRISPR-Cas Systems Across Bacterial Species
| Bacterial Species | QS System | CRISPR-Cas Type | Regulatory Mechanism | Functional Impact |
|---|---|---|---|---|
| Serratia sp. ATCC39006 | LuxIR-type (SmaI/SmaR) | I-E, I-F, III-A | SmaR repression in absence of C4-HSL | 20-500x reduced interference; impaired adaptation |
| Pseudomonas aeruginosa | LasIR/RhlIR | I-F | AHL-mediated regulation | Alters phage resistance evolution |
| Streptococcus agalactiae | CovRS | II-A | CovR repression of P2cas promoter | Enhances immunity against mutated targets |
Virulence Regulator Control of CRISPR-Cas Expression
Beyond canonical QS systems, master virulence regulators directly control CRISPR-Cas expression in important pathogens. In Streptococcus agalactiae (Group B Streptococcus), the CovR (control of virulence) two-component system regulator binds to and represses a distal promoter (P2cas) of the cas operon, integrating CRISPR-Cas9 immunity within the virulence regulatory network [27].
CovR-mediated repression provides several fitness advantages:
- Prevention of autoimmunity: Tight control minimizes off-target Cas9 cleavage and toxicity
- Immune memory enhancement: CovR inactivation increases potency against old spacers and stimulates new spacer acquisition
- Adaptation to phage escape mutants: Elevated Cas9 expression improves targeting of mutated protospacers that would otherwise evade immunity [27]
This regulatory connection demonstrates how pathogens coordinate immune function with virulence expression, potentially allowing enhanced defense during specific stages of infection when virulence genes are derepressed.
CRISPR-Cas Regulation of Biofilm Formation and Virulence
The relationship between CRISPR-Cas systems and virulence is bidirectional, with CRISPR components directly influencing biofilm formation and pathogenicity in some species. In Acinetobacter baumannii, Cas3 of the type I-Fa system upregulates biofilm formation and virulence by enhancing production of extracellular polymeric substances (EPS) and key virulence factors like outer membrane protein A (OmpA) [14].
Mechanistic studies show that cas3 deletion:
- Reduces biofilm thickness and structural complexity
- Decreases adhesion to and invasion of A549 human alveolar epithelial cells by >50%
- Attenuates virulence in Galleria mellonella and murine infection models
- Alters carbon metabolism and oxidative phosphorylation pathways [14]
These findings demonstrate that in some pathogens, CRISPR-Cas components have been co-opted for virulence functions beyond their canonical immune roles, creating an additional fitness trade-off between defense and pathogenic capability.
Quantitative Analysis of Fitness Trade-Offs
The regulation of CRISPR-Cas systems by virulence and QS pathways reflects fundamental fitness trade-offs between immunity costs and pathogenic requirements. Quantitative measurements reveal the magnitude of these trade-offs across bacterial species and infection contexts.
Table 2: Quantitative Fitness Trade-Offs in CRISPR-Cas Immune Systems
| Fitness Parameter | Measurement | Experimental System | Biological Significance |
|---|---|---|---|
| Interference efficiency | 20-500x reduction in QS-deficient mutants | Serratia sp. ATCC39006 | QS enhances defense at high cell density |
| Biofilm formation | 60-70% reduction in Δcas3 mutants | A. baumannii ATCC19606 | Cas3 enhances biofilm matrix production |
| Host cell invasion | 50% reduction in Δcas3 mutants | A. baumannii in A549 cells | CRISPR components facilitate pathogenesis |
| In vivo survival | 50% survival vs. 0% in Δcas3-infected larvae | G. mellonella infection model | Cas3 significantly enhances virulence |
| Spacer acquisition | Impaired in QS-deficient backgrounds | Serratia adaptation assays | QS regulates immune memory formation |
The metabolic costs of CRISPR-Cas immunity manifest through multiple mechanisms:
- Energetic investment: Basal expression and maintenance of Cas proteins and CRISPR arrays
- Infection-induced costs: Expression of phage genes prior to clearance in CRISPR immunity
- Opportunity costs: Resource allocation away from growth and virulence functions
These costs create selective environments where surface-based resistance mutations (which typically carry fixed fitness costs) may be favored over CRISPR immunity (which has infection-induced costs) under high phage exposure [28].
Experimental Approaches and Methodologies
Assessing CRISPR-Cas Regulation by Quorum Sensing
Protocol: Measuring QS-Dependent CRISPR Interference Efficiency
Strain Construction
- Create isogenic QS mutant (e.g., ΔsmaI) and complementation strains
- For Serratia: Use allelic exchange to delete smaI (signal synthase)
- Complement with plasmid-borne smaI or synthetic C4-HSL (100-200 nM)
Interference Assay
- Introduce target plasmids containing protospacers with appropriate PAM sequences
- Use conjugation or transformation to deliver targeted and control plasmids
- Culture strains to high cell density (OD600 ~0.8-1.0) to ensure QS activation
Quantification
- Calculate transformation efficiency (CFU/μg DNA) for targeted vs. non-targeted plasmids
- Determine immunity index: Log₂(transformation efficiency targeted/control)
- Compare indices between WT, QS mutant, and complemented strains [1]
Key Reagents:
- QS signal molecules: C4-HSL for Serratia (100-200 nM working concentration)
- Antibiotics for selection of transformants
- Conjugation or transformation equipment
Evaluating Virulence Regulator Control of CRISPR-Cas
Protocol: CRISPR-Cas Regulation by CovR in S. agalactiae
Genetic Manipulation
- Create ΔcovR mutant using allelic replacement
- Construct P2cas promoter-reporter fusions (e.g., lacZ)
- Generate Cas9-FLAG tagged strains for protein quantification
Binding Assays
- Perform electrophoretic mobility shift assays (EMSAs) with purified CovR and P2cas DNA
- Use DNase I footprinting to identify precise binding sites
- Validate in vivo binding via ChIP-seq or ChIP-qPCR
Functional Immunity Assays
- Transform with vectors containing protospacers matching CRISPR spacers
- Introduce mismatches at various positions to assess targeting stringency
- Calculate immunity indices as described in Section 4.1 [27]
Investigating CRISPR Components in Virulence
Protocol: Assessing Cas3 Role in Biofilm and Virulence
Mutant Construction
- Create cas3 deletion mutant via homologous recombination
- Complement with plasmid-borne cas3 under native promoter
- Verify constructs by PCR and sequencing
Biofilm Assays
- Quantify biofilm formation via crystal violet staining (OD570 measurement)
- Analyze biofilm architecture using confocal laser scanning microscopy (CLSM)
- Stain with SYTO9 (cells) and Alexa Fluor-dextran (EPS matrix)
Virulence Assessment
- Evaluate adhesion/invasion in A549 epithelial cells (MOI 100, 2h infection)
- Determine larval survival in Galleria mellonella model (10⁶ CFU/larva)
- Quantify bacterial loads in murine organs (spleen, liver, lung) after infection [14]
QS-CRISPR Regulatory Pathway
Research Reagent Solutions
Table 3: Essential Research Reagents for Investigating CRISPR-Virulence Connections
| Reagent/Category | Specific Examples | Application/Function |
|---|---|---|
| QS Modulators | C4-HSL, 3OC12-HSL, Baicalein | Activate or inhibit QS signaling pathways |
| Genetic Tools | dCas9 plasmids, gRNA vectors, allelic exchange systems | CRISPRi, gene knockout, complementation |
| Reporter Systems | β-galactosidase, fluorescent proteins, luciferase | Promoter activity quantification |
| Antibodies | Anti-FLAG, protein-specific antibodies | Protein detection and quantification |
| Biofilm Assays | Crystal violet, SYTO9, Alexa Fluor-dextran | Biofilm mass and architecture analysis |
| Infection Models | A549 cells, Galleria mellonella, murine models | Virulence and pathogenicity assessment |
Therapeutic Implications and Future Directions
The intricate connections between CRISPR-Cas immunity and virulence regulation present novel opportunities for antimicrobial development. Several promising approaches emerge from current research:
QS Inhibition Strategies: Chemical inhibition of QS in P. aeruginosa with Baicalein decreases phage adsorption rates and alters the evolution of CRISPR immunity, potentially enhancing phage therapy efficacy [28]. However, the therapeutic outcome depends critically on whether QS inhibition upregulates or downregulates the targeted phage receptors.
CRISPR-Based Antimicrobials: Engineered CRISPR-Cas systems can selectively target virulence genes or antibiotic resistance determinants in pathogens while sparing commensal bacteria. For example, liposomal Cas9 formulations reduced P. aeruginosa biofilm biomass by >90% in vitro [11] [25].
Virulence Disruption: Targeting the regulatory nodes connecting CRISPR and virulence pathways could attenuate pathogenicity without directly killing bacteria, potentially reducing selection for resistance. The Cas3-mediated virulence enhancement in A. baumannii suggests that CRISPR components themselves could be therapeutic targets [14].
Future research should focus on mapping the complete regulatory networks connecting CRISPR-Cas systems with virulence and QS pathways across diverse pathogens, developing delivery systems for CRISPR-based antimicrobials that can penetrate biofilms, and exploring combination therapies that exploit the fitness trade-offs between immunity and virulence.
CRISPR-Cas systems are not autonomous defense modules but are intricately connected with the broader regulatory networks controlling bacterial virulence and social behavior. The balance between immunity costs and virulence requirements represents a fundamental fitness trade-off that pathogens must navigate to successfully colonize host environments. Understanding these connections at molecular, physiological, and evolutionary levels provides both fundamental insights into bacterial pathogenesis and practical avenues for developing novel antimicrobial strategies that exploit these natural trade-offs. The experimental frameworks and reagents outlined in this whitepaper provide researchers with the tools necessary to investigate these connections across diverse bacterial species and infection contexts.
Precision Engineering: CRISPR Tools for Disrupting Quorum Sensing and Biofilm Integrity
CRISPR-Cas9 and CRISPRi for Targeted Gene Knockout and Silencing
Bacterial biofilms represent a significant challenge in both clinical and industrial settings, contributing to persistent infections, antimicrobial resistance, and biofouling. These structured microbial communities are embedded in a protective extracellular polymeric substance (EPS) that confers resistance to conventional antibiotics and environmental stresses [11] [29]. The formation and maintenance of biofilms are tightly regulated by complex genetic networks and quorum sensing (QS) pathways—a cell-cell communication system that enables bacteria to coordinate gene expression in a population-density-dependent manner [29].
CRISPR-based technologies have emerged as transformative tools for dissecting and disrupting these pathways with unprecedented precision. While the CRISPR-Cas9 system enables permanent gene knockout (KO) through targeted DNA cleavage, CRISPR interference (CRISPRi) provides reversible gene silencing without altering the DNA sequence [30] [31]. This technical guide explores the mechanisms, applications, and experimental protocols for both platforms, with specific emphasis on their utility in bacterial quorum sensing and biofilm formation research for drug development and microbial ecology studies.
Core Molecular Mechanisms
The fundamental distinction between CRISPR-Cas9 and CRISPRi lies in the enzymatic activity of the Cas protein and the resulting genetic outcome:
CRISPR-Cas9 for Gene Knockout: Utilizes an active Cas9 nuclease that creates double-strand breaks (DSBs) in target DNA. Cellular repair through error-prone non-homologous end joining (NHEJ) often introduces insertion/deletion mutations (indels) that disrupt the gene's reading frame, resulting in permanent knockout [31] [32]. This approach is ideal for complete ablation of gene function.
CRISPRi for Gene Silencing: Employs a catalytically dead Cas9 (dCas9) with point mutations (D10A and H840A for SpCas9) that inactivate nuclease domains while preserving DNA-binding capability [30] [31]. When fused to repressor domains like the Krüppel-associated box (KRAB), dCas9 blocks transcription initiation or elongation through steric hindrance, achieving reversible gene knockdown without DNA damage [31] [33].
Table 1: Comparative Analysis of CRISPR-Cas9 and CRISPRi Platforms
| Feature | CRISPR-Cas9 Knockout | CRISPR Interference (CRISPRi) |
|---|---|---|
| Cas Protein | Active Cas9 nuclease | Catalytically dead Cas9 (dCas9) |
| Mechanism | DNA cleavage → DSB repair → indels | Steric hindrance of transcription + epigenetic repression |
| Genetic Outcome | Permanent gene knockout | Reversible gene silencing |
| Efficiency | Varies; influenced by repair mechanisms | Typically 60-80% repression with dCas9 alone; >90% with fused repressors [30] [33] |
| Applications in Biofilm Research | Essential gene validation, resistance gene disruption | Titratable studies of essential genes, quorum sensing pathway modulation |
| Advantages | Complete, permanent loss of function | Reversible, titratable, no genotoxic stress [34] [32] |
| Limitations | Potential for off-target editing, cytotoxic DNA damage response | Requires sustained effector expression, position-dependent efficiency |
Applications in Quorum Sensing and Biofilm Research
CRISPR-Cas9 and CRISPRi enable precise manipulation of the genetic circuits governing biofilm development and virulence:
Quorum Sensing Disruption: CRISPRi can target and silence central components of QS systems, including autoinducer synthases (e.g., LuxI homologs) and transcriptional regulators (e.g., LuxR homologs), effectively blocking cell-cell communication without eliminating bacterial populations [29] [25].
Biofilm Matrix Degradation: CRISPR-Cas9 can knockout genes encoding structural components of the EPS matrix, such as exopolysaccharide biosynthesis proteins and extracellular DNA (eDNA) release mechanisms, compromising biofilm integrity [11] [25].
Antibiotic Resensitization: Both platforms can target antibiotic resistance genes (e.g., beta-lactamases, efflux pumps) within biofilms. Recent advances combine CRISPR systems with nanoparticle delivery to enhance penetration through the protective EPS barrier, with liposomal Cas9 formulations demonstrating >90% reduction in Pseudomonas aeruginosa biofilm biomass in vitro [11].
Diagram 1: CRISPR Technology Applications in Biofilm Research
Experimental Design and Workflow
Guide RNA Design Considerations
The design of single guide RNAs (sgRNAs) differs significantly between CRISPR-Cas9 and CRISPRi applications, primarily due to their distinct mechanisms of action:
CRISPR-Cas9 sgRNA Design: For effective gene knockout, sgRNAs should target early exons of protein-coding genes to maximize the probability of frameshift mutations that disrupt the entire coding sequence. Optimal targeting avoids regions with homologous sequences to minimize off-target effects [32].
CRISPRi sgRNA Design: CRISPRi efficiency is highly dependent on the sgRNA binding position relative to the transcriptional start site (TSS). The optimal targeting window spans from -50 to +300 base pairs from the TSS, with the most effective repression occurring within the first 100 bp downstream of the TSS [31] [32]. For CRISPR activation (CRISPRa), which is beyond this guide's scope but shares design principles, the optimal window is -400 to -50 bp upstream of the TSS [30].
Table 2: Quantitative Performance of CRISPR Technologies Against Biofilms
| Target/Application | CRISPR Platform | Delivery System | Efficiency | Reference Model |
|---|---|---|---|---|
| P. aeruginosa biofilm | Cas9 | Liposomal nanoparticles | >90% biomass reduction | In vitro [11] |
| E. coli adhesion genes | CRISPR-Cas9 with HDR | Plasmid | Significant reduction in biofilm formation on urinary catheters | Clinical application model [25] |
| Antibiotic resistance genes | dCas9-KRAB (CRISPRi) | Gold nanoparticles | 3.5× higher editing efficiency vs. non-carrier systems | In vitro [11] |
| Pro-inflammatory cytokines | dCas9-KRAB (CRISPRi) | Lentiviral transduction | >70% mRNA reduction sustained for 72 hours | Human PBMCs [33] |
| Foodborne pathogen biofilms | CRISPR-Cas9 | Bacteriophage | ~3-log reduction of target pathogens | Food processing surfaces [25] |
Delivery Strategies for Biofilm Environments
Efficient delivery of CRISPR components remains a significant challenge in biofilm research due to the protective EPS matrix. Recent advances have focused on engineered delivery systems:
Nanoparticle-Mediated Delivery: Inorganic and organic nanoparticles can enhance CRISPR component stability and biofilm penetration. Gold nanoparticles complexed with CRISPR-Cas9 systems have demonstrated 3.5-fold higher editing efficiency compared to non-carrier systems, while lipid-based nanoparticles protect nucleic acids from degradation in the biofilm microenvironment [11].
Bacteriophage Delivery: Engineered bacteriophages can package CRISPR systems and selectively infect target bacteria within multispecies biofilms. This approach offers species-specific delivery while naturally penetrating biofilm structures [25].
Conjugative Plasmids: Self-transmissible plasmids can facilitate interbacterial transfer of CRISPR constructs, enabling the spread of antimicrobial agents throughout biofilm communities. This approach has been used to eliminate plasmid-borne antibiotic resistance genes, such as mcr-1, from Escherichia coli populations [25].
Research Reagent Solutions
Table 3: Essential Research Reagents for CRISPR Biofilm Studies
| Reagent/Category | Function | Examples & Specifications |
|---|---|---|
| dCas9 Repressor Fusion Proteins | Transcriptional repression in CRISPRi | dCas9-KRAB (most common), dCas9-ZIM3 (enhanced repression) [34] |
| CRISPRa Activator Systems | Transcriptional activation | dCas9-VP64, SAM system (Synergistic Activation Mediator) [30] [31] |
| Nanoparticle Delivery Systems | Enhanced biofilm penetration & delivery | Gold nanoparticles, lipid nanoparticles, polymer-based nanoparticles [11] |
| Optimized sgRNA Libraries | Targeted genetic screens | Genome-wide libraries (3-10 sgRNAs/gene), focused biofilm-related gene sets [32] |
| Bacterial CRISPR Delivery Vectors | Expression of CRISPR components in bacteria | Conjugative plasmids, phage-integrated systems, inducible promoters [25] |
Advanced Methodologies and Protocols
CRISPRi-Mediated Silencing of Quorum Sensing Genes
The following protocol adapts established CRISPRi methodologies for targeting quorum sensing systems in biofilm-forming bacteria:
sgRNA Design and Cloning:
- Identify transcriptional start sites for target QS genes (e.g., lasI, rhlI in P. aeruginosa) using bacterial genome databases.
- Design sgRNAs complementary to the -50 to +300 bp window relative to the TSS.
- Clone sgRNA sequences into a bacterial expression vector containing a dCas9-KRAB expression cassette.
Delivery and Transformation:
- Introduce the CRISPRi construct into target bacteria via electroporation or conjugation.
- For refractory strains, consider nanoparticle-assisted transformation or phage transduction.
Validation of Silencing Efficiency:
- Quantify mRNA reduction of target genes using RT-qPCR 24-48 hours post-transformation.
- Assess functional consequences by measuring autoinducer production (e.g., C4-HSL, 3-oxo-C12-HSL for P. aeruginosa) using HPLC-MS or bioreporter assays.
- Evaluate biofilm formation phenotypes using crystal violet staining or confocal microscopy.
Durability Assessment:
- Monitor silencing persistence over multiple bacterial generations through continuous culture and periodic sampling.
- Assess reversibility by measuring gene expression recovery after removal of selection pressure.
Diagram 2: CRISPRi Experimental Workflow for Quorum Sensing Genes
Combinatorial Approaches for Enhanced Efficacy
Recent advances have demonstrated the superior efficacy of combinatorial CRISPR systems that integrate multiple mechanisms of action:
CRISPRgenee System: This novel approach simultaneously combines Cas9 nuclease activity and dCas9-KRAB repression against the same target gene, achieving more complete loss-of-function than either system alone. The system uses truncated sgRNAs (15-nt) to maintain repression while allowing DNA cleavage, resulting in improved depletion efficiency and reduced sgRNA performance variance [34].
Dual-Targeting Strategies: Implementing multiple sgRNAs against different components of the same regulatory pathway (e.g., targeting both autoinducer synthases and response regulators in QS systems) can produce synergistic effects and reduce the likelihood of resistance development.
Future Directions and Translational Potential
The integration of CRISPR technologies with emerging bioengineering approaches presents exciting opportunities for advanced biofilm control:
AI-Guided sgRNA Design: Machine learning algorithms are being employed to predict optimal sgRNA sequences and target sites based on epigenetic accessibility, sequence composition, and predicted secondary structure, potentially increasing success rates in diverse bacterial species [25].
Integrated Diagnostic-Therapeutic Systems: CRISPR-based biosensors (e.g., Cas12a, Cas13a) can detect specific biofilm-forming pathogens or resistance genes, while simultaneously activating therapeutic CRISPR systems for targeted elimination, creating "smart" antimicrobial systems [35] [25].
Precision Microbiome Engineering: CRISPR systems programmed to selectively eliminate pathogenic strains while preserving commensal microbiota offer promising approaches for managing biofilm-related dysbiosis without broad-spectrum antibiotics [20].
As these technologies mature, they hold significant potential for developing next-generation anti-biofilm strategies that overcome the limitations of conventional antibiotics, particularly for device-associated infections and chronic bacterial persistence where biofilms play a central pathogenic role.
The escalating global health crisis of antibiotic-resistant bacterial infections is profoundly driven by the formation of biofilms—structured communities of microorganisms embedded in a protective extracellular matrix [24]. Biofilms provide an ecological advantage against environmental stressors, making this the most common life-cycle stage for many bacteria and a significant source of antimicrobial-resistant strains in clinical settings [36]. Within the context of a broader thesis on the role of CRISPR in bacterial quorum sensing and biofilm formation research, this technical guide addresses the critical foundation of effective CRISPR intervention: the strategic design of guide RNAs (gRNAs) for precision targeting of key regulatory genes.
The inherent resistance of biofilm-associated infections to conventional antimicrobial therapies necessitates novel therapeutic strategies [24]. The CRISPR-Cas system has emerged as a revolutionary tool that can be programmed to disrupt the complex regulatory networks governing biofilm development [25]. Unlike broad-spectrum antibiotics, CRISPR-based approaches can be designed to target specific genes with precision, potentially resensitizing bacteria to traditional antibiotics or preventing biofilm formation altogether [24] [37]. The efficacy of these approaches hinges on the rational selection of target genes and the optimal design of gRNAs, which direct the Cas nuclease to specific genomic sequences [38]. This guide provides researchers and drug development professionals with a comprehensive framework for gRNA design against quorum sensing genes and biofilm regulators, incorporating quantitative data, experimental protocols, and essential bioinformatic tools.
Biofilm Formation and Key Regulatory Circuits
Biofilm development is a complex, multi-stage process tightly regulated by interconnected genetic networks. Understanding these circuits is prerequisite to effective target selection.
Stages of Biofilm Development
The biofilm life cycle follows a defined progression [36] [39]:
- Initial Attachment: Planktonic cells reversibly adhere to a surface.
- Irreversible Attachment: Cells form strong adhesions via adhesins and pili.
- Microcolony Formation: Cells proliferate and begin producing extracellular polymeric substances (EPS).
- Maturation: A complex, three-dimensional architecture develops with characteristic mushroom-like structures and water channels.
- Dispersion: Cells detach from the biofilm to colonize new surfaces.
Core Regulatory Systems
Three key regulatory systems intricately control the transition from planktonic to sessile biofilm lifestyles, and they represent prime targets for CRISPR intervention.
Quorum Sensing (QS) is a cell-density dependent communication system where bacteria produce, release, and detect small signaling molecules called autoinducers [39]. This system regulates a broad range of genes, including those for virulence factor production and biofilm formation [28]. Key QS components include:
- LuxI/LuxR-type systems: In Gram-negative bacteria, LuxI-type synthases produce acyl-homoserine lactone (AHL) autoinducers, which bind to LuxR-type receptor proteins to activate gene transcription [28].
- AIO-2/LuxS system: The luxS gene encodes the synthase for Autoinducer-2 (AIO-2), a universal signaling molecule involved in interspecies communication and biofilm maturation [37].
Cyclic di-GMP (c-di-GMP) is a ubiquitous second messenger that promotes biofilm formation by inversely regulating motility and EPS production [36] [40]. High intracellular c-di-GMP concentrations repress motility and stimulate matrix production, cementing the sessile lifestyle.
Small Non-coding RNAs (sRNAs) are post-transcriptional regulators that fine-tune gene expression in response to environmental cues, including stress [36] [40]. They often act by base-pairing with target mRNAs, affecting their translation and stability, typically with the help of the Hfq RNA chaperone. sRNAs are pivotal in the decision to "stay" (form biofilm) or "go" (remain planktonic).
The diagram below illustrates the interconnected relationships between these core regulatory systems during biofilm development:
Strategic Target Gene Selection
The selection of appropriate target genes is the most critical step in designing an effective CRISPR-based anti-biofilm strategy. Ideal targets are master regulators whose disruption causes significant impairment of biofilm formation or stability. The table below summarizes high-value target genes, their mechanisms, and the phenotypic consequences of their disruption.
Table 1: High-Value Target Genes for Anti-Biofilm gRNA Design
| Target Gene | Regulatory System | Function | Effect of Disruption | Bacterial Species |
|---|---|---|---|---|
| luxS [37] | Quorum Sensing | AI-2 Autoinducer synthase | Inhibits biofilm maturation; reduces EPS production | E. coli, others |
| csgD [40] | sRNA/c-di-GMP | Transcriptional regulator of curli fibrils | Decreases curli and cellulose production | E. coli, Salmonella |
| AbaI [38] | Quorum Sensing | AHL Autoinducer synthase | Reduces biofilm formation and virulence | Acinetobacter baumannii |
| lasI/rhlI [28] | Quorum Sensing | AHL Synthases | Downregulates virulence and CRISPR-cas expression | Pseudomonas aeruginosa |
| rsmA [40] | sRNA | RNA-binding protein regulator | Alters EPS synthesis and biofilm development | P. aeruginosa |
Selection Criteria and Quantitative Considerations
When prioritizing targets, researchers should consider:
- Network Centrality: Genes like csgD and luxS act as hubs, and their disruption has cascading effects. CRISPRi inhibition of luxS in E. coli has been experimentally validated to reduce biofilm formation [37].
- Phenotypic Impact: Targets should be empirically linked to crucial phenotypes. For instance, targeting the AbaI gene in A. baumannii disrupts QS and is predicted to reduce biofilm-associated virulence [38].
- Conservation and Specificity: For narrow-spectrum applications, target unique genes. For broader applications, consider conserved genes like luxS.
- Therapeutic Context: The target must be relevant in the infection model. For example, in P. aeruginosa, QS inhibition not only affects virulence but also influences the evolution of phage resistance by modulating the expression of the Type IV pilus, a common phage receptor [28].
gRNA Design and Optimization Workflow
A rigorous, multi-stage workflow is essential for designing highly efficient and specific gRNAs.
gRNA Design Principles
The gRNA directs the Cas nuclease to its genomic target via a 20-nucleotide spacer sequence. Key design principles include:
- Protospacer Adjacent Motif (PAM): The target site must be adjacent to the PAM sequence, which is 5'-NGG-3' for the most commonly used S. pyogenes Cas9 [37] [41].
- Seed Sequence: The 10-12 nucleotides proximal to the PAM are critical for binding specificity and tolerance to mismatches [41].
- On-Target Efficiency: Algorithms predict efficiency based on GC content (prefer 40-80%), nucleotide composition, and position-specific features.
- Off-Target Minimization: The gRNA sequence should be unique within the genome, with no or minimal near-complementary sites, especially in the seed region [41].
The following workflow outlines the key steps from target identification to experimental validation:
Protocol: gRNA Cloning and Delivery
This protocol details the experimental steps for cloning gRNAs into expression vectors, based on established methods [37] [41].
Materials:
- pdCas9 plasmid (e.g., Addgene #44249) and pgRNA plasmid (e.g., Addgene #44251) [37].
- Chemically competent E. coli cells (e.g., Top10 for cloning, target strain for expression).
- Electra cloning kit or similar restriction-ligation reagents.
- Antibiotics (Ampicillin, Chloramphenicol).
- Inducer (Anhydrotetracycline, aTc).
Method:
- gRNA Insert Design: For your target gene (e.g., luxS), synthesize three sets of primers containing a 20 bp sequence complementary to the target site immediately preceding a 5'-CCN-3' PAM, flanked by 35 nt of the dCas9 handle sequence [37].
- Cloning into pgRNA Vector: Perform inverse PCR on the pgRNA plasmid using the phosphorylated primers. Digest the PCR product with DpnI to remove the template, purify, and ligate using a blunt-end ligation kit to create circular plasmids (e.g., pgRNA-LV1, pgRNA-LV2) [37].
- Transformation and Sequencing: Transform the ligated plasmids into competent E. coli Top10 cells. Screen colonies by PCR and confirm successful cloning by Sanger sequencing.
- Co-transformation: Isolate the confirmed pgRNA plasmids and co-transform them with the pdCas9 plasmid into your target bacterial strain (e.g., E. coli AK-117).
- Induction: To activate the CRISPRi system, grow the knockdown strain in media supplemented with the appropriate antibiotics and 2 μM aTc [37].
Experimental Validation and Phenotypic Assessment
After creating knockdown strains, a multi-tiered validation approach is required to confirm target suppression and its functional consequences.
Table 2: Key Assays for Validating Anti-Biofilm gRNA Efficacy
| Assay Type | Specific Method | Measured Outcome | Interpretation |
|---|---|---|---|
| Molecular Confirmation | qRT-PCR [37] | mRNA expression level of target gene (e.g., luxS) | Confirms transcriptional knockdown. |
| Biomass Quantification | Crystal Violet Staining [37] | Total adhered biofilm biomass | Measures overall reduction in biofilm formation. |
| Metabolic Activity | XTT Reduction Assay [37] | Metabolic activity of biofilm cells | Assesses biofilm viability post-treatment. |
| Structural Analysis | Scanning Electron Microscopy (SEM) [37] | 3D architecture and integrity of biofilm | Visualizes disruption of biofilm microstructure. |
| Functional Disruption | Gene Expression Panels | Downstream virulence and matrix genes | Verifies cascade effect on regulatory network. |
Advanced Delivery and Synergistic Strategies
For translational applications, efficient delivery of CRISPR components is paramount. Nanoparticles (NPs) present an innovative solution, serving as effective carriers for CRISPR/Cas9 components while exhibiting intrinsic antibacterial properties [24].
- Nanoparticle Carriers: Liposomal Cas9 formulations have been shown to reduce P. aeruginosa biofilm biomass by over 90% in vitro, while gold nanoparticle carriers can enhance editing efficiency by up to 3.5-fold compared to non-carrier systems [24].
- Synergistic Therapies: These hybrid platforms enable co-delivery with antibiotics or antimicrobial peptides, producing superior biofilm disruption and antibacterial effects [24].
- CRISPR Diagnostics Integration: For a comprehensive strategy, Cas12a/Cas13-based biosensors (e.g., DETECTR, SHERLOCK) can be used for rapid, on-site detection and serotyping of biofilm-forming pathogens before and after intervention [25].
The Scientist's Toolkit: Research Reagent Solutions
A curated list of essential materials and tools is critical for implementing the protocols described in this guide.
Table 3: Essential Research Reagents and Tools for gRNA-Based Biofilm Research
| Reagent / Tool | Function / Description | Example Source / Specification |
|---|---|---|
| dCas9 Plasmids | Catalytically "dead" Cas9 for CRISPRi; blocks transcription without cutting DNA. | pdCas9 (Addgene #44249) [37] |
| sgRNA Cloning Vector | Plasmid for expression of gene-specific single guide RNA (sgRNA). | pgRNA (Addgene #44251) [37] |
| Electra Cloning Kit | Enzyme mix for rapid, seamless assembly of gRNA inserts into daughter vectors. | ATUM Bio (Catalog #EKT-03) [41] |
| ATUM gRNA Design Tool | Web-based algorithm for designing gRNAs with predicted high specificity and efficiency. | https://www.atum.bio [41] |
| Anhydrotetracycline (aTc) | Small-molecule inducer for high-precision, temporal control of dCas9/sgRNA expression. | Final working concentration: 2 μM [37] |
| Liposomal Nanoparticles | Advanced delivery system for CRISPR components to enhance biofilm penetration and editing efficiency. | Formulations shown to reduce biofilm by >90% [24] |
Strategic gRNA design targeting master regulators of quorum sensing and biofilm formation represents a powerful application of CRISPR technology in the fight against antibiotic-resistant infections. By systematically selecting central nodes in the regulatory network—such as luxS, csgD, and AbaI—and following a rigorous workflow encompassing in silico design, experimental validation, and advanced delivery methods, researchers can develop highly specific and effective anti-biofilm strategies. The integration of these precision genetic tools with nanoparticle delivery and synergistic antimicrobials holds immense potential for translating CRISPR-based biofilm control from the laboratory to clinical and industrial settings, ultimately contributing to a broader solution for the global antimicrobial resistance crisis.
Nanoparticle-CRISPR Hybrid Systems for Enhanced Biofilm Penetration and Delivery
The escalating crisis of antibiotic-resistant infections, largely driven by biofilm-associated bacteria, demands innovative therapeutic strategies. This technical guide explores the development of nanoparticle-CRISPR hybrid systems engineered to overcome the fundamental limitations of conventional antimicrobials against biofilms. By integrating the precision of CRISPR-based gene editing with the enhanced delivery capabilities of advanced nanomaterials, these platforms represent a paradigm shift in targeting the genetic underpinnings of antibiotic resistance, quorum sensing mechanisms, and biofilm integrity. We provide a comprehensive analysis of the mechanisms, experimental data, and methodological protocols underpinning this emerging technology, contextualized within the broader scientific inquiry into CRISPR's role in bacterial quorum sensing and biofilm formation research.
Biofilm-mediated infections present a formidable challenge in clinical settings due to their inherent tolerance to antimicrobials. Biofilms are structured communities of microorganisms encapsulated within a self-produced matrix of extracellular polymeric substances (EPS) that can exhibit up to 1000-fold greater tolerance to antibiotics compared to their planktonic counterparts [11]. This protective matrix limits antibiotic penetration, enhances horizontal gene transfer, and creates heterogeneous microenvironments with differential metabolic activity, facilitating bacterial survival [42] [11].
The integration of CRISPR/Cas9 gene-editing technology with nanoparticle-based delivery systems has emerged as a revolutionary approach to combat biofilm-associated antibiotic resistance. This hybrid strategy enables precise targeting of genetic resistance determinants while simultaneously overcoming physical biofilm barriers, offering a synergistic antimicrobial strategy that addresses both genetic and phenotypic resistance mechanisms [42] [11].
Technical Mechanisms of Action
CRISPR/Cas9 Targeting Strategies for Biofilm Disruption
The CRISPR/Cas9 system provides unparalleled precision in targeting specific genetic elements that govern biofilm formation and maintenance. The system consists of two key components: the Cas9 nuclease, which introduces double-strand breaks in DNA, and a guide RNA (gRNA) that directs Cas9 to specific genomic sequences [11]. In the context of biofilm disruption, three primary targeting strategies have been developed:
- Disruption of Antibiotic Resistance Genes: By designing gRNAs to target and disrupt genes encoding antibiotic resistance mechanisms (e.g., bla, mecA, ndm-1), CRISPR/Cas9 can resensitize biofilm-embedded bacteria to conventional antibiotics [11].
- Interference with Quorum Sensing Pathways: Quorum sensing (QS) is a bacterial cell-to-cell communication system that coordinates biofilm formation and virulence factor expression. CRISPR/Cas9 can target and disrupt key genes in QS pathways (e.g., LasIR and RhlIR in Pseudomonas aeruginosa), effectively silencing bacterial communication and preventing biofilm maturation [43].
- Targeting Biofilm-Regulating Factors: Essential structural and regulatory components of biofilms, including genes responsible for EPS production, adhesion proteins, and transcriptional regulators, can be precisely edited to compromise biofilm integrity [42] [44].
Nanoparticle-Mediated Delivery Enhancement
Nanoparticles (NPs) serve as transformative carriers for CRISPR/Cas9 components, addressing critical delivery challenges while exhibiting intrinsic antibacterial properties. Different classes of nanoparticles offer distinct advantages for biofilm penetration and CRISPR delivery [42] [45]:
Table 1: Nanoparticle Platforms for CRISPR Delivery Against Biofilms
| Nanoparticle Type | Key Advantages | CRISPR Delivery Efficacy | Intrinsic Anti-biofilm Properties |
|---|---|---|---|
| Lipid-based NPs | Biocompatibility, encapsulation efficiency, fusion with bacterial membranes | Liposomal Cas9 formulations reduce P. aeruginosa biofilm biomass by >90% in vitro [42] | Membrane disruption, controlled release in biofilm microenvironment |
| Gold NPs | Surface functionalization, optical properties, photothermal activation | 3.5-fold increase in editing efficiency compared to non-carrier systems [42] | Enhanced penetration through EPS matrix |
| Polymeric NPs | Tunable degradation rates, stimuli-responsive release | High payload capacity for Cas9 ribonucleoprotein complexes [45] | Mucoadhesive properties for prolonged retention |
| Hybrid LNP-SNAs | Dense DNA coating for enhanced cellular uptake, organ targeting | 3x more effective cell entry, 3x higher editing efficiency, >60% improvement in precise DNA repairs [46] | Architecture-dependent uptake across cell types |
The mechanism of nanoparticle-mediated delivery involves multiple enhanced functionalities: improved cellular uptake through surface receptor interactions, protection of genetic material from degradation within the biofilm environment, increased target specificity through surface modifications, and controlled release triggered by biofilm-specific stimuli (e.g., pH, enzymes) [42] [11] [45].
Visualizing the Hybrid System Mechanism
The following diagram illustrates the coordinated mechanism of nanoparticle-CRISPR hybrid systems for enhanced biofilm penetration and delivery:
Diagram 1: Hybrid System Mechanism for Biofilm Disruption
Quantitative Efficacy Data
Recent advances in nanoparticle-CRISPR hybrid systems have demonstrated significant efficacy against resilient biofilms. The quantitative data below highlights the performance of various platforms:
Table 2: Quantitative Efficacy of Nanoparticle-CRISPR Systems Against Biofilms
| Platform | Target Organism | Biofilm Reduction | Editing Efficiency | Synergistic Effects |
|---|---|---|---|---|
| Liposomal CRISPR-Cas9 | Pseudomonas aeruginosa | >90% reduction in biomass in vitro [42] | Not specified | Co-delivery with antibiotics enhances biofilm disruption |
| CRISPR-Gold Nanoparticle Hybrids | Model bacterial systems | Significant disruption | 3.5x increase compared to non-carrier systems [42] | Synergistic action with antibiotics demonstrated |
| LNP-SNAs (Lipid Nanoparticle Spherical Nucleic Acids) | Various human cell types (validation studies) | Enhanced penetration capability | 3x higher than standard LNPs, >60% improvement in precise DNA repairs [46] | Reduced cellular toxicity, modular targeting |
The synergistic potential of these systems is particularly noteworthy. Hybrid platforms enable co-delivery of CRISPR components with antibiotics or antimicrobial peptides, creating multifaceted approaches that attack bacterial populations through both genetic disruption and traditional antimicrobial mechanisms [42] [11]. This combination strategy is essential for addressing the complex nature of biofilm-associated resistance, which involves both genetic determinants and physical protection mechanisms.
Experimental Protocols and Methodologies
Protocol: Liposomal CRISPR-Cas9 Preparation and Biofilm Assay
This protocol details the methodology for developing liposomal CRISPR-Cas9 formulations and evaluating their efficacy against bacterial biofilms, based on established procedures with reported >90% biofilm reduction [42].
Materials Required:
- Cas9 protein and guide RNA (designed against target biofilm genes)
- Lipid mixture (e.g., DOTAP, DOPE, cholesterol)
- Biofilm-forming bacterial strain (e.g., P. aeruginosa)
- Standard culture media and biofilm growth substrates
- Confocal laser scanning microscopy (CLSM) equipment
- Quantitative PCR and viability assay reagents
Procedure:
Liposome Preparation
- Prepare lipid film by evaporating chloroform from lipid mixture under nitrogen stream
- Hydrate lipid film with CRISPR-Cas9 complex (preassembled Cas9 and gRNA) in appropriate buffer
- Subject to freeze-thaw cycles and extrude through polycarbonate membranes (100-400 nm)
- Purify using size exclusion chromatography to remove unencapsulated CRISPR components
- Characterize particle size (Zetasizer), zeta potential, and encapsulation efficiency
Biofilm Cultivation
- Grow bacterial cultures to mid-log phase in suitable media
- Inoculate biofilm substrates (e.g., peg lids, chambered coverslips) with bacterial suspension
- Incubate for 24-48 hours to allow mature biofilm formation
- Verify biofilm development using microscopy techniques
Treatment and Assessment
- Apply liposomal CRISPR-Cas9 formulations to established biofilms
- Include appropriate controls (empty liposomes, free CRISPR, untreated)
- Incubate for predetermined time periods (typically 4-24 hours)
- Assess biofilm biomass using crystal violet staining or CLSM
- Evaluate bacterial viability through colony-forming unit (CFU) counts
- Analyze genetic editing efficiency via sequencing of target loci
Protocol: Gold Nanoparticle-CRISPR Conjugation and Evaluation
This protocol outlines the methodology for creating gold nanoparticle-CRISPR conjugates that demonstrate 3.5-fold enhanced editing efficiency [42].
Materials Required:
- Gold nanoparticles (15-30 nm)
- Thiol-modified DNA linkers for Cas9/gRNA complex attachment
- Cas9 ribonucleoprotein (RNP) complexes with target-specific gRNAs
- Bacterial transformation or electroporation equipment
- Gel electrophoresis supplies for validation
Procedure:
Surface Functionalization
- Incubate gold nanoparticles with thiolated DNA linkers (16-24 hours)
- Purify functionalized nanoparticles from excess linkers using centrifugation
- Characterize functionalization success through UV-Vis spectroscopy and dynamic light scattering
CRISPR Complex Conjugation
- Preassemble Cas9 RNP complexes with target-specific gRNAs
- Incubate functionalized gold nanoparticles with RNP complexes
- Allow conjugation through complementary DNA hybridization
- Purify conjugates and confirm using gel shift assays
Efficacy Evaluation
- Deliver conjugates to bacterial cultures or biofilms using optimized methods
- Include controls (naked CRISPR, unconjugated nanoparticles)
- Assess editing efficiency through restriction fragment length polymorphism or sequencing
- Quantify biofilm penetration using fluorescently labeled components
- Evaluate phenotypic effects on biofilm structure and antibiotic susceptibility
Visualizing Experimental Workflow
The standardized workflow for developing and testing nanoparticle-CRISPR hybrid systems is illustrated below:
Diagram 2: Experimental Workflow for Hybrid System Evaluation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Nanoparticle-CRISPR Biofilm Studies
| Reagent Category | Specific Examples | Function/Purpose | Key Considerations |
|---|---|---|---|
| CRISPR Components | Cas9 protein, sgRNAs targeting quorum sensing genes (lasR, rhlR), repair templates | Precision editing of biofilm-related genes | gRNA design critical for specificity; off-target effects must be assessed |
| Nanoparticle Platforms | Lipid nanoparticles (DOTAP/DOPE), gold nanoparticles, polymeric NPs (PLGA) | CRISPR delivery and biofilm penetration | Size, surface charge, and functionalization affect penetration efficiency |
| Biofilm Assay Tools | Crystal violet, confocal microscopy with BacLight viability stains, Calgary biofilm device | Biofilm quantification and visualization | Multiple assessment methods recommended for comprehensive analysis |
| Bacterial Strains | P. aeruginosa PA14, CRISPRI-mutant libraries, clinical biofilm-forming isolates | Relevant biofilm models | Include both laboratory strains and clinical isolates for translational relevance |
| Quorum Sensing Inhibitors | Baicalein, furanones, AHL analogs | Comparative controls for QS disruption | Mechanism of action differs from genetic targeting by CRISPR |
| Characterization Equipment | Dynamic light scattering, transmission electron microscopy, HPLC | Nanoparticle and conjugate validation | Thorough characterization essential for reproducible results |
Integration with Quorum Sensing Research
The application of nanoparticle-CRISPR hybrid systems intersects significantly with quorum sensing (QS) research, as QS mechanisms fundamentally regulate biofilm formation and virulence in many pathogenic bacteria. The CRISPR-Cas system itself is regulated by QS in some bacterial species, creating a complex interplay that can be exploited therapeutically [28] [43].
Research has demonstrated that QS controls the expression of the CRISPR-cas immune system in Pseudomonas aeruginosa [28]. This relationship suggests that CRISPR-based approaches can be designed to target the very regulatory systems that may control their natural bacterial counterparts. Furthermore, inhibiting QS with chemical inhibitors like Baicalein has been shown to influence phage adsorption rates and affect the evolution of CRISPR immunity in bacterial populations, highlighting the complex connections between these systems [28].
The strategic integration of nanoparticle-CRISPR technologies with QS research enables:
- Precision targeting of QS master regulators (e.g., LasI/LasR in P. aeruginosa)
- Disruption of autoinducer synthesis and reception pathways
- Intervention in the coordination of biofilm development and dispersal
- Potential to resensitize persistent cells to conventional antibiotics
This approach represents a significant advancement over traditional QS inhibition strategies by permanently disrupting QS circuitry at the genetic level rather than temporarily interfering with signaling molecules.
Challenges and Future Perspectives
Despite the promising results, several challenges remain in translating nanoparticle-CRISPR hybrid systems into clinical applications. Key limitations include optimizing delivery platforms for specific biofilm types, minimizing potential off-target effects, ensuring long-term safety, and navigating regulatory pathways for these complex therapeutic entities [42]. The specificity of CRISPR editing requires careful gRNA design to avoid unintended genetic consequences, and nanoparticle delivery systems must be refined for optimal penetration into different biofilm architectures.
Future research directions should focus on:
- Developing stimuli-responsive nanoparticles that release CRISPR payloads in response to biofilm-specific signals
- Engineering targeted delivery systems with specificity for particular bacterial species within polymicrobial biofilms
- Exploring CRISPR-based approaches that target multiple genetic elements simultaneously
- Conducting comprehensive in vivo efficacy and toxicity studies
- Investigating the potential of CRISPR-nanoparticle systems to prevent biofilm formation on medical devices
The convergence of nanotechnology and gene editing represents a frontier in antimicrobial strategy, particularly against recalcitrant biofilm-associated infections. As these platforms evolve, they hold immense potential to address the growing crisis of antibiotic resistance by providing precision tools to disrupt the genetic foundations of bacterial persistence and virulence.
The escalating crisis of antimicrobial resistance (AMR), particularly within biofilm-associated infections, necessitates the development of innovative therapeutic strategies. The CRISPR-Cas system, a revolutionary gene-editing tool, presents a highly specific weapon to disarm bacterial defense mechanisms. This whitepaper explores the paradigm of co-delivering CRISPR constructs with conventional antibiotics, a synergistic combination that resensitizes resistant bacteria to traditional treatments. We detail the molecular mechanisms, focusing on the disruption of quorum sensing and biofilm integrity, and provide a comprehensive technical guide on nanoparticle-mediated delivery systems. Supported by quantitative data, standardized protocols, and visual workflows, this document serves as a foundational resource for researchers and drug development professionals aiming to design next-generation antimicrobials that overcome the formidable challenge of biofilm-driven resistance.
Biofilms are structured communities of microorganisms encapsulated within a self-produced extracellular polymeric substance (EPS) matrix, acting as a primary driver of antimicrobial resistance in chronic and device-associated infections [11] [47]. This EPS matrix creates a formidable physical and physiological barrier, reducing antibiotic penetration by up to 1,000-fold compared to planktonic cells and fostering heterogeneous bacterial populations with recalcitrant "persister" cells [11] [44]. Conventional antibiotics, which predominantly target actively growing cells, are often ineffective against these dormant, biofilm-embedded populations.
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) system, derived from a prokaryotic adaptive immune system, has emerged as a precision genetic tool to combat this challenge [48] [49]. Its application extends beyond traditional gene editing to include CRISPR interference (CRISPRi) for targeted gene repression without altering the DNA sequence [25]. By programming CRISPR-Cas systems to disrupt key genetic determinants of pathogenicity—such as antibiotic resistance genes (e.g., blaNDM, mecA, mcr-1), quorum sensing (QS) circuits, and biofilm-regulating factors—researchers can selectively resensitize bacterial populations, making them vulnerable again to conventional antibiotics [11] [50] [51]. This whitepaper frames this approach within a broader thesis on manipulating bacterial quorum sensing and biofilm formation, detailing the technical execution of co-delivering CRISPR constructs and antibiotics using advanced nanocarriers.
Mechanisms of Action: Precision Strikes Against Bacterial Defenses
The synergistic effect of CRISPR and antibiotics is achieved through a multi-pronged attack on the bacterial resistance apparatus. The table below summarizes key target genes and their functional consequences when disrupted by CRISPR.
Table 1: Key Genetic Targets for CRISPR-Based Sensitization of Biofilm-Forming Bacteria
| Target Category | Example Genes | Function of Gene Product | Effect of CRISPR Disruption |
|---|---|---|---|
| Antibiotic Resistance | mecA, blaNDM-1, mcr-1 |
Confers resistance to beta-lactams, carbapenems, and colistin. | Restores susceptibility to the corresponding antibiotic [50] [51]. |
| Quorum Sensing | lasI/R, rhlI/R |
Synthases and receptors for acyl-homoserine lactone (AHL) signaling molecules in P. aeruginosa. | Attenuates virulence factor production and biofilm maturation [11] [44]. |
| Biofilm Matrix | pelA, pslA, algD |
Involved in the synthesis of polysaccharide components of the EPS. | Compromises biofilm structural integrity, enhancing antibiotic penetration [25] [47]. |
| Global Regulators | rpoS, csgD |
Stress response and curli fiber production regulators in E. coli. | Reduces bacterial fitness and ability to form robust biofilms [25]. |
The logical flow of this synergistic mechanism, from cellular delivery to bactericidal effect, is illustrated in the following pathway:
Diagram 1: Synergistic antibacterial mechanism. The pathway illustrates how nanoparticle-mediated co-delivery of CRISPR constructs and antibiotics leads to synergistic biofilm eradication.
Delivery Platforms: Overcoming the Biofilm Barrier with Nanocarriers
The clinical translation of CRISPR-based antimicrobials is critically dependent on efficient delivery systems. Nanoparticles (NPs) offer an innovative solution, protecting CRISPR components from degradation and enhancing their penetration through the dense EPS matrix [11] [49]. Different nanocarriers have been engineered to co-encapsulate CRISPR payloads and antibiotics, as summarized below.
Table 2: Nanoparticle Platforms for Co-delivery of CRISPR and Antibiotics
| Nanoparticle Type | Key Characteristics | CRISPR Payload Format | Antibiotic Example | Documented Efficacy |
|---|---|---|---|---|
| Liposomal NPs | Biocompatible, fuse with bacterial membranes. | Plasmid DNA or ribonucleoprotein (RNP) complex. | Ciprofloxacin, Tobramycin. | >90% reduction in P. aeruginosa biofilm biomass in vitro [11] [42]. |
| Gold Nanoparticles (AuNPs) | Conjugatable surface, tunable size, photothermal properties. | Covalently conjugated RNP complex. | Ampicillin, Vancomycin. | 3.5-fold increase in gene-editing efficiency vs. non-carrier systems [11]. |
| Polymeric NPs | High stability, controlled release profile. | Plasmid DNA encapsulated via electrostatic interaction. | Levofloxacin. | Successful in vivo resensitization of MRSA to methicillin [51]. |
| Bacteriophage-NP Hybrids | Natural bacterial targeting, engineered for enhanced penetration. | Cas9/sgRNA encoded in phage genome. | Not specified. | Effective targeting and killing of specific pathogens in mixed communities [25] [51]. |
Quantitative Evidence of Synergistic Efficacy
Robust experimental data validates the superior efficacy of the combinatorial approach over mono-therapies. The following table consolidates key quantitative findings from recent studies.
Table 3: Quantitative Efficacy of CRISPR-Antibiotic Co-Delivery Systems
| Pathogen & Model | CRISPR Target | Co-delivered Antibiotic | Nanocarrier | Key Synergistic Outcome |
|---|---|---|---|---|
| Pseudomonas aeruginosa (in vitro) | Quorum sensing genes (lasI/R) |
Tobramycin | Liposomal NPs | >90% reduction in biofilm biomass; >3-log reduction in viable cell count [11]. |
| Escherichia coli (in vitro) | Colistin resistance gene (mcr-1) |
Colistin | Gold Nanoparticles (AuNPs) | Restored susceptibility, leading to complete bacterial eradication at sub-MIC colistin doses [50] [51]. |
| Staphylococcus aureus (in vivo) | Methicillin resistance gene (mecA) |
Methicillin | Polymer-derivatized Cas9 RNP complex | Significant reduction in bacterial load in murine skin abscess model; resensitization to antibiotic observed [51]. |
| Mixed Biofilm Community (in vitro) | Species-specific virulence genes | Broad-spectrum antibiotic | Phage-guided NPs | Precatile elimination of target pathogen without disrupting the broader microbial community [25]. |
Experimental Protocols: A Guide to Key Methodologies
For researchers seeking to replicate and build upon these findings, this section provides detailed protocols for core experimental procedures.
Protocol: Formulation of Liposomal CRISPR-Antibiotic Nanoparticles
This protocol describes the preparation of co-encapsulating liposomes using a thin-film hydration and extrusion method [11].
- Lipid Film Formation: Dissolve a lipid mixture (e.g., DOPC:Cholesterol:DOTAP at a molar ratio of 55:40:5) in chloroform in a round-bottom flask. Remove the organic solvent under reduced pressure using a rotary evaporator to form a thin, uniform lipid film on the flask wall.
- Hydration with Active Agents: Hydrate the dried lipid film with an ammonium sulfate solution (250 mM, pH 5.5) containing the CRISPR payload (e.g., 100 µg of pre-assembled Cas9-sgRNA RNP complex) and the antibiotic (e.g., 50 µg of tobramycin). Vortex vigorously for 1 hour above the lipid transition temperature (e.g., 50°C) to form large, multilamellar vesicles (LMVs).
- Size Reduction and Purification: Extrude the hydrated liposome suspension through a series of polycarbonate membranes (e.g., 400 nm, 200 nm, and finally 100 nm) using a mini-extruder to obtain small, unilamellar vesicles (SUVs) with a uniform size distribution.
- Remote Loading (Optional): For ionizable antibiotics, dialyze the extruded liposomes against a HEPES-buffered saline (HBS, pH 7.4) to create a transmembrane pH gradient, which can enhance antibiotic loading.
- Purification and Storage: Purify the final formulation from non-encapsulated materials using size-exclusion chromatography (e.g., Sephadex G-50) or dialysis. Filter-sterilize the liposome suspension (0.22 µm pore size) and store at 4°C for short-term use.
Protocol: In Vitro Assessment of Anti-Biofilm Efficacy
This standard assay quantifies the ability of the formulation to disrupt and kill bacteria within a pre-established biofilm [11] [47].
- Biofilm Formation: Grow the target bacterial strain (e.g., P. aeruginosa PAO1) in a suitable medium (e.g., Tryptic Soy Broth) in a 96-well flat-bottom polystyrene plate for 24-48 hours at 37°C to allow for robust biofilm formation on the well walls.
- Treatment: Carefully aspirate the planktonic culture and rinse the biofilm gently with phosphate-buffered saline (PBS). Add the experimental formulations: (a) PBS control, (b) antibiotic alone, (c) CRISPR-NPs alone, and (d) CRISPR-Antibiotic NP combination. Incubate for a specified period (e.g., 24 hours) at 37°C.
- Biofilm Biomass Quantification (Crystal Violet Assay):
- Aspirate the treatment, rinse the biofilm with PBS, and air-dry.
- Fix the biofilm with 99% methanol for 15 minutes, then discard and air-dry.
- Stain with 0.1% crystal violet solution for 20 minutes.
- Rinse extensively with water to remove unbound dye.
- Elute the bound dye with 33% acetic acid.
- Measure the absorbance of the eluent at 595 nm. Lower absorbance indicates reduced biofilm biomass.
- Viable Cell Count (Colony Forming Units - CFU):
- After treatment in a separate plate, aspirate the treatment and rinse with PBS.
- Scrape the biofilm from the wells into PBS using a pipette tip and vigorous pipetting.
- Vortex the suspension vigorously to disaggregate cells.
- Serially dilute the suspension in PBS and plate on solid agar medium.
- Incubate plates overnight at 37°C and count the resulting colonies. Report as Log₁₀ CFU/mL.
The workflow for designing and executing such an experiment is systematic and can be visualized as follows:
Diagram 2: Experimental workflow for anti-biofilm testing. The flowchart outlines the key steps from gRNA design to data analysis for evaluating synergistic combinations.
The Scientist's Toolkit: Essential Research Reagents
The successful implementation of the protocols and strategies outlined above relies on a suite of key reagents and tools.
Table 4: Essential Research Reagents for CRISPR-Antibiotic Synergy Studies
| Reagent / Tool Category | Specific Examples | Critical Function & Notes |
|---|---|---|
| CRISPR-Cas Systems | S. pyogenes Cas9 (SpCas9), Cas12a (Cpf1), dCas9 for CRISPRi. | SpCas9 is most widely used. dCas9 enables gene repression without cleavage for modulating QS [25] [48]. |
| Guide RNA Design Tools | CRISPOR, CHOPCHOP, Benchling. | Algorithms identify specific, high-efficiency gRNA sequences with minimal off-target effects for target genes (e.g., lasR, mecA) [51]. |
| Nanocarrier Components | Cationic lipids (DOTAP, DLIN-MC3-DMA), polymers (PLGA, PEI), gold nanorods. | Cationic materials enhance complexation with nucleic acids; PLGA allows for sustained release of antibiotics [11] [49]. |
| Antibiotics for Co-delivery | Tobramycin, Colistin, Ciprofloxacin, Vancomycin. | Selection is based on the resistance profile of the target pathogen and the specific resistance gene being disrupted by CRISPR. |
| Biofilm Assay Kits | Commercial crystal violet kits, resazurin-based viability assays (AlamarBlue). | Standardize the quantification of biofilm biomass and metabolic activity, enabling high-throughput screening [47]. |
| Bacterial Strains | WT pathogens (PAO1, MRSA USA300), isogenic mutant strains. | Mutants (e.g., lasI knockout) serve as essential controls to validate the mechanism of action of QS-targeting constructs. |
The co-delivery of CRISPR constructs and conventional antibiotics represents a paradigm shift in our approach to treating resilient biofilm-mediated infections. By leveraging the precision of CRISPR to surgically disable bacterial resistance and virulence machinery and synergistically combining it with the brute force of traditional antibiotics, this strategy offers a path to revitalizing our antimicrobial arsenal. While challenges in in vivo delivery efficiency, safety, and regulatory approval remain, advancements in nanoparticle design and a deepening understanding of bacterial genetics are rapidly overcoming these hurdles. The integration of this approach with artificial intelligence for predictive gRNA design and the development of smart, stimulus-responsive nanocarriers will further define the future of precision antimicrobial therapy, turning the tide against antimicrobial resistance.
In Vitro and In Vivo Models for Validating Anti-Biofilm Efficacy
The persistence of biofilm-associated infections poses a significant challenge in clinical settings, accounting for approximately 65-80% of all microbial infections and demonstrating dramatically increased antibiotic tolerance—up to 1,000-fold greater than their planktonic counterparts [24] [44]. This resilience necessitates robust experimental models for validating novel anti-biofilm strategies, including emerging CRISPR-based technologies that target bacterial quorum sensing (QS) and biofilm formation pathways. The validation pipeline for these innovative therapeutic approaches requires carefully designed in vitro and in vivo models that can accurately assess efficacy across different biological complexities [52]. This technical guide provides researchers and drug development professionals with comprehensive methodologies for evaluating anti-biofilm interventions, with particular emphasis on models relevant to assessing CRISPR-based approaches that target the molecular mechanisms underpinning biofilm-mediated resistance.
Biofilm Biology and CRISPR Targeting Strategies
Biofilm Architecture and Resistance Mechanisms
Bacterial biofilms are structured microbial communities encased within a self-produced extracellular polymeric substance (EPS) composed of polysaccharides, proteins, and extracellular DNA (eDNA) [24] [44]. This complex matrix creates a physical barrier that limits antibiotic penetration while establishing heterogeneous microenvironments with varying metabolic activity, pH, and oxygen tension [24]. The biofilm life cycle progresses through initial attachment, microcolony formation, maturation, and active dispersal phases, each regulated by distinct genetic networks and signaling pathways [25].
A critical mechanism driving biofilm resilience is phenotypic heterogeneity, where subpopulations of dormant "persister" cells exhibit exceptional antibiotic tolerance [44]. Additionally, the biofilm matrix facilitates horizontal gene transfer (HGT), accelerating the dissemination of antibiotic resistance genes (ARGs) among bacterial populations [53]. These combined mechanisms—physical barrier function, metabolic heterogeneity, and enhanced genetic exchange—render conventional antibiotic therapies largely ineffective against biofilm-associated infections.
CRISPR-Based Interventions Against Biofilms
CRISPR-Cas systems offer precision approaches for combating biofilms through multiple strategic applications:
Precision Gene Editing: CRISPR-Cas9 enables targeted disruption of essential biofilm-related genes, including those encoding adhesion proteins, EPS synthesis enzymes, and QS regulatory components [25] [44].
Gene Regulation: Catalytically inactive Cas9 (dCas9) systems facilitate CRISPR interference (CRISPRi) and CRISPR activation (CRISPRa) for tunable modulation of biofilm gene expression without permanent genetic alterations [25].
Antimicrobial Programming: Sequence-specific targeting of bacterial chromosomes or antibiotic resistance genes allows selective elimination of pathogens or resensitization to conventional antibiotics [25] [53].
Mobile Genetic Element Control: Native CRISPR-Cas systems in bacteria can restrict HGT of ARGs, while engineered versions can precisely degrade resistance plasmids [53].
The following diagram illustrates key CRISPR targeting strategies for disrupting biofilm formation and function:
In Vitro Models for Anti-Biofilm Evaluation
In vitro models provide controlled, reproducible systems for initial screening of anti-biofilm efficacy. Standardization is critical, as methodological variations significantly impact quantitative outcomes—a single antimicrobial agent tested across different laboratories showed up to 3,000-fold differences in reported efficacy [52].
Standardized Quantitative Assays
Microtiter Plate-Based Assays represent the foundational approach for high-throughput screening of anti-biofilm compounds:
Table 1: Core In Vitro Biofilm Assessment Methods
| Method | Key Measurements | Applications | Considerations |
|---|---|---|---|
| Crystal Violet Staining | Total biofilm biomass | High-throughput screening of anti-biofilm compounds | Does not distinguish live/dead cells |
| Calgary Biofilm Device | Minimum Biofilm Eradication Concentration (MBEC) | Antibiotic susceptibility testing | Standardized surface area for reproducibility |
| AntiBioVol Assay | Efficacy of volatile compounds against pre-formed biofilms | Testing essential oils, disinfectants | Uses 24-well plates for controlled vapor exposure [54] |
| Confocal Laser Scanning Microscopy (CLSM) | 3D architecture, biofilm thickness, live/dead distribution | Structural analysis of biofilm disruption | Requires fluorescent staining; specialized equipment |
The AntiBioVol assay represents a specialized approach for evaluating volatile anti-biofilm compounds. This method employs two 24-well plates—Plate A containing agar plugs and Plate B hosting biofilm-grown agar discs—sealed together to create a controlled vapor chamber [54]. This system enables quantitative assessment of biofilm reduction following exposure to test compounds, with specific advantages including standardized surface area, controlled atmosphere, and compatibility with high-replication experimental designs.
Advanced In Vitro Systems
More sophisticated in vitro models incorporate relevant surface materials and hydrodynamic conditions:
Flow Cell Systems: Continuous nutrient flow mimics in vivo conditions, enabling real-time imaging of biofilm development and treatment responses [52].
Biofilm Reactors: Rotating disk and CDC biofilm reactors generate high-throughput, reproducible biofilms for efficacy testing under controlled shear stress [52].
Co-culture Models: Incorporating multiple bacterial species or host cells better represents the complex interactions in clinical biofilm infections.
In Vivo Models for Anti-Biofilm Validation
In vivo models provide essential assessment of anti-biofilm efficacy in biologically complex environments, incorporating host immune responses and tissue-specific factors.
Non-Mammalian Models
Galleria mellonella (wax moth larvae) offer an ethically favorable, cost-effective initial in vivo screening platform:
Infection Protocol: Larvae (200-300 mg weight) are inoculated with 1.0×10^6 CFU of biofilm-forming bacteria via proleg injection, followed by anti-biofilm treatment administration [14].
Endpoint Measurements: Survival rates over 96 hours, melanization response as an immune indicator, and bacterial burden quantification through homogenate plating [14].
CRISPR Application: This model successfully demonstrated reduced virulence of Acinetobacter baumannii following cas3 gene knockout, with larval survival increasing from 10% (wildtype) to 50% (Δcas3 mutant) at 96 hours post-infection [14].
Mammalian Models
Mammalian systems provide critical translational data on anti-biofilm efficacy in immunologically complex environments:
Table 2: Mammalian In Vivo Models for Biofilm Research
| Model System | Infection Site | Key Parameters | Relevance to CRISPR Studies |
|---|---|---|---|
| Murine Catheter Model | Subcutaneous catheter | Bacterial load on explanted device, histopathology | Testing surface-coated CRISPR formulations |
| Mouse Pneumonia Model | Lung | Bacterial load in homogenates, cytokine levels, inflammation scoring | Assessing CRISPR delivery to respiratory biofilms |
| Mouse Bacteremia Model | Systemic circulation | Survival curves, organ bacterial loads, serum cytokine profiling | Evaluating efficacy against disseminated infections |
For murine bacteremia models, animals are infected intravenously with approximately 1×10^7 CFU of biofilm-forming bacteria. Anti-biofilm treatments are administered systemically, with efficacy assessed through survival monitoring, bacterial enumeration in spleen and liver, and cytokine profiling [14]. These models have demonstrated the critical role of specific CRISPR-Cas components in bacterial virulence—deletion of the cas3 gene in A. baumannii significantly reduced bacterial loads in murine organs and decreased serum cytokine levels, confirming its importance in pathogenesis [14].
Experimental Workflow for CRISPR-Based Anti-Biofilm Testing
The following diagram outlines a comprehensive workflow for evaluating CRISPR-based anti-biofilm strategies:
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Anti-Biofilm CRISPR Research
| Reagent/Category | Specific Examples | Research Application |
|---|---|---|
| CRISPR Delivery Systems | Liposomal nanoparticles, Gold nanoparticles, Phage-based vectors | Enhancing cellular uptake of CRISPR components through biofilm matrix [24] |
| Essential Controls | Non-targeting gRNA, Empty vector, Cas9-only | Distinguishing specific from non-specific effects in CRISPR experiments |
| Biofilm Staining Reagents | Crystal violet, SYTO9/propidium iodide, Alexa Fluor-dextran | Quantifying biomass and visualizing viability/architecture [14] |
| Model Organisms | Galleria mellonella, Specific pathogen-free mice | Providing tiered in vivo testing platforms with increasing complexity [14] |
| Culture Media | Brain Heart Infusion (BHI), Tryptic Soy Broth (TSB) | Supporting robust biofilm growth under standardized conditions [54] |
Data Documentation and Reporting Standards
Comprehensive reporting enables appropriate interpretation and replication of anti-biofilm studies:
Quantitative Metrics: Report log reduction values, surface area-to-volume ratios, and biofilm areal cell density for all efficacy experiments [52].
Methodological Details: Document specific experimental parameters including surface materials, hydrodynamic conditions, and biofilm maturation time [52].
Control Data: Include benchmark antimicrobial agents to contextualize novel treatment efficacy and validate experimental conditions [52].
Dose-Response Relationships: Conduct measurements across multiple treatment concentrations or exposure durations to establish robust efficacy profiles [52].
Advanced imaging approaches, particularly Confocal Laser Scanning Microscopy (CLSM) with appropriate fluorescent tags (e.g., SYTO9 for bacterial cells, Alexa Fluor-conjugated dextran for EPS matrix), provide critical three-dimensional structural data on biofilm disruption following CRISPR-based interventions [14].
Robust validation of anti-biofilm therapies requires complementary in vitro and in vivo models that assess efficacy across biological complexities. CRISPR-based technologies represent a promising frontier for precision targeting of biofilm regulation mechanisms, particularly quorum sensing pathways and virulence gene networks. Standardized experimental approaches, appropriate model selection, and comprehensive reporting are essential for generating translatable data on anti-biofilm efficacy. As CRISPR delivery systems advance—particularly nanoparticle-based platforms that enhance penetration through the biofilm matrix—these validation frameworks will be crucial for translating innovative approaches into clinical solutions for persistent biofilm-associated infections.
Navigating the Hurdles: Off-Target Effects, Delivery Efficiency, and Specificity Optimization
The application of CRISPR-Cas systems has expanded beyond basic genome editing to become a pivotal tool in bacterial physiology research, including the study of quorum sensing (QS) and biofilm formation [25] [20]. Biofilms, which are structured communities of microorganisms embedded in an extracellular polymeric substance, are a major contributor to antimicrobial resistance and chronic infections [11] [55]. The precision of CRISPR-Cas tools allows researchers to dissect the complex genetic networks that govern biofilm development and bacterial communication [25]. However, the safety and reliability of these CRISPR-based interventions are contingent upon their specificity. Unintended off-target effects can lead to erroneous scientific conclusions in basic research and pose significant safety risks in therapeutic development [56] [57]. Therefore, a rigorous assessment of off-target activity is indispensable. This analysis focuses on three prominent off-target discovery methods—GUIDE-seq, CIRCLE-seq, and modern in silico predictors—evaluating their methodologies, performance, and applicability within the specific context of CRISPR-based biofilm and QS research.
Off-target detection methods can be broadly classified into cellular (in cellula), biochemical (in vitro), and computational (in silico) approaches [56]. Cellular methods, such as GUIDE-seq, detect edits within the native cellular environment, capturing the influence of chromatin structure, DNA repair pathways, and cellular physiology. In contrast, biochemical methods like CIRCLE-seq use purified genomic DNA, offering ultra-sensitive and comprehensive mapping of potential cleavage sites in the absence of cellular context. Computational predictors have evolved from simple sequence alignment tools to sophisticated deep-learning models that can forecast off-target activity prior to experimentation [58] [59]. The following sections provide a detailed examination of each method, with a particular emphasis on their utility for research involving bacterial pathogens and biofilm-forming species.
Table 1: Key Characteristics of Off-Target Discovery Methods
| Method | Approach | Input Material | Detection Context | Key Strengths | Primary Limitations |
|---|---|---|---|---|---|
| GUIDE-seq [56] | Cellular (Unbiased) | Living cells (edited) | Native chromatin & repair | Reflects true cellular activity; identifies biologically relevant edits | Requires efficient delivery; less sensitive; may miss rare sites |
| CIRCLE-seq [56] | Biochemical (Unbiased) | Purified genomic DNA | Naked DNA (no chromatin) | Ultra-sensitive; comprehensive; standardized | May overestimate cleavage; lacks biological context |
| In Silico Predictors (e.g., CCLMoff, DNABERT-Epi) [58] [59] | Computational | Genome sequence & sgRNA | Predicted sites (based on models) | Fast, inexpensive; useful for guide design & prior knowledge | Predictions only; may not capture all biological variables |
Detailed Analysis of GUIDE-seq
Methodology and Workflow
GUIDE-seq (Genome-wide, Unbiased Identification of DSBs Enabled by Sequencing) is a cellular-based method that directly captures and sequences double-strand breaks (DSBs) within living cells [56]. The core of this technique involves the incorporation of a double-stranded oligonucleotide tag into DSBs generated by the CRISPR-Cas9 nuclease during the cell's repair process.
Experimental Protocol:
- Co-delivery: The cells of interest (e.g., bacterial cultures or eukaryotic cell lines used to model bacterial pathways) are co-transfected with three components:
- Plasmids encoding the Cas9 nuclease and the target-specific sgRNA.
- The proprietary GUIDE-seq oligonucleotide tag.
- Tag Integration: When Cas9 induces a DSB, the cellular repair machinery incorporates the oligonucleotide tag into the break site.
- Genomic DNA Extraction & Library Prep: Genomic DNA is harvested and sheared. DNA fragments containing the integrated tag are then enriched and prepared for next-generation sequencing (NGS) using a tag-specific primer.
- Sequencing & Data Analysis: The resulting libraries are sequenced. Bioinformatic analysis identifies the genomic sequences flanking the integrated tags, thereby mapping the locations of both on-target and off-target DSBs genome-wide [56].
GUIDE-seq Workflow: From delivery to genome-wide DSB mapping.
Application in Biofilm and Quorum Sensing Research
GUIDE-seq is particularly valuable in a bacterial research context because it identifies off-target effects within the native physiological conditions of the cell. When using CRISPRi/a (interference/activation) to modulate the expression of QS genes like luxR or lasI, or biofilm structural genes such as pel or psl, GUIDE-seq can confirm that the observed phenotypic changes (e.g., altered biofilm architecture or virulence) are a direct result of on-target regulation and not confounding off-target mutations [25]. Its ability to detect edits in a chromatin context is especially relevant for studying host-pathogen interactions in eukaryotic infection models, where bacterial genes are introduced.
Detailed Analysis of CIRCLE-seq
Methodology and Workflow
CIRCLE-seq (Circularization for In vitro Reporting of Cleavage Effects by Sequencing) is a highly sensitive, biochemical assay performed on purified genomic DNA [56]. Its key innovation is the circularization of DNA and enzymatic enrichment of cleavage products, which allows for the detection of extremely rare off-target sites.
Experimental Protocol:
- Genomic DNA Preparation & Circularization: High-molecular-weight genomic DNA is extracted from the target cells or bacterial strain. This DNA is then fragmented and circularized using DNA ligase.
- In Vitro Cleavage: The circularized DNA library is treated with a pre-assembled Cas9-sgRNA ribonucleoprotein (RNP) complex under optimal reaction conditions.
- Exonuclease Enrichment: A crucial step follows where an exonuclease is added to digest linear, non-cleaved DNA. The key advantage is that cleaved DNA fragments, which become linearized upon Cas9 cutting, are protected from exonuclease digestion because they remain part of the larger circular molecules. This step dramatically enriches for sequences that were cut by Cas9.
- Linearization & Library Prep: The enriched, cleaved fragments are then released from the circular backbone (e.g., by re-cutting with the original restriction enzyme used for fragmentation). These fragments are used to construct an NGS library.
- Sequencing & Analysis: High-throughput sequencing and subsequent analysis reveal the sequences of all DNA ends generated by Cas9 cleavage, providing a genome-wide profile of potential off-target sites [56].
CIRCLE-seq Workflow: In vitro cleavage and enzymatic enrichment.
Application in Biofilm and Quorum Sensing Research
CIRCLE-seq's primary advantage is its exceptional sensitivity, which allows it to identify potential off-target sites that might be missed by less sensitive cellular assays. For foundational research aimed at characterizing a novel Cas nuclease or sgRNA for use in biofilm studies, CIRCLE-seq provides a comprehensive landscape of its cleavage preferences. This is crucial for establishing the reliability of genetic tools before they are deployed in complex experiments, such as engineering probiotic strains to express CRISPR-based antimicrobials that target specific biofilm formation genes in pathogens [11] [55]. However, researchers must be cautious, as its biochemical nature lacks cellular context and may identify sites that are not cleaved in vivo.
Detailed Analysis of In Silico Predictors
Evolution and Methodology
In silico prediction tools have rapidly advanced from simple alignment-based algorithms to sophisticated deep learning models that leverage large-scale genomic and epigenetic data [58] [59]. Early tools like Cas-OFFinder and MIT CRISPR design relied on sequence homology and mismatch counting. The current state-of-the-art, exemplified by tools such as CCLMoff and DNABERT-Epi, utilizes transformer-based architectures pre-trained on vast genomic datasets.
Model Architecture and Workflow:
- CCLMoff: This framework employs a pretrained RNA language model (RNA-FM) to capture mutual sequence information between the sgRNA and potential target DNA sites. It formulates off-target prediction as a question-answering task, where the sgRNA is the "question" and a candidate DNA sequence is the "answer." The model is trained on a comprehensive dataset compiled from 13 different genome-wide off-target detection techniques, enabling strong generalization [58].
- DNABERT-Epi: This model integrates a DNA foundation model (DNABERT), which is pre-trained on the entire human genome, with epigenetic features such as H3K4me3, H3K27ac, and ATAC-seq data. The integration of chromatin accessibility information has been shown to significantly enhance predictive accuracy for off-target effects in cellular environments [59].
Application in Biofilm and Quorum Sensing Research
The primary utility of in silico predictors lies in their speed and cost-effectiveness during the sgRNA design phase. Before committing to laborious wet-lab experiments, researchers can screen dozens of candidate sgRNAs targeting a QS gene (e.g., rhlR in P. aeruginosa) to select the one with the lowest predicted off-target risk. Furthermore, the ability of models like DNABERT-Epi to incorporate epigenetic data makes them increasingly relevant for studies in bacterial species where chromatin structure influences gene expression. For translational applications, such as designing CRISPR-based antimicrobials that precisely target antibiotic resistance genes within a biofilm community, these tools are indispensable for maximizing on-target specificity from the outset [25] [11].
Table 2: Performance Comparison of Off-Target Detection Methods
| Performance Metric | GUIDE-seq | CIRCLE-seq | In Silico Predictors |
|---|---|---|---|
| Sensitivity | Moderate to High (in cells) | Very High (in vitro) | Varies; High for modern models |
| Specificity | High (biologically relevant) | Lower (may include non-physiological sites) | Varies; improving with deep learning |
| Throughput | Medium | High | Very High |
| Workflow Complexity | High (requires cell culture & delivery) | Medium (biochemical assay) | Low (computational) |
| Cost | High | Medium | Low |
| Best Use Case | Validation of biologically relevant off-targets | Broad, sensitive discovery during tool characterization | Preliminary sgRNA screening and design |
Integrated Workflow for Off-Target Assessment in Biofilm Research
To ensure the highest level of precision in CRISPR-based biofilm and QS research, a sequential, multi-method approach is recommended by leading authorities, including the FDA [56]. The following integrated workflow leverages the strengths of each method:
- sgRNA Design & In Silico Screening: Initiate the process with a state-of-the-art in silico predictor (e.g., CCLMoff or DNABERT-Epi) to design and rank multiple sgRNAs against the target gene (e.g., a quorum sensing regulator). Select the top candidate with the lowest predicted off-target risk for experimental validation [58] [59].
- Broad Experimental Discovery: Perform CIRCLE-seq using genomic DNA purified from the relevant bacterial strain or host cell line. This step provides a sensitive, genome-wide map of all potential off-target sites for the selected sgRNA, establishing a comprehensive "risk list" [56].
- Biological Validation: Translate the potential off-target sites identified by CIRCLE-seq into biologically relevant findings using GUIDE-seq. This confirms which of the in vitro sites are actually cleaved in a living cellular environment, under conditions that mimic the intended experimental setup [56].
- Final Functional Assays: With a validated understanding of off-target activity, proceed with functional CRISPR experiments (e.g., gene knockout with Cas9 or gene repression with dCas9) and confidently attribute phenotypic changes in biofilm formation or virulence to the on-target genetic modification [25].
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of the described methods requires a suite of specialized reagents and tools.
Table 3: Key Research Reagent Solutions for Off-Target Analysis
| Reagent / Tool | Function | Example Application |
|---|---|---|
| GUIDE-seq Oligonucleotide | A double-stranded oligo that integrates into Cas9-induced DSBs, serving as a tag for sequencing library preparation [56]. | The core reagent for the GUIDE-seq protocol to capture breaks in living cells. |
| Cas9 Nuclease (High-Purity) | The engineered nuclease protein that, when complexed with sgRNA, induces DSBs at specific genomic sites. | Essential for all biochemical (CIRCLE-seq) and many cellular (GUIDE-seq) off-target detection assays. |
| Next-Generation Sequencing Platform | Enables high-throughput sequencing of enriched DNA libraries to map off-target sites genome-wide. | Required for the final readout of GUIDE-seq, CIRCLE-seq, and related methods. |
| CCLMoff Software | A deep learning framework for predicting CRISPR/Cas9 off-target effects, leveraging a pretrained RNA language model [58]. | Used for in silico sgRNA design and pre-screening to select optimal guides with minimal predicted off-target effects. |
| Epigenetic Data (e.g., ATAC-seq) | Provides information on chromatin accessibility, which can be integrated into models like DNABERT-Epi to improve off-target prediction in cellular contexts [59]. | Enhances the accuracy of computational predictions for experiments in eukaryotic cells or complex microbial communities. |
The precision of CRISPR-Cas systems has made them invaluable for dissecting intricate bacterial processes like quorum sensing and biofilm formation. However, this precision is not absolute, making rigorous off-target assessment a critical component of the research pipeline. GUIDE-seq, CIRCLE-seq, and modern in silico predictors each offer distinct advantages and limitations. GUIDE-seq provides the gold standard for biological validation, CIRCLE-seq offers unmatched sensitivity for discovery, and in silico tools deliver unparalleled speed for design. An integrated approach, combining computational prediction with biochemical and cellular validation, is the most robust strategy to ensure the fidelity and safety of CRISPR-based interventions. As the field progresses towards clinical applications, such as CRISPR-engineered probiotics or sequence-specific antimicrobials, the standardized use of these complementary off-target discovery tools will be paramount in translating laboratory findings into reliable therapeutic strategies against biofilm-associated infections.
The application of the CRISPR-Cas9 system to study bacterial quorum sensing and biofilm formation represents a paradigm shift in microbial genetics. However, the utility of this powerful tool is contingent upon its precision. In the context of complex bacterial communities, off-target effects can lead to misleading phenotypic data and confound the interpretation of gene function in pathways critical for biofilm maturation and cell-cell communication. The inherent challenge stems from the fact that standard Cas9 nucleases, when directed by a guide RNA (gRNA), can tolerate mismatches between the gRNA and target DNA sequence, potentially cleaving non-intended genomic sites [60]. This review details two foundational strategies—high-fidelity Cas9 variants and truncated gRNAs (trugRNAs)—that work synergistically to enhance the specificity of CRISPR-Cas9 interventions, thereby ensuring the accuracy of findings in biofilm and quorum sensing research.
Core Mechanisms for Enhancing CRISPR-Cas9 Specificity
Truncated gRNAs (trugRNAs)
Truncated gRNAs represent a simple yet powerful modification to the standard CRISPR-Cas9 system to minimize off-target interactions. Conventional gRNAs contain 20 nucleotides of target-complementary sequence. Tru-gRNAs are shortened at the 5' end of this complementary region, typically to 17 or 18 nucleotides [60].
- Mechanism of Action: The reduction in length decreases the energy of hybridization between the gRNA and its DNA target. This makes the binding event more sensitive to mismatches, particularly at the 3' end of the target sequence, which is crucial for cleavage. A shorter complementary region means that any single mismatch constitutes a more significant percentage of the total binding energy, making off-target binding at sites with subtle sequence variations less thermodynamically favorable [60].
- Efficacy: Research has demonstrated that tru-gRNAs can reduce undesired mutagenesis at off-target sites by 5,000-fold or more without compromising on-target editing efficiency. For some specific off-target sites, the improvement in specificity can exceed 10,000-fold [60].
Table 1: Performance Comparison of Standard gRNAs vs. Truncated gRNAs
| Feature | Standard gRNA (20 nt) | Truncated gRNA (17-18 nt) |
|---|---|---|
| Complementarity Length | 20 nucleotides | 17-18 nucleotides |
| On-Target Efficiency | High | Comparable to standard gRNAs |
| Off-Target Mutation Frequency | Can be significant | Up to 5,000-fold reduction |
| Mismatch Sensitivity | Moderate | Highly increased, especially to 3' end mismatches |
| Applicability | Universal | Requires empirical validation for each target site |
High-Fidelity Cas9 Variants
While tru-gRNAs modify the RNA component, high-fidelity Cas9 variants are engineered versions of the Cas9 protein with mutated amino acids that tighten the enzyme's fidelity. These variants are designed to be less tolerant of gRNA-DNA mismatches.
- Mechanism of Action: These mutations often involve residues that contact the DNA non-target strand. The alterations destabilize the R-loop formation—a key intermediate step in Cas9 activation—unless the gRNA-DNA pairing is perfectly complementary. This creates a higher energy barrier for cleavage, ensuring that only true on-target sites are cut [60].
- Synergy with Tru-gRNAs: The specificity-enhancing effects of high-fidelity Cas9 variants and tru-gRNAs are complementary. Using them in combination can lead to multiplicative reductions in off-target effects, providing a robust solution for applications requiring the highest level of precision, such as dissecting genetic networks in quorum sensing [60].
Experimental Protocols for Specificity Optimization
Designing and Testing Truncated gRNAs
A systematic approach is required to implement tru-gRNAs effectively in bacterial biofilm studies.
Protocol: In vitro Evaluation of Tru-gRNA Efficacy
- Target Selection: Identify the gene of interest (e.g., a quorum sensing regulator like lasR in P. aeruginosa).
- gRNA Design: Design a standard 20-nucleotide gRNA. Generate a series of truncated versions (18-nt, 17-nt) from the 5' end of the target-complementary sequence.
- Cloning: Clone each gRNA construct into an appropriate CRISPR-Cas9 delivery vector suitable for your bacterial system.
- Transformation: Introduce the vectors into the target bacterial strain.
- On-Target Efficiency Assay:
- Quantify editing efficiency at the intended locus using a mismatch detection assay (e.g., T7 Endonuclease I) or by high-throughput sequencing of the target region.
- Select tru-gRNAs that maintain ≥80% of the standard gRNA's editing efficiency.
- Off-Target Profiling:
- Use in silico prediction tools (e.g., Cas-OFFinder) to identify potential off-target sites in the genome.
- Amplify and sequence these putative off-target loci from treated and control cultures.
- Calculate the indel mutation frequency to confirm the reduction in off-target activity with the selected tru-gRNA.
Specificity Validation in Biofilm Models
Once high-specificity gRNA/Cas9 combinations are identified, their functional impact must be validated in a biologically relevant context.
Protocol: Functional Validation in a Biofilm Assay
- Strain Engineering: Create bacterial strains harboring the high-fidelity CRISPR-Cas9 system targeting a biofilm-related gene (e.g., for EPS production).
- Biofilm Cultivation: Grow the engineered and control strains in a standardized biofilm model (e.g., using a microtiter plate or a flow-cell system).
- Phenotypic Analysis:
- Biomass Quantification: Use crystal violet staining to measure total biofilm biomass.
- Viability Assessment: Perform colony-forming unit (CFU) counts or use a live/dead viability stain coupled with confocal microscopy.
- Structural Analysis: Visualize the biofilm architecture using Scanning Electron Microscopy (SEM) or Confocal Laser Scanning Microscopy (CLSM).
- Transcriptional Analysis: Use RT-qPCR to measure the expression of downstream genes in the targeted pathway (e.g., other quorum sensing-controlled genes) to confirm the specific knockdown effect.
The Scientist's Toolkit: Essential Reagents for Specificity-Focused Research
Table 2: Key Research Reagent Solutions for Optimizing CRISPR-Cas9 Specificity
| Reagent / Tool | Function & Application | Justification |
|---|---|---|
| High-Fidelity Cas9 Expression Vector | Provides a genetically encoded source of a fidelity-enhanced Cas9 nuclease (e.g., eSpCas9, SpCas9-HF1). | Foundation for reducing off-target cleavage while maintaining robust on-target activity in bacterial cells. |
| Modular gRNA Cloning Kit | Enables rapid and efficient cloning of both standard and truncated gRNA sequences. | Essential for the parallel construction and testing of multiple gRNA designs for a single target. |
| T7 Endonuclease I Assay Kit | Detects indel mutations at the target site by recognizing and cleaving DNA heteroduplexes. | A cost-effective and rapid method for initial screening of on-target editing efficiency. |
| Next-Generation Sequencing (NGS) Service/Library Prep Kit | Provides a comprehensive, genome-wide view of both on-target and off-target editing events. | The gold standard for unbiased, sensitive assessment of CRISPR-Cas9 specificity; critical for validating tru-gRNAs and high-fidelity variants. |
| In silico Off-Target Prediction Software (e.g., Cas-OFFinder) | Identifies potential off-target genomic sites based on sequence similarity to the gRNA. | Guides the experimental design for off-target validation, allowing researchers to focus on the most likely problematic loci. |
Visualizing Specificity Enhancement Strategies
The following diagrams illustrate the core concepts and workflows for optimizing CRISPR-Cas9 specificity.
Mechanism of Truncated gRNAs
Specificity Optimization Workflow
The precision required to deconvolute the genetic underpinnings of bacterial quorum sensing and biofilm formation demands CRISPR-Cas9 tools of the highest specificity. The combined use of truncated gRNAs and high-fidelity Cas9 variants provides a robust, widely applicable strategy to minimize off-target effects, thereby ensuring that observed phenotypic changes are directly attributable to the targeted genetic modification. As CRISPR-based technologies continue to evolve, integrating these specificity-enhanced systems with advanced delivery mechanisms, such as nanoparticles [11], will further empower researchers to conduct definitive functional genomics studies within complex microbial communities.
Bacterial biofilms represent a significant obstacle in treating persistent infections, largely due to their complex extracellular polymeric substance (EPS) matrix that limits antibiotic penetration and enhances antimicrobial resistance. This protective matrix creates microenvironments where bacteria can survive at antibiotic concentrations up to 1000-fold higher than those required to eliminate their planktonic counterparts [24]. Within this context, Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-based gene editing has emerged as a revolutionary tool for precision targeting of the fundamental genetic mechanisms governing biofilm formation and maintenance [61] [24]. The CRISPR-Cas system, derived from bacterial immune mechanisms against viral DNA, utilizes a guide RNA (gRNA) to direct Cas nucleases to specific genetic sequences, enabling targeted disruption of antibiotic resistance genes, quorum sensing (QS) pathways, and biofilm-regulating factors [62] [24].
However, the clinical application of CRISPR-based antibacterials faces significant delivery challenges, particularly in efficiently traversing biofilm barriers to reach bacterial populations. Nanoparticles present an innovative solution, serving as advanced carriers for CRISPR/Cas9 components while exhibiting intrinsic antibacterial properties [24]. The synergistic integration of CRISPR technology with engineered nanoparticles creates a multifaceted approach that attacks biofilms through both genetic disruption and enhanced physical penetration. This combination strategy not only improves the precision of CRISPR/Cas9 targeting but also overcomes the physical barriers posed by biofilm matrices, thereby potentially revolutionizing the treatment outcomes of resistant infections [24]. This technical guide explores the engineering principles behind nanoparticle design for optimized biofilm penetration and CRISPR delivery, providing researchers with methodologies and frameworks to advance this promising field.
Nanoparticle Engineering for Enhanced Biofilm Penetration
Material Composition and Surface Properties
The strategic design of nanoparticle materials directly dictates their penetration capability through biofilm matrices. Lipid-based nanoparticles (LNPs), particularly those incorporating ionizable lipids, have demonstrated exceptional efficacy in encapsulating and delivering large CRISPR components. These LNPs undergo a charge-based mechanism where they remain neutral in the bloodstream to reduce toxicity but become positively charged in acidic environments like endosomes, facilitating endosomal escape and payload release [63]. The four-component structure of LNPs—consisting of ionizable lipids, phospholipids, cholesterol, and PEG-lipids—can be precisely tuned to optimize encapsulation efficiency, stability, and biofilm interaction [63] [24]. For instance, incorporating ester bonds or disulfide bridges into ionizable lipids enhances biodegradability and reduces potential toxicity, as demonstrated by lipids like FTT5 and ssPalmO-Phe [63].
Surface functionalization plays an equally critical role in penetration efficiency. Polyethylene glycol (PEG) coating provides a "stealth" effect that reduces opsonization and extends circulation time, while enabling smaller particle sizes that enhance diffusion through the heterogeneous biofilm architecture [64] [63]. Additionally, engineering nanoparticles with targeting ligands such as lectins or peptides that recognize specific bacterial surface components can further improve localization and cellular uptake within biofilms [64]. Recent advances have also explored the incorporation of biofilm matrix-degrading enzymes like depolymerase directly onto nanoparticle surfaces, creating a penetration strategy that actively dismantles structural barriers while delivering therapeutic payloads [65].
Table 1: Nanoparticle Types and Their Characteristics for Biofilm Penetration
| Nanoparticle Type | Key Components | Size Range | Penetration Advantages | CRISPR Payload Capacity |
|---|---|---|---|---|
| Lipid Nanoparticles (LNPs) | Ionizable lipids, Phospholipids, Cholesterol, PEG-lipids [63] | 50-200 nm [64] | Enhanced cellular uptake, endosomal escape, tunable surface charge [63] | High (Cas9 mRNA, sgRNA, RNP complexes) [63] |
| Gold Nanoparticles (AuNPs) | Gold core, surface functionalized with polymers or peptides [64] | 10-100 nm | Plasmonic properties for photothermal activation, excellent biocompatibility [64] | Moderate (typically smaller CRISPR systems) [24] |
| Polymeric Nanoparticles | PLGA, Chitosan, PEI [64] | 80-250 nm | Controlled release kinetics, mucosal adhesion (chitosan) [64] | High (can encapsulate large CRISPR complexes) [64] |
| Hybrid Systems | Lipid-polymer composites, exosome-inspired [64] [63] | 100-200 nm | Combined advantages of multiple materials, enhanced targeting [64] | High with improved stability [64] |
Size, Charge, and Morphology Optimization
The physical parameters of nanoparticles—particularly size, surface charge, and shape—fundamentally influence their ability to navigate the complex architecture of biofilms. Size optimization is critical, with particles in the 50-200 nm range demonstrating optimal penetration through the water channels and heterogenous matrix of typical biofilms [64]. This size range represents a strategic balance, sufficiently small to navigate structural barriers yet large enough to carry meaningful CRISPR payloads.
Surface charge engineering represents another crucial design consideration. While slightly positive charges (typically +5 to +30 mV) enhance interaction with negatively charged bacterial membranes and biofilm components, excessively cationic surfaces can lead to non-specific binding and limited penetration depth [24]. The development of charge-switching nanoparticles that adapt their surface properties in response to environmental stimuli like pH offers a sophisticated approach to this challenge, maintaining stability during circulation while enhancing biofilm adhesion and cellular uptake upon arrival at the target site [63].
Morphological innovations beyond traditional spherical nanoparticles show particular promise for biofilm penetration. Anisotropic particles with elongated or rod-like shapes demonstrate enhanced diffusion capabilities through biological barriers, while some studies suggest that smaller, deformable nanoparticles can better navigate the constrictive pores within biofilm matrices [64]. These engineering considerations collectively contribute to the development of nanoparticles specifically optimized for the unique challenges of biofilm penetration.
CRISPR Delivery for Targeting Quorum Sensing and Biofilm Formation
Molecular Targets in Quorum Sensing and Biofilm Regulation
The precision of CRISPR-based therapeutics enables strategic disruption of key genetic pathways governing biofilm development and maintenance. Quorum sensing (QS) genes, which facilitate bacterial cell-to-cell communication and coordinate population-level behaviors like biofilm formation, represent particularly valuable targets. The luxS gene, involved in autoinducer-2 (AI-2) synthesis, has been successfully targeted using CRISPR interference (CRISPRi) in Escherichia coli, resulting in significantly inhibited biofilm formation [62]. This approach utilizes a catalytically inactive dCas9 protein that binds to target DNA without cleaving it, thereby blocking transcription through steric hindrance [61] [62].
Additional high-value targets include adhesion genes such as fimH, which encodes for type 1 fimbrial adhesion critical for initial surface attachment, and bolA, a transcriptional regulator influencing curli amyloid fibers and cellulose production [66]. Knockout studies demonstrate that ΔfimH, ΔluxS, and ΔbolA strains exhibit biofilm reduction of 75-84% compared to wild-type strains, accompanied by significant reductions in EPS production [66]. Beyond these targets, the csgD gene, which regulates curli fiber production and biofilm architecture, and pel genes, responsible for Pel exopolysaccharide biosynthesis, offer additional strategic points for intervention [61] [67].
Table 2: Key Genetic Targets for CRISPR-Based Biofilm Disruption
| Target Gene | Function | CRISPR Approach | Observed Biofilm Reduction | Additional Phenotypic Effects |
|---|---|---|---|---|
| luxS | AI-2 autoinducer synthesis in quorum sensing [62] | CRISPRi with dCas9 [62] | 77.51% [66] | Disrupted cell communication, reduced virulence [62] |
| fimH | Adhesion protein for surface attachment [66] | CRISPR/Cas9-HDR knockout [66] | 78.37%-84.17% [66] | Impaired initial attachment [66] |
| bolA | Transcriptional regulator for curli and cellulose [66] | CRISPR/Cas9-HDR knockout [66] | 75.39%-78.24% [66] | Reduced EPS matrix production [66] |
| csgD | Master regulator of biofilm formation [61] | CRISPRi [61] | Not quantified in results | Downregulation of curli and cellulose production [61] |
| pel genes | Pel exopolysaccharide biosynthesis [67] | Not specified in results | Not quantified in results | Compromised biofilm structural integrity [67] |
CRISPR System Selection and Delivery Formats
The selection of appropriate CRISPR systems represents a critical consideration in anti-biofilm strategy formulation. While the standard Streptococcus pyogenes Cas9 (SpCas9) remains widely utilized, its relatively large size (∼4.2 kb) presents packaging challenges for certain nanoparticle platforms [68]. Smaller alternatives such as Staphylococcus aureus Cas9 (SaCas9, ∼3.2 kb) provide more accommodating dimensions for nanoparticle encapsulation while maintaining robust editing capabilities [68]. Beyond traditional nucleases, advanced CRISPR tools including base editors and prime editors offer more precise genetic modifications without creating double-strand breaks, potentially reducing unintended consequences in complex microbial communities [68].
The format of CRISPR component delivery significantly influences editing efficiency and specificity. Plasmid DNA encoding both Cas9 and guide RNA represents the most straightforward approach but raises concerns about prolonged nuclease expression and potential immune responses [24]. mRNA-based delivery offers transient expression with reduced off-target risks, while preassembled Cas9 ribonucleoprotein (RNP) complexes provide the most rapid activity onset and clearance, minimizing off-target effects [63] [24]. Research indicates that RNP delivery via lipid nanoparticles demonstrates particularly high efficiency in bacterial systems, with one study reporting up to 90% reduction in Pseudomonas aeruginosa biofilm biomass using liposomal Cas9 RNP formulations [24].
Diagram Title: CRISPR-Nanoparticle System for Biofilm Disruption
Experimental Protocols and Methodologies
Protocol: CRISPRi-Mediated luxS Gene Suppression in E. coli
The following detailed protocol outlines the methodology for targeted suppression of the luxS gene using CRISPR interference, adapted from established approaches with enhancements for improved efficacy [62].
Materials and Reagents:
- pdCas9 plasmid (expressing catalytically dead Cas9)
- pgRNA plasmid (for sgRNA expression)
- E. coli clinical strain with high biofilm-forming capacity
- Luria Bertani (LB) broth and agar
- Antibiotics: ampicillin (100 μg/mL), chloramphenicol (25 μg/mL)
- Inducer: anhydrotetracycline (aTc, 2 μM)
- Transformation reagents (competent cells, heat shock materials)
- RNA extraction kit and qRT-PCR reagents
Procedure:
- sgRNA Design and Cloning:
- Design three 20-nucleotide complementary sequences targeting the luxS gene, adjacent to 5'-CCN-3' PAM sequences [62].
- Synthesize forward and reverse primers containing the target sequences with 35-nt overlaps for the dCas9 handle.
- Perform inverse PCR amplification of the pgRNA plasmid using phosphorylated primers.
- Digest template DNA with DpnI, purify PCR products, and perform blunt-end ligation.
- Transform ligated products into E. coli Top10 competent cells and verify through colony PCR and sequencing.
Co-transformation and Strain Creation:
- Isolate verified pgRNA-luxS plasmids and co-transform with pdCas9 plasmid into the target E. coli strain.
- Plate on selective media containing both ampicillin and chloramphenicol.
- Pick single colonies and validate dCas9 expression via SDS-PAGE.
Gene Suppression and Validation:
- Inoculate knockdown strains in LB broth supplemented with antibiotics and 2 μM aTc.
- Incubate at 37°C with shaking (220 rpm) for 16-24 hours.
- Extract total RNA and perform qRT-PCR to quantify luxS suppression compared to wild-type controls.
Biofilm Assessment:
- Quantify biofilm formation using crystal violet assay: stain with 0.1% crystal violet, elute with ethanol-acetone (80:20), measure OD590 [66] [62].
- Assess metabolic activity within biofilms using XTT reduction assay.
- Visualize biofilm architecture and EPS production using scanning electron microscopy (SEM).
Protocol: Lipid Nanoparticle Formulation for CRISPR Delivery
This protocol details the preparation of lipid nanoparticles optimized for encapsulation and delivery of CRISPR components to bacterial biofilms, incorporating recent advances in the field [63] [24].
Materials and Reagents:
- Ionizable lipid (e.g., FTT5, DLin-MC3-DMA)
- Helper lipids: DSPC, cholesterol
- PEG-lipid (e.g., DMG-PEG2000)
- CRISPR payload (Cas9 mRNA, sgRNA, or RNP complexes)
- Ethanol (absolute)
- Citrate buffer (10 mM, pH 4.0)
- Dialysis cassettes (MWCO 100 kDa)
- Microfluidic device or alternative mixing apparatus
Procedure:
- Lipid Solution Preparation:
- Dissolve ionizable lipid, DSPC, cholesterol, and PEG-lipid in ethanol at molar ratios of 50:10:38.5:1.5.
- Final lipid concentration should be 10-20 mM in ethanol.
Aqueous Phase Preparation:
- Dilute CRISPR components (Cas9 mRNA at 0.2 mg/mL or RNP complexes) in 10 mM citrate buffer (pH 4.0).
Nanoparticle Formation:
- Utilize microfluidic mixing with total flow rate of 12 mL/min and flow rate ratio of 3:1 (aqueous:ethanol).
- Alternatively, employ rapid pipetting or vortex-based methods for small-scale preparations.
- Immediately dilute formed nanoparticles in phosphate-buffered saline (PBS).
Purification and Characterization:
- Dialyze against PBS (pH 7.4) for 4-6 hours to remove ethanol and exchange buffer.
- Sterilize using 0.22 μm filters.
- Characterize particle size (target 80-150 nm), polydispersity index (<0.2), zeta potential (+5 to +15 mV), and encapsulation efficiency (>90%).
- Verify CRISPR activity retention through gel retardation assays.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for CRISPR-Nanoparticle Biofilm Studies
| Reagent/Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| CRISPR Plasmids | pdCas9, pgRNA [62] | Express dCas9 and target-specific sgRNA | Compatible origin and resistance; inducible promoters preferred |
| Lipid Components | Ionizable lipids (FTT5, DLin-MC3-DMA) [63] | LNP structural and functional core | Biodegradable designs (esters, disulfides) enhance safety |
| Polymer Materials | PLGA, Chitosan, PEI [64] | Polymeric NP matrix for CRISPR delivery | Molecular weight and charge density affect efficacy/toxicity |
| Metallic NPs | Gold nanoparticles (AuNPs) [64] | Photothermal-controlled release | Size and surface chemistry tune plasmonic properties |
| Characterization Kits | Zeta potential analyzers, dialysis membranes | NP physicochemical characterization | Size, PDI, and encapsulation efficiency critical for performance |
| Biofilm Assays | Crystal violet, XTT, SYTO-9/propidium iodide | Quantify biofilm biomass and viability | Combine multiple assays for comprehensive assessment |
| Imaging Tools | SEM, CLSM, TEM [66] [62] | Visualize biofilm structure and NP penetration | Sample preparation preserves native biofilm architecture |
Quantitative Efficacy Data and Performance Metrics
The integration of nanoparticle delivery systems with CRISPR technology demonstrates quantitatively superior performance compared to conventional delivery approaches. Recent studies directly comparing delivery modalities reveal that liposomal Cas9 formulations reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro, substantially outperforming non-vectored CRISPR delivery [24]. Similarly, gold nanoparticle-CRISPR hybrids demonstrate a 3.5-fold increase in gene-editing efficiency compared to non-carrier systems while promoting synergistic action with antibiotics [24].
Specific gene targeting results further validate this approach, with CRISPR/Cas9-mediated knockout of adhesion and quorum sensing genes in E. coli achieving 75-84% reduction in biofilm formation [66]. Microscopic analyses corroborate these quantitative findings, showing that mutant strains lack the extensive extracellular polymeric substance matrix observed in wild-type strains embedded in mature biofilms [66]. The application of these engineered strains to urinary catheters demonstrates significantly reduced adherence, cell aggregation, and biofilm formation compared to wild-type strains, highlighting the translational potential of this methodology [66].
Diagram Title: NP-CRISPR Development Pipeline
The strategic integration of engineered nanoparticles with CRISPR-based technologies represents a paradigm shift in our approach to combating biofilm-associated infections. By synergizing the physical penetration capabilities of nanocarriers with the genetic precision of CRISPR systems, researchers can now target the fundamental mechanisms of biofilm formation and antibiotic resistance with unprecedented specificity. The methodologies and data presented in this technical guide provide a foundation for advancing this promising field, highlighting both the substantial progress already made and the challenges that remain.
Looking forward, several emerging trends suggest exciting directions for this field. The development of increasingly sophisticated stimulus-responsive nanoparticles that activate CRISPR release in response to specific biofilm microenvironments (e.g., hypoxia, low pH, or particular enzymes) promises enhanced spatial and temporal control over therapeutic activity [64] [24]. Similarly, the creation of multi-targeting gRNA arrays that simultaneously disrupt multiple genetic pathways within biofilms may help prevent compensatory resistance mechanisms from emerging [61] [66]. As these technologies mature, their translation toward clinical application will require careful attention to safety profiles, manufacturing scalability, and regulatory considerations. Through continued interdisciplinary collaboration between materials science, microbiology, and molecular biology, the integrated CRISPR-nanoparticle platform holds exceptional potential to overcome the persistent challenge of biofilm-mediated resistance ushering in a new era of precision antimicrobial therapeutics.
The application of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) systems in bacterial research has revolutionized our ability to interrogate and manipulate microbial physiology, particularly in the contexts of quorum sensing and biofilm formation. However, a significant challenge persists: the variable efficacy of CRISPR interventions across different bacterial strains and CRISPR system types. This technical guide examines the molecular foundations of this variability, providing researchers and drug development professionals with frameworks to predict, measure, and overcome heterogeneity in CRISPR performance. Understanding these differences is particularly crucial when targeting social bacterial behaviors like quorum sensing, where pathway redundancy and strain-specific regulatory networks can confound standardized approaches. The efficacy of CRISPR systems is not uniform but is influenced by a complex interplay of genetic, structural, and delivery factors that vary substantially across microbial targets [69] [20].
CRISPR System Diversity: Mechanisms and Implications
CRISPR-Cas systems demonstrate remarkable diversity in nature, which directly impacts their suitability and efficacy for different applications. These systems are broadly classified into two classes based on their effector molecules, with distinct functional characteristics that influence their experimental performance [70] [20].
Table 1: Classification and Characteristics of Major CRISPR Systems
| Class | Type | Signature Nuclease | Target | Subunit Composition | Prevalence | Key Features |
|---|---|---|---|---|---|---|
| Class 1 | I | Cas3 | DNA | Multi-protein | ~90% of CRISPR loci in bacteria and archaea | Contains Cas3 with helicase and nuclease activity [70] |
| III | Cas10 | DNA/RNA | Multi-protein | Found in bacteria and archaea | Recognizes and cleaves both DNA and RNA [70] | |
| Class 2 | II | Cas9 | DNA | Single protein | ~10% of CRISPR loci (bacteria only) | Requires tracrRNA; most widely used in genome editing [70] [20] |
| V | Cas12 | DNA | Single protein | Found in bacteria | Requires tracrRNA; targets DNA [70] | |
| VI | Cas13 | RNA | Single protein | Found in bacteria | Only Class 2 system that targets RNA; enables RNA editing [70] |
The distribution of these systems across prokaryotes is uneven, with approximately 40% of sequenced bacteria and over 80% of archaea encoding at least one CRISPR-Cas system [20]. This natural variation constitutes the first layer of strain-to-strain differences researchers encounter. Class 1 systems, utilizing multi-protein effector complexes, represent the majority of naturally occurring CRISPR systems and are particularly prevalent in archaea. In contrast, Class 2 systems rely on a single effector protein (e.g., Cas9, Cas12, Cas13) and are found exclusively in bacteria [70] [20]. This fundamental architectural difference impacts both the delivery logistics and the functional efficacy when these systems are deployed against target bacteria.
Molecular Determinants of Variable Efficacy
Guide RNA (sgRNA) Efficiency and Sequence Specificity
The efficiency of CRISPR-Cas systems depends critically on the guide RNA sequence and its interaction with target genomic loci. Nucleotide composition significantly influences sgRNA efficacy, with specific patterns correlating with successful targeting. Research has identified a preference for guanine and avoidance of thymine near the Protospacer Adjacent Motif (PAM) sequence, cytosine preference near the cut site, and overall GC content as major determinants of efficiency [69]. The seed region—the 5-10 bases nearest the PAM sequence—is particularly critical for target recognition and cleavage efficiency [69].
Quantum chemical properties of the sgRNA-target DNA hybrid further contribute to efficacy variation. Advanced modeling reveals that electron density distributions, characterized by Highest Occupied Molecular Orbital-Lowest Unoccupied Molecular Orbital (HOMO-LUMO) gaps, influence the kinetic stability of the hybridization complex [69]. Additionally, hydrogen bonding and π-stacking interactions between aromatic rings in nucleotide bases affect thermodynamic stability and complex formation. These quantum properties vary with sequence composition and can predict sgRNA efficiency across different bacterial strains [69].
Cellular Context and Delivery Challenges
Bacterial cellular context introduces another layer of variability in CRISPR efficacy. Prokaryotic cells have simpler chromatin structure than eukaryotic cells, potentially making target regions more accessible [69]. However, differences in DNA repair mechanisms significantly impact CRISPR outcomes. Mammalian cells employ active non-homologous end-joining (NHEJ) systems, while in prokaryotes, double-strand breaks are often lethal in the absence of efficient repair systems [69]. This fundamental difference means sgRNA activity in bacteria correlates strongly with cellular survival, creating stronger selective pressure against efficient targeting.
Delivery efficiency varies substantially across bacterial strains due to differences in cell envelope composition, efflux pump activity, and innate immune mechanisms. Nanoparticle-based delivery systems have emerged as promising vehicles to overcome these barriers, with liposomal Cas9 formulations demonstrating over 90% reduction of Pseudomonas aeruginosa biofilm biomass in vitro, and gold nanoparticle carriers enhancing editing efficiency up to 3.5-fold compared to non-carrier systems [11]. The extracellular polymeric substance (EPS) matrix of biofilms presents a particular challenge, restricting penetration of CRISPR components and contributing to strain-dependent efficacy [11] [25].
Quantitative Analysis of Efficacy Variables
Table 2: Factors Influencing Strain-to-Strain and System-to-System Efficacy
| Factor Category | Specific Variable | Impact on Efficacy | Experimental Workaround |
|---|---|---|---|
| sgRNA Design | Seed region composition (5-10 bp near PAM) | Critical for target recognition and cleavage efficiency [69] | Favor guanine, avoid thymine near PAM; prefer cytosine near cut site |
| GC content | Moderate GC (40-60%) generally optimal; extremes reduce efficiency [69] | Adjust sgRNA design to maintain moderate GC content across target region | |
| HOMO-LUMO gap | Affects kinetic stability of sgRNA-DNA hybrid [69] | Quantum chemical modeling to predict stable hybridization | |
| Cellular Context | Chromatin accessibility | More accessible in prokaryotes; still varies by genomic location [69] | Choose target sites with minimal nucleosome occupancy or chromatin structure |
| DNA repair mechanisms | NHEJ absent in many bacteria; double-strand breaks often lethal [69] | Employ CRISPRi/a with dCas9 for non-lethal gene regulation | |
| Efflux pumps & membrane permeability | Varies by strain; impacts delivery efficiency [11] | Utilize nanoparticle carriers to enhance uptake and avoid efflux | |
| Delivery Method | Nanoparticle type | Liposomal: >90% biofilm reduction; Gold: 3.5× editing efficiency [11] | Match nanoparticle characteristics to target strain and biofilm status |
| Biofilm penetration | EPS matrix reduces component penetration [11] [25] | Engineer nanoparticles with biofilm-degrading enzymes or surface modifications |
Experimental Protocols for Assessing Efficacy
sgRNA Efficiency Profiling in Bacterial Strains
Purpose: To quantitatively measure and compare CRISPR sgRNA efficiency across multiple bacterial strains targeting homologous genes in quorum sensing pathways.
Materials:
- Bacterial strains of interest (e.g., Pseudomonas aeruginosa, Escherichia coli, Staphylococcus aureus)
- CRISPR-Cas9 plasmid system appropriate for target strains
- Custom sgRNA library targeting conserved regions of quorum sensing genes (e.g., lasI, rhlI, luxS)
- Electroporation or conjugation equipment for plasmid delivery
- Selection antibiotics appropriate for plasmid and bacterial strain
- Next-generation sequencing platform
- Quantum chemistry computational resources (optional)
Methodology:
- Design sgRNAs targeting homologous quorum sensing genes across test strains, noting position-specific sequence features and calculating quantum chemical properties [69].
- Introduce CRISPR-Cas9-sgRNA constructs into each bacterial strain via optimized transformation methods (electroporation, conjugation, or nanoparticle delivery) [11].
- Culture transformed bacteria under appropriate conditions with selection pressure for 24-72 hours.
- Extract genomic DNA and amplify target regions for sequencing.
- Quantify editing efficiency by calculating the percentage of sequencing reads with indels or modifications at target sites.
- Correlate editing efficiency with sgRNA sequence features, including nucleotide composition, GC content, and predicted quantum chemical properties [69].
- For advanced modeling, employ iterative Random Forest (iRF) machine learning to identify strain-specific determinants of sgRNA efficiency [69].
Assessing Biofilm Disruption Efficacy Across Strains
Purpose: To evaluate strain-dependent differences in CRISPR-mediated biofilm disruption targeting quorum sensing pathways.
Materials:
- Crystal violet staining solution
- Microtiter plate biofilm assay system
- Confocal laser scanning microscope (CLSM)
- RNA extraction and qRT-PCR equipment
- Nanoparticle delivery systems (liposomal, gold, or polymeric) [11]
Methodology:
- Engineer CRISPR-Cas systems targeting key quorum sensing genes (e.g., lasR, rhlR, luxR) in isogenic or clinical strains.
- Deliver CRISPR constructs using nanoparticle-based systems optimized for biofilm penetration [11].
- Quantify biofilm biomass using crystal violet staining at 24, 48, and 72-hour time points.
- Visualize biofilm architecture and thickness using CLSM with appropriate stains.
- Extract RNA from biofilm cells and quantify expression of target quorum sensing genes and downstream virulence factors using qRT-PCR.
- Compare efficacy across strains and correlate with strain characteristics (e.g., EPS composition, growth rate, natural competence).
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Addressing Efficacy Variation
| Reagent/Solution | Function | Application Context | Considerations for Strain Variability |
|---|---|---|---|
| CRISPR-Cas9 System (Type II) | DNA cleavage with sgRNA guidance | Broad-spectrum gene editing in transformable strains | Requires functional PAM (NGG); efficiency varies by strain [70] [20] |
| CRISPR-Cas12 System (Type V) | DNA cleavage with different PAM requirements | Alternative to Cas9 with different PAM preferences | TTTN PAM may work in strains where Cas9 fails [70] |
| dCas9 CRISPRi/a Systems | Gene repression/activation without cleavage | Essential gene study; metabolic engineering; quorum sensing modulation | Avoids lethal cleavage; better for comparative studies across strains [25] |
| Liposomal Nanoparticles | Encapsulation and delivery of CRISPR components | Enhanced biofilm penetration; P. aeruginosa studies | >90% biofilm biomass reduction demonstrated [11] |
| Gold Nanoparticles | Non-viral delivery vehicle for CRISPR components | Improving editing efficiency in recalcitrant strains | 3.5× enhancement in editing efficiency reported [11] |
| CRISPRviz/CrisprVi Software | Visualization and analysis of CRISPR sequences | Identifying native CRISPR systems in target strains | Helps design strain-specific guides; identifies natural immunity [71] |
| Iterative Random Forest (iRF) | Explainable AI for sgRNA efficiency prediction | Predicting strain-specific sgRNA performance | Identifies quantum chemical features influencing efficacy [69] |
Visualization and Computational Analysis Approaches
Advanced visualization and computational tools are essential for understanding and predicting efficacy variations across bacterial strains and CRISPR systems. CrisprVi software enables researchers to visually analyze CRISPR direct repeats and spacers, providing insights into the locations, orders, and components of CRISPR sequences [71]. This tool facilitates the identification of consensus sequences across strains, which can inform sgRNA design for broad-spectrum applications.
Machine learning approaches, particularly iterative Random Forest (iRF), enable researchers to identify complex patterns in sgRNA efficiency data [69]. By incorporating quantum chemical properties and sequence features into predictive models, researchers can develop strain-specific sgRNA design rules that account for the molecular interactions governing CRISPR efficacy. These computational approaches are particularly valuable for optimizing CRISPR interventions in quorum sensing pathways, where precision and efficiency are critical for disrupting social behaviors without imposing excessive selective pressure.
The variable efficacy of CRISPR systems across bacterial strains and system types presents both challenges and opportunities for researchers targeting quorum sensing and biofilm formation. By understanding the molecular determinants of this variability—from sgRNA sequence features and quantum chemical properties to delivery efficiency and cellular context—researchers can develop more predictable and robust CRISPR interventions. The integration of nanoparticle delivery systems, advanced computational modeling, and strain-specific optimization strategies will enhance our ability to harness CRISPR technologies for precise manipulation of bacterial social behaviors. As these approaches mature, they will accelerate both fundamental research into quorum sensing mechanisms and the development of novel therapeutic strategies targeting biofilm-associated infections.
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) proteins constitute an adaptive immune system in prokaryotes, providing sequence-specific defense against invasive genetic elements like bacteriophages (phages) and plasmids [72] [15]. This defense system incorporates short sequences from invaders into the host genome, creating a molecular memory that guides the cleavage of future infections. In response, phages have evolved sophisticated countermeasures, primarily Anti-CRISPR (Acr) proteins, which are small, potent inhibitors that inactivate the CRISPR-Cas machinery through diverse mechanisms [73]. This dynamic arms race drives the molecular evolution of both bacterial defense and phage offense, with significant implications for bacterial physiology, particularly in the contexts of quorum sensing (QS) and biofilm formation [74] [15]. Understanding these escape mechanisms is critical for developing novel antimicrobial strategies and leveraging CRISPR technologies against antibiotic-resistant infections.
Core Bacterial Escape Mechanisms
Anti-CRISPR Proteins: Structure and Direct Inhibition
Anti-CRISPR proteins are the primary mechanism by which phages directly counteract CRISPR-Cas immunity. They are highly diverse and often specific to particular types and subtypes of CRISPR-Cas systems. Their mechanisms of action include:
- Direct Blockage of DNA Binding: AcrIE3, identified in a Pseudomonas phage, inhibits the Type I-E CRISPR-Cas system in Pseudomonas aeruginosa. Structural and biochemical analyses reveal that AcrIE3 employs an all-helical fold with a negatively charged surface to selectively bind the Cas8e subunit of the Cascade complex. This binding site is the protospacer adjacent motif (PAM) recognition pocket, effectively preventing the Cascade complex from recognizing and binding to its target DNA [73].
- Diversity and Specificity: The discovery of AcrIE3 highlights that PAM interaction sites are primary targets for divergent Acr inhibitors. The precise steric and charge complementarity allows these small proteins to efficiently jam the recognition machinery of large, multi-protein Cas complexes [73].
Evolutionary Genomic Escape Strategies
Beyond direct protein inhibition, phages evolve genomic characteristics that reduce their susceptibility to CRISPR recognition and cleavage over evolutionary time.
- PAM Sequence Depletion: A comprehensive analysis of P. aeruginosa and its phages demonstrated that viral genomes targeted by the predominant Type I-F CRISPR-Cas system (which recognizes a 5′-CC PAM) have a significantly lower frequency of CC dinucleotides than non-targeted viruses. This indicates a strong selective pressure to mutate PAM sequences, thereby rendering the phage "invisible" to the bacterial immune system [72].
- Genomic Nucleotide Composition Bias: The same study found that phages targeted by the Type I-F system also possess a lower overall G+C content. This genome-wide mutational bias contributes to the systematic depletion of CC motifs, representing a long-term, passive escape strategy [72].
Table 1: Primary Bacterial Escape Mechanisms from CRISPR-Cas Immunity
| Escape Mechanism | Molecular Basis | Example | Outcome |
|---|---|---|---|
| Anti-CRISPR (Acr) Proteins | Direct binding and inhibition of Cas proteins or complexes. | AcrIE3 binding to Cas8e in Type I-E system [73]. | Immediate, potent shutdown of immune function. |
| PAM Sequence Depletion | Mutation of protospacer adjacent motifs (PAMs) in the phage genome. | Lower CC dinucleotide frequency in phages targeted by P. aeruginosa I-F system [72]. | Phage evades initial recognition by the Cascade complex. |
| Genomic GC Content Reduction | Genome-wide mutation pressure leading to fewer PAM sequences. | Lower G+C content in targeted phages [72]. | Long-term evolutionary reduction in targetability. |
The Interplay with Quorum Sensing and Biofilm Formation
The function of CRISPR-Cas systems is not isolated but is integrated into the broader regulatory networks of the bacterial cell, notably quorum sensing (QS)—a cell-cell communication process that controls collective behaviors in a cell-density-dependent manner.
Quorum Sensing Regulation of CRISPR-Cas
Research in P. aeruginosa has established a direct link between QS and CRISPR-Cas activity. The bacterium uses QS to upregulate the expression of cas genes at high cell density [74]. This regulatory logic ensures that the energy-intensive CRISPR-Cas immune system is maximally active precisely when large, dense bacterial populations are at the highest risk for phage infection. Consequently, QS activation enhances both CRISPR-Cas targeting of foreign DNA and the acquisition of new spacers (adaptation) [74]. This relationship also presents a therapeutic avenue; pro- and anti-quorum sensing compounds can be used to modulate CRISPR-Cas activity, potentially suppressing it to enhance phage therapy efficacy [74].
CRISPR-Cas in Biofilm Development and Virulence
The role of CRISPR-Cas systems in biofilm formation and virulence is complex and appears to be strain and system-dependent, indicating a sophisticated, context-specific regulatory function beyond immunity.
- Biofilm Inhibition: In P. aeruginosa, the Type I-F system is implicated in the loss of biofilm formation upon lysogenic infection by phage DMS3. This CRISPR-dependent inhibition requires the presence of a specific PAM within the bacterial genome, suggesting a mechanism of endogenous gene regulation that alters bacterial physiology and social behavior [15].
- Biofilm Enhancement: Conversely, in Acinetobacter baumannii, the Cas3 protein of the Type I-Fa system acts as a positive regulator of biofilm formation and virulence. Deletion of cas3 led to significantly reduced biofilm biomass, thickness, and downregulation of key virulence factors like the outer membrane protein A (OmpA). In mouse infection models, the Δcas3 strain showed markedly reduced pathogenicity [14].
- Dual Regulatory Roles: Another study in A. baumannii on the I-Fb system showed that the transcriptional regulators H-NS and BaeR suppress cas3 expression. Deletion of cas3 in this context resulted in increased biofilm formation, elevated production of extracellular matrix (PNAG), and enhanced epithelial cell adhesion. This indicates that in this specific subtype, the CRISPR-Cas system represses virulence traits, and its suppression leads to a hypervirulent phenotype [17].
Table 2: Contrasting Roles of CRISPR-Cas in Bacterial Physiology and Virulence
| Bacterial Species & System | Effect on Biofilm/Virulence | Proposed Mechanism |
|---|---|---|
| Pseudomonas aeruginosa (Type I-F) | Inhibits biofilm formation [15]. | Self-targeting of endogenous genes via CRISPR interference. |
| Acinetobacter baumannii (Type I-Fa) | Enhances biofilm formation and virulence; Δcas3 is attenuated [14]. | Regulation of virulence factors (e.g., OmpA) and metabolic pathways (e.g., carbon metabolism). |
| Acinetobacter baumannii (Type I-Fb) | Suppresses biofilm and virulence; Δcas3 shows enhanced adhesion [17]. | BaeR and H-NS regulators suppress Cas3, which normally inhibits biofilm/EPS and pilus expression. |
Experimental Analysis of Anti-CRISPR Mechanisms
Structural Analysis of AcrIE3
Objective: To determine the high-resolution structure of the Anti-CRISPR protein AcrIE3 and elucidate its mechanism of inhibition against the Type I-E Cascade complex.
Protocol:
- Protein Expression and Purification: Clone the gene encoding AcrIE3 into an expression vector (e.g., pET series). Transform into E. coli BL21(DE3) and induce protein expression with IPTG. Purify the protein using affinity chromatography (e.g., Ni-NTA for His-tagged protein), followed by size-exclusion chromatography.
- Crystallization and Data Collection: Set up crystallization trials for purified AcrIE3 using commercial sparse matrix screens. Optimize initial hits. Flash-cool the crystal in liquid nitrogen and collect X-ray diffraction data at a synchrotron facility.
- Structure Determination: Solve the phase problem using molecular replacement or single-wavelength anomalous dispersion (SAD). Build and refine the atomic model of AcrIE3.
- Mutational Analysis: Conduct site-directed mutagenesis of surface-exposed, negatively charged residues on AcrIE3. Test the binding affinity of these mutants for the Cascade complex using techniques like Surface Plasmon Resonance (SPR) or Isothermal Titration Calorimetry (ITC).
- Biochemical Validation: Perform Electrophoretic Mobility Shift Assays (EMSAs) to demonstrate that wild-type AcrIE3, but not the binding-deficient mutants, inhibits the binding of the Cascade complex to its target DNA substrate [73].
Genomic Analysis of PAM Evasion
Objective: To statistically analyze the frequency of PAM sequences in phage genomes that are targeted versus not targeted by a specific bacterial CRISPR-Cas system.
Protocol:
- Data Collection: Retrieve a large number of bacterial genomes (e.g., 7,876 P. aeruginosa genomes) from public databases like NCBI. Identify and classify CRISPR-Cas systems and their spacers using bioinformatics tools like CRISPRCasFinder and CRISPRCasTyper [72].
- Spacer-PROTOSPACER Matching: Align spacer sequences from the bacterial CRISPR arrays against a database of viral genomes using BLASTn to identify "targeted" phages (those with matching protospacers).
- PAM Prediction and Frequency Analysis: For each CRISPR-Cas subtype (e.g., I-F), predict the PAM consensus sequence (e.g., 5′-CC) using a tool like Spacer2PAM. For the viral genomes, calculate the observed frequency of the PAM dinucleotide. Compare this frequency to the expected frequency based on the overall nucleotide composition of the viral genome.
- Statistical Testing: Apply statistical tests (e.g., Chi-squared test) to determine if the depletion of PAM sequences in targeted phages is significant compared to non-targeted phages [72].
The Scientist's Toolkit: Key Research Reagents and Solutions
Table 3: Essential Reagents for Investigating CRISPR Escape Mechanisms
| Research Reagent / Tool | Function & Application | Key Characteristics |
|---|---|---|
| Heterologous Expression Systems (e.g., E. coli BL21(DE3) with pET vectors) | High-yield expression and purification of recombinant Cas and Acr proteins for structural and biochemical studies [73] [17]. | Allows for production of tagged proteins (e.g., His-tag) for simplified purification. |
| Surface Plasmon Resonance (SPR) | Label-free analysis of binding kinetics (e.g., affinity, on/off rates) between Acr proteins and Cas protein targets [73]. | Provides quantitative data on protein-protein interactions. |
| Electrophoretic Mobility Shift Assay (EMSA) | Determines if a protein (e.g., Acr) can bind to a protein-DNA complex (e.g., Cascade-target DNA) and inhibit complex formation [73] [17]. | A classic, accessible gel-based technique. |
| Bioinformatics Suites (e.g., CRISPRCasFinder, Spacer2PAM) | Identifies CRISPR arrays and cas genes in genomic data and predicts PAM sequences from spacer matches [72]. | Essential for large-scale genomic analysis and hypothesis generation. |
| Confocal Laser Scanning Microscopy (CLSM) | Visualizes 3D architecture, biomass, and thickness of bacterial biofilms, often using fluorescent stains (e.g., SYTO9, dextran conjugates) [14]. | Critical for assessing phenotypic consequences of CRISPR/Acr manipulation on biofilms. |
| Quorum Sensing Modulators (synthetic AIs, QS inhibitors) | Pro- and anti-quorum sensing compounds used to investigate the regulatory link between QS and CRISPR-Cas activity [74]. | Allows functional dissection of cell-density-dependent immune regulation. |
The evolutionary arms race between bacteria and phages has given rise to sophisticated escape mechanisms like Anti-CRISPR proteins and genomic PAM depletion. These are not merely isolated defensive and offensive tactics but are deeply intertwined with the core physiology of the bacterium, including its social communication (quorum sensing) and community structure (biofilm formation). The dual role of CRISPR-Cas systems—acting as both a immune defense and a regulator of virulence—adds a layer of complexity that must be considered when developing CRISPR-based antimicrobials or phage therapies. Future research must focus on discovering novel Acrs, unraveling the precise molecular pathways by which CRISPR-Cas systems regulate gene expression, and exploiting the QS-CRISPR link to manipulate bacterial behavior. Understanding these multifaceted escape mechanisms will be pivotal in advancing our ability to combat persistent biofilm-driven infections and manage the growing crisis of antibiotic resistance.
Bench to Bedside: Validating CRISPR Strategies and Comparative Analysis with Alternative Therapies
The escalating crisis of antibiotic-resistant bacterial infections poses a formidable challenge to global health, with biofilms playing a pivotal role in bacterial persistence and treatment failure [11]. These structured microbial communities, encased within a self-produced extracellular polymeric substance (EPS), exhibit dramatically enhanced tolerance to antimicrobial agents—often up to 1000-fold greater than their planktonic counterparts [11]. Within the context of infectious disease management, accurate assessment of therapeutic outcomes against biofilms is paramount for developing effective treatment strategies.
The emergence of CRISPR-based technologies has revolutionized antibacterial research, providing unprecedented precision in targeting bacterial virulence mechanisms [75]. These systems enable selective disruption of antibiotic resistance genes, quorum-sensing pathways, and biofilm-regulating factors [11]. However, the accurate evaluation of these innovative therapies demands robust, standardized metrics for quantifying biofilm biomass and bacterial viability reduction. This technical guide provides researchers and drug development professionals with comprehensive methodologies for assessing anti-biofilm therapeutic efficacy, with particular emphasis on CRISPR-mediated interventions. We present detailed protocols, analytical frameworks, and standardized metrics essential for validating novel anti-biofilm strategies in the era of precision antimicrobials.
Core Metrics and Quantitative Assessment
The evaluation of anti-biofilm therapeutic efficacy rests on two fundamental pillars: the quantification of biofilm structural integrity (biomass) and the measurement of microbial survival (viability). A comprehensive assessment typically integrates both structural and viability metrics to provide a complete picture of therapeutic impact [76].
Table 1: Core Metrics for Biofilm Biomass Quantification
| Metric | Methodology | Key Output Parameters | Advantages | Limitations |
|---|---|---|---|---|
| Crystal Violet Staining | Colorimetric assay measuring bound dye after biofilm fixation | Optical Density (OD~595nm~), Percentage biomass reduction | High-throughput, cost-effective, well-established | Does not differentiate live/dead cells; measures total adhered biomass only [76] |
| Confocal Laser Scanning Microscopy (CLSM) | Fluorescent staining and 3D image reconstruction | Biofilm thickness (µm), Biovolume (µm³), Surface coverage (%) | Provides 3D structural data, can be combined with viability stains | Requires specialized equipment, lower throughput, complex data analysis [77] [14] |
| Scanning Electron Microscopy (SEM) | High-resolution imaging of dehydrated samples | Ultrastructural morphology, matrix architecture, cell arrangement | Exceptional spatial resolution, reveals biofilm topography | Requires extensive sample preparation, artifacts possible, no viability data [76] |
| Dual-Staining Method (Maneval's) | Sequential Congo red and Maneval's staining for light microscopy | Matrix thickness, cell-matrix differentiation, structural integrity | Cost-effective, distinguishes cells (magenta-red) from matrix (blue) | Semi-quantitative, requires manual interpretation [76] |
Table 2: Core Metrics for Bacterial Viability Assessment
| Metric | Methodology | Key Output Parameters | Advantages | Limitations |
|---|---|---|---|---|
| Colony Forming Units (CFUs) | Serial dilution and plating of dispersed biofilms | CFU/mL, Log~10~ reduction, Percentage viability loss | Direct measure of cultivable cells, considered gold standard | Labor-intensive, misses viable but non-culturable cells, biofilm dispersal critical [14] |
| Flow Cytometry with Vital Stains | Fluorogenic dye analysis of single-cell suspensions | Percentage live/dead cells, fluorescence intensity distributions | Rapid, quantitative, high-throughput capability | Requires biofilm disaggregation, potential staining artifacts [77] |
| Metabolic Assays (Resazurin/XTT) | Measurement of metabolic activity via dye reduction | Fluorescence/absorbance units, percentage metabolic inhibition | High sensitivity, can monitor real-time changes | Does not directly correlate with cell number, affected by metabolic state [11] |
| CLSM with Viability Stains | Dual staining with SYTO9 (live) and propidium iodide (dead) | Live/dead cell ratios, spatial distribution of viability | Provides spatial context of viability within biofilm structure | Photo-bleaching, dye penetration issues in thick biofilms [14] |
Advanced Methodologies for CRISPR Anti-Biofilm Research
CRISPR-Mediated Biofilm Disruption: Experimental Workflow
The evaluation of CRISPR-based anti-biofilm therapeutics requires specialized methodologies that account for their unique mechanism of action, which involves genetic disruption of biofilm-related pathways rather than direct bactericidal activity [11]. The workflow typically progresses from genetic construction to phenotypic assessment.
Detailed Protocol: Assessing CRISPR-Nanoparticle Efficacy
Objective: Evaluate the efficacy of CRISPR-nanoparticle conjugates against mature Pseudomonas aeruginosa biofilms using integrated biomass and viability metrics [11].
Materials and Reagents:
- 24-well polystyrene tissue culture plates
- Mueller Hinton broth or appropriate bacterial growth medium
- CRISPR-loaded nanoparticles (e.g., liposomal Cas9-gRNA or gold nanoparticle hybrids)
- Phosphate Buffered Saline (PBS), pH 7.4
- Crystal violet solution (0.1% w/v)
- Acetic acid (30% v/v)
- SYTO9 and propidium iodide fluorescent stains
- DNAse I for biofilm dispersion
- Tryptic Soy Agar plates
Procedure:
Week 1: Biofilm Establishment
- Inoculum Preparation: Grow P. aeruginosa overnight in suitable broth. Dilute to 1×10^6 CFU/mL in fresh medium.
- Biofilm Formation: Add 1 mL bacterial suspension per well of 24-well plate. Incubate statically for 24-48 hours at 37°C to establish mature biofilms.
- Washing: Gently wash established biofilms twice with PBS to remove non-adherent cells.
Week 1: Therapeutic Intervention
- Treatment Application: Add CRISPR-nanoparticle formulations in fresh medium at predetermined concentrations. Include appropriate controls:
- Untreated biofilm control
- Nanoparticle-only control
- Conventional antibiotic control
- Free CRISPR system (without nanoparticles)
- Incubation: Incubate treated biofilms for 4-24 hours at 37°C, depending on experimental design.
Week 1: Post-Treatment Analysis
- Biofilm Biomass Assessment (Crystal Violet):
- Remove medium and gently wash wells twice with PBS
- Fix biofilms with methanol for 15 minutes
- Stain with 0.1% crystal violet for 20 minutes
- Wash extensively to remove unbound dye
- Solubilize bound dye in 30% acetic acid
- Measure OD~595nm~ using plate reader
- Calculate percentage biomass reduction relative to untreated control
- Bacterial Viability Assessment (CFU Enumeration):
- In parallel wells, remove treatment and wash gently with PBS
- Add 1 mL PBS containing DNAse I (10 µg/mL) and sonicate briefly (5-10 seconds at low power) to disperse biofilm
- Serially dilute dispersed biofilm suspension in PBS
- Plate dilutions on appropriate agar medium
- Incubate plates at 37°C for 24-48 hours
- Count CFUs and calculate log~10~ reduction compared to untreated control
Advanced Assessment: CLSM Analysis
- Biofilm Imaging:
- Grow biofilms on sterile coverslips placed in 24-well plates
- Apply treatments as described above
- Stain with SYTO9 (5 µM) and propidium iodide (15 µM) for 30 minutes in dark
- Image using CLSM with appropriate filter sets
- Acquire Z-stacks at 1-2 µm intervals through entire biofilm thickness
- Analyze using image analysis software (e.g., ImageJ, IMARIS, COMSTAT) to determine:
- Total biovolume (µm³/µm²)
- Average thickness (µm)
- Live/dead cell ratios throughout depth
- Surface coverage (%)
Expected Outcomes: Effective CRISPR-nanoparticle treatments targeting biofilm-related genes (e.g., quorum-sensing regulators) typically demonstrate 70-90% biomass reduction and 2-4 log~10~ reductions in viability [11]. The spatial analysis should reveal structural collapse and increased dead cell regions, particularly in deeper biofilm layers where nanoparticle penetration is often limited.
CRISPRi for Functional Gene Analysis in Biofilms
CRISPR interference (CRISPRi) provides a powerful approach for validating potential anti-biofilm targets by enabling precise gene knockdown without permanent mutation [78]. This methodology is particularly valuable for identifying essential genes involved in biofilm formation and maintenance.
Protocol Overview:
- CRISPRi System Design: Utilize dCas9 and target-specific gRNAs against genes of interest (e.g., gacA, alg44, or c-di-GMP regulators) [78].
- Transformation: Introduce CRISPRi plasmids into target bacterial strains using appropriate methods.
- Biofilm Phenotyping: Assess knockdown effects using microtiter plate assays and CLSM as described above.
- Molecular Validation: Confirm target gene knockdown using RT-qPCR and assess downstream effects on relevant pathways.
Key Parameters: Effective CRISPRi-mediated gene silencing should produce dose-dependent reductions in target gene expression (typically 60-90% knockdown) with corresponding phenotypic changes in biofilm architecture and viability [78].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Research Reagent Solutions for CRISPR-Biofilm Studies
| Reagent Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| CRISPR Delivery Systems | Liposomal nanoparticles, Gold nanoparticles, Bacteriophages [11] | Deliver CRISPR components to bacterial cells within biofilms | Efficiency of biofilm penetration; stability; loading capacity |
| Fluorescent Stains | SYTO9, Propidium iodide, CellTrace dyes, Calcofluor white [77] [76] | Differentiate live/dead cells; label specific biofilm components | Photostability; penetration depth; spectral compatibility |
| Biofilm Matrix Dyes | Congo red, Alexa Fluor-conjugated dextran, FITC-labeled lectins [76] [14] | Visualize and quantify extracellular polymeric substances | Specificity for matrix components; background staining |
| Gene Expression Tools | dCas9 CRISPRi systems, Reporter constructs [78] | Modulate and monitor gene expression in biofilm contexts | Knockdown efficiency; temporal control; promoter compatibility |
| Biofilm Dispersal Agents | DNAse I, Dispersin B, Proteases [11] | Facilitate biofilm breakdown for analysis | Activity against specific matrix components; effect on viability |
Signaling Pathways in CRISPR-Biofilm Research
Understanding the molecular pathways targeted by CRISPR-based interventions is essential for interpreting experimental outcomes and designing effective therapeutic strategies.
Data Interpretation and Reporting Standards
Normalization and Statistical Considerations
Accurate interpretation of anti-biofilm efficacy data requires appropriate normalization and statistical analysis:
- Baseline Normalization: Express biomass and viability data as percentages relative to untreated controls
- Positive Controls: Include appropriate benchmark compounds (conventional antibiotics, matrix-disrupting agents)
- Time-Course Analyses: Monitor therapeutic effects at multiple time points (4, 8, 24, 48 hours) to capture dynamics
- Spatial Considerations: Account for spatial heterogeneity in biofilms through multiple sampling regions or imaging fields
- Dose-Response Relationships: Establish concentration-dependent effects for CRISPR therapeutics
Minimum Reporting Standards
For publication and comparative analysis, include these essential parameters:
- Biofilm Growth Conditions: Medium composition, incubation time, temperature, surface material
- Biofilm Maturity: Age of biofilms at treatment initiation
- Treatment Exposure: Duration, concentration, volume
- Control Data: Untreated, vehicle-only, and appropriate positive controls
- Replication: Minimum of three biological replicates with technical triplicates
- Normalization Method: Detailed description of normalization approach
- Statistical Analysis: Specific tests used with significance thresholds
The rigorous assessment of biofilm biomass and bacterial viability represents a critical component in the development of CRISPR-based anti-biofilm therapies. The methodologies outlined in this technical guide provide a standardized framework for evaluating therapeutic efficacy across multiple dimensions—from structural disruption to functional consequences. As CRISPR technologies continue to evolve toward clinical application, these metrics will play an increasingly vital role in validating intervention strategies, optimizing delivery systems, and ultimately addressing the pervasive challenge of biofilm-associated infections. The integration of quantitative biomass assessment with sophisticated viability metrics enables comprehensive characterization of anti-biofilm agents, accelerating the translation of CRISPR-based therapeutics from laboratory research to clinical implementation.
Abstract Biofilm-associated infections represent a significant challenge in clinical settings due to their inherent resistance to antimicrobial treatments. This whitepaper provides a comparative analysis of conventional antibiotic strategies and the emerging paradigm of CRISPR-based gene editing for combating biofilm-mediated resistance. Conventional approaches, while enhanced by adjunctive therapies, primarily target the structural integrity of the biofilm and actively growing cells. In contrast, CRISPR/Cas9 technology offers a precision tool to disrupt the genetic foundations of biofilm formation, antibiotic resistance, and bacterial persistence. Framed within the context of quorum sensing and biofilm research, this analysis synthesizes current efficacy data, detailed methodologies, and essential research tools, highlighting the potential of integrated therapeutic strategies to overcome one of modern medicine's most persistent obstacles.
Bacterial biofilms are structured communities of microbial cells enclosed in a self-produced matrix of extracellular polymeric substances (EPS) that adhere to biological or inert surfaces [11] [47]. This EPS matrix, composed of polysaccharides, proteins, and extracellular DNA (eDNA), creates a formidable biological barrier that severely limits the efficacy of conventional antimicrobial agents [75]. Biofilm-forming bacteria can exhibit up to 1000-fold greater tolerance to antibiotics compared to their free-floating (planktonic) counterparts, leading to persistent and chronic infections that are notoriously difficult to eradicate [11]. This resistance is multifactorial, arising from physical barrier-mediated limited antibiotic penetration, reduced metabolic activity of embedded cells, and the presence of persistent "dormant" phenotypes [75] [47].
The global health impact of biofilm-associated infections is profound, complicating treatments involving medical implants, causing chronic wounds, and perpetuating lung infections in cystic fibrosis patients. The ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species) are of particular concern, as they frequently form robust biofilms on medical devices and tissues [47]. The escalating crisis of antimicrobial resistance (AMR) has therefore prompted a critical re-evaluation of conventional antibiotic monotherapies and spurred the development of novel, targeted approaches, with CRISPR-based genetic interventions standing at the forefront of this innovation [11] [75] [79].
Conventional Antibiotics: Mechanisms, Limitations, and Adjunctive Strategies
Conventional antibiotics target essential bacterial cellular processes, but their efficacy is dramatically impaired against biofilms.
Table 1: Conventional Antibiotic Classes and Their Efficacy Against Biofilms
| Antibiotic Class | Primary Mechanism | Efficacy Against Planktonic Cells | Key Limitations Against Biofilms |
|---|---|---|---|
| β-lactams (e.g., Penicillin) | Inhibits cell wall synthesis | High | Ineffective against dormant cells; degraded by β-lactamases concentrated in biofilm matrix [11] [80] |
| Aminoglycosides (e.g., Gentamicin) | Inhibits protein synthesis | High | Reduced penetration through anionic EPS; requires aerobic metabolism for uptake [11] |
| Fluoroquinolones (e.g., Ciprofloxacin) | Inhibits DNA replication | High | Reduced efficacy against slow-growing or non-growing persister cells within biofilms [11] [81] |
| Macrolides (e.g., Erythromycin) | Inhibits protein synthesis | Moderate to High | Requires active growth for optimal efficacy; limited activity in biofilm depths [80] |
Enhancing Conventional Therapies: Synergistic and Anti-Biofilm Approaches
Research has focused on strategies to potentiate existing antibiotics. These include:
- Antibiotic Potentiators: Compounds like octyl gallate (OG), a food-grade antioxidant, can enhance antibiotic activity. OG increases bacterial cell wall permeability, improving penicillin and bacitracin's access to cellular targets, resulting in an 8-fold and 4-fold reduction in their minimum inhibitory concentration (MIC), respectively, against Staphylococcus epidermidis biofilms [80].
- Natural Biofilm Disruptors: Molecules such as Raspberry Ketone (RK) inhibit Salmonella enterica biofilm formation by disrupting the rdar morphotype (associated with curli fimbriae and cellulose) and impairing bacterial motility, without affecting planktonic cell growth [80].
- Antimicrobial Proteins and Peptides (AMPPs): Semi-purified hemolymph protein extract (HPE) from the Sydney Rock Oyster demonstrates synergistic activity with antibiotics like ampicillin and ciprofloxacin, improving their effectiveness by 2 to 32-fold against various pathogens, including Streptococcus pneumoniae and Pseudomonas aeruginosa [81].
CRISPR/Cas9: A Precision Genetic Arsenal Against Biofilms
The CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas9 system has emerged as a revolutionary tool for precision genome editing. Its application extends to targeted disruption of bacterial virulence, resistance mechanisms, and biofilm integrity [11] [75] [79].
Mechanisms of CRISPR Action Against Biofilms
CRISPR/Cas9 can be programmed to target specific genetic sequences crucial for biofilm resilience:
- Targeting Antibiotic Resistance Genes: The system can be directed to introduce double-strand breaks in chromosomal genes like bla (β-lactamase) or mecA (methicillin resistance), resensitizing bacteria to conventional antibiotics [11] [75].
- Disrupting Quorum Sensing (QS) Pathways: QS is a cell-density-dependent communication system that regulates biofilm formation and virulence. CRISPR/Cas9 can knockout key QS regulator genes (e.g., lasI, rhlI in P. aeruginosa or AbaI in A. baumannii), effectively "deafening" the bacteria and preventing coordinated biofilm development [11] [38].
- Interfering with Biofilm Structural Genes: Genes responsible for producing EPS components (e.g., alginate, cellulose, curli) can be disrupted, compromising the biofilm's structural integrity and making it more susceptible to antimicrobials and immune clearance [11] [75].
Table 2: Quantitative Efficacy of CRISPR-Based and Nanoparticle-Enhanced Strategies
| Therapeutic Strategy | Target Pathogen | Key Experimental Findings | Proposed Mechanism of Action |
|---|---|---|---|
| Liposomal CRISPR-Cas9 | Pseudomonas aeruginosa | >90% reduction in biofilm biomass in vitro [11] | Targeted disruption of resistance or QS genes within biofilm-embedded cells |
| CRISPR-Gold Nanoparticle Hybrids | Model bacterial pathogens | 3.5-fold increase in gene-editing efficiency compared to non-carrier systems [11] | Enhanced cellular uptake and controlled release of CRISPR components |
| β-caryophyllene Gold Nanoparticles (β-c-AuNPs) | S. aureus and C. albicans (mixed biofilm) | MIC of 512 µg/mL; significant reduction in mature biofilm CFUs [80] | Physical disruption of biofilm matrix and microbial cell membranes |
Direct Comparative Analysis: Mechanisms and Outcomes
The fundamental difference between the two approaches lies in their strategy: conventional antibiotics primarily apply a selective pressure that often fails within the biofilm niche, whereas CRISPR/Cas9 seeks to rewrite the genetic code that enables biofilm survival.
Table 3: Comparative Analysis of Conventional Antibiotics vs. CRISPR-Based Therapy
| Feature | Conventional Antibiotics | CRISPR/Cas9 Therapy |
|---|---|---|
| Primary Target | Vital cellular processes (e.g., cell wall, protein synthesis) | Specific genetic sequences (e.g., resistance, QS, virulence genes) |
| Specificity | Low to moderate (can affect commensal flora) | Very high (defined by guide RNA sequence) |
| Efficacy Against Dormant Cells | Low | Potentially high (targets DNA regardless of metabolic state) |
| Risk of Resistance Development | High (driven by selection pressure) | Potentially lower, but off-target effects must be minimized [11] |
| Delivery Challenge | Low (systemic or topical delivery) | High (requires efficient delivery into bacterial cells within biofilm) [11] [75] |
| Therapeutic Strategy | Broad-spectrum inhibition | Precision genetic engineering |
Experimental Protocols for Key Analyses
To evaluate anti-biofilm strategies, standardized and reproducible experimental models are essential.
Protocol: Assessing Synergy Between Adjuvants and Antibiotics
This protocol is adapted from studies on octyl gallate (OG) and oyster hemolymph protein extract (HPE) [80] [81].
- Checkerboard Titration Assay:
- Prepare serial two-fold dilutions of the antibiotic (e.g., penicillin) in a 96-well microtiter plate along one axis.
- Prepare serial two-fold dilutions of the potentiator (e.g., OG or HPE) along the other axis.
- Inoculate each well with a standardized suspension of the target bacterium (e.g., S. epidermidis at ~10^5 CFU/mL).
- Incubate the plate at 37°C for 18-24 hours.
- Determine the Minimum Inhibitory Concentration (MIC) of each agent alone and in combination.
- Data Analysis:
- Calculate the Fractional Inhibitory Concentration (FIC) index: FIC = (MIC of antibiotic in combination / MIC of antibiotic alone) + (MIC of potentiator in combination / MIC of potentiator alone).
- Interpret the FIC index: ≤0.5 indicates synergy; >0.5 to ≤4 indicates indifference; >4 indicates antagonism.
- Biofilm Inhibition Assay:
- Treat pre-formed biofilms in microtiter plates with sub-MIC concentrations of the synergistic combinations.
- Stain with crystal violet to quantify total biomass or use metabolic assays (e.g., resazurin) to assess cell viability.
Protocol: Evaluating CRISPR-Based Biofilm Disruption
This protocol is based on research utilizing nanoparticle-delivered CRISPR systems [11].
- sgRNA Design and Complex Formation:
- Design sgRNAs to target specific genes of interest (e.g., lasI in P. aeruginosa for QS disruption). Computational tools like CHOPCHOP are used for this purpose [38].
- Formulate CRISPR-Cas9 ribonucleoproteins (RNPs) or encode them in plasmids.
- Complex the CRISPR machinery with a delivery nanoparticle, such as gold nanoparticles (AuNPs) or lipid-based nanoparticles.
- Treatment of Biofilms:
- Grow mature biofilms (for 24-48 hours) on a relevant surface (e.g., peg lids, glass coverslips).
- Treat the biofilms with the CRISPR-nanoparticle formulation. Include controls: untreated biofilm, nanoparticle-only, and scrambled sgRNA.
- Incubate for a specified period (e.g., 4-24 hours) to allow for bacterial uptake and gene editing.
- Efficacy Assessment:
- Biomass Quantification: Use crystal violet staining to measure total biofilm reduction.
- Viability Assessment: Perform colony-forming unit (CFU) counts after disrupting the biofilm by sonication and vortexing.
- Gene Editing Confirmation: Extract genomic DNA from treated biofilms and use sequencing to confirm indels or gene knockout at the target locus.
- Functional Analysis: For QS targets, quantify the reduction in virulence factor production (e.g., pyocyanin, elastase) in the biofilm supernatant.
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for Anti-Biofilm Research
| Reagent / Material | Function in Research | Example Application |
|---|---|---|
| Octyl Gallate (OG) | Antibiotic potentiator that increases cell wall permeability. | Synergy studies with β-lactam antibiotics against staphylococcal biofilms [80]. |
| Raspberry Ketone (RK) | Natural compound that inhibits biofilm matrix formation. | Studying curli and cellulose inhibition in Salmonella biofilms [80]. |
| Gold Nanoparticles (AuNPs) | Carrier for CRISPR component delivery; intrinsic antimicrobial properties. | Enhancing cellular uptake of CRISPR-Cas9 RNPs for targeted gene editing in biofilms [11] [80]. |
| Hemolymph Protein Extract (HPE) | Natural source of antimicrobial proteins/peptides (AMPPs). | Investigating synergy with conventional antibiotics against respiratory pathogens [81]. |
| Benzalkonium Chloride (BAC) | Quaternary ammonium compound and disinfectant. | Evaluating biofilm inhibition and eradication on abiotic surfaces (e.g., against Campylobacter) [80]. |
| Quorum Sensing Inhibitors (e.g., Baicalein) | Chemical inhibitors of bacterial communication. | Probing the link between QS, CRISPR immunity, and phage therapy [28]. |
Visualizing Concepts and Workflows
Diagram 1: Comparative strategic workflow for conventional and CRISPR-based anti-biofilm approaches.
Diagram 2: Key stages of bacterial biofilm development, from initial attachment to dispersion [47].
The comparative analysis unequivocally demonstrates that conventional antibiotics and CRISPR/Cas9 gene editing offer distinct and potentially complementary approaches to the biofilm problem. Conventional therapies, though hampered by penetration and efficacy issues, remain a cornerstone of treatment, especially when enhanced by synergistic adjuvants. Meanwhile, CRISPR/Cas9 technology represents a paradigm shift, moving from broad-spectrum cytotoxicity to precision genetic disruption of the very mechanisms that underpin biofilm resilience and antibiotic resistance.
The future of anti-biofilm therapy likely lies in integrated, combinatorial strategies. A promising approach involves using CRISPR to resensitize a bacterial population to a conventional antibiotic, which is then administered to clear the infection. The significant challenge of efficient, targeted delivery of CRISPR components in complex clinical environments is being actively addressed through advanced nanoparticle platforms [11]. As these technologies mature, the fusion of genetic precision with traditional antimicrobials holds the promise of effectively dismantling biofilms, offering new hope in the fight against persistent and drug-resistant infections.
The escalating crisis of antibiotic resistance has catalyzed the development of next-generation antimicrobial strategies, with CRISPR-based technologies, phage therapy, and quorum sensing inhibitors (QSIs) emerging as leading contenders. Each approach operates through distinct molecular mechanisms and offers unique advantages and limitations for combating biofilm-associated and resistant infections. This technical analysis provides a comparative evaluation of these platforms, focusing on their efficacy in disrupting quorum sensing and biofilm formation, with specific emphasis on translational applications for research and drug development. Quantitative metrics reveal that integrated approaches, particularly CRISPR-nanoparticle systems, demonstrate superior biofilm reduction capabilities (>90%) in controlled settings, while QSIs and engineered phages offer promising anti-virulence strategies with potentially lower resistance selection pressures.
Table 1: Quantitative Comparison of Core Anti-Biofilm Technologies
| Technology | Primary Mechanism | Biofilm Reduction (In Vitro) | Key Molecular Target | Therapeutic Specificity | Delivery Challenges |
|---|---|---|---|---|---|
| CRISPR-Cas | Gene editing & disruption | >90% [24] | Resistance genes, QS regulators (e.g., LasR, RhlR) [25] | Sequence-specific | High (requires efficient delivery vector) |
| Phage Therapy | Bacterial lysis & biofilm degradation | Variable (strain-dependent) | Surface receptors (e.g., Type IV pili) [82] | Species/strain-specific | Low to Moderate |
| Quorum Sensing Inhibitors (QSIs) | Signal interference & virulence attenuation | Not quantified in results | AHL signals, LasI/RhlI synthases [83] | Pathway-specific | Low |
In-Depth Technology Analysis
CRISPR-Cas Systems: Precision Genetic Warfare
CRISPR-Cas systems have evolved from a bacterial adaptive immune mechanism into a programmable platform for precision antimicrobial therapy. The technology leverages Cas nucleases guided by RNA molecules to target and disrupt specific genetic sequences fundamental to bacterial virulence, antibiotic resistance, and biofilm formation [25].
1.1.1 Core Mechanisms and Applications The primary application of CRISPR-Cas against bacterial pathogens involves the targeted disruption of essential genes through Double-Strand Breaks (DSBs). This can be directed against: (1) Antibiotic resistance genes (e.g., blaNDM-1, mecA), resensitizing bacteria to conventional antibiotics; (2) Quorum sensing regulators (e.g., LasR, RhlR in P. aeruginosa), effectively silencing bacterial communication; and (3) Biofilm structural genes (e.g., alginate biosynthesis operons), compromising biofilm integrity [24] [44].
Alternative modalities like CRISPR interference (CRISPRi) using catalytically dead Cas9 (dCas9) enable reversible gene silencing without permanent DNA damage. This is particularly valuable for functional genomics studies of essential genes and for transiently suppressing virulence factors during infection [25].
1.1.2 Experimental Protocol: CRISPR-Cas9-Mediated Biofilm Disruption
- Objective: To disrupt the LasI/LasR quorum sensing circuit in Pseudomonas aeruginosa and assess subsequent biofilm inhibition.
- Materials:
- pCas9-gRNA plasmid (constitutively expressing Cas9 and gRNA targeting lasR)
- P. aeruginosa PAO1-GFP (constitutively expressing GFP for biomass quantification)
- Cationic lipid nanoparticles (LNPs) or conjugative plasmids for delivery
- LB medium, 96-well polystyrene plates, confocal microscopy supplies
- Methodology:
- gRNA Design: Design and clone gRNAs with 20-nt spacers complementary to the lasR gene promoter and coding sequence.
- Delivery: Transform P. aeruginosa with the pCas9-gRNA construct via conjugation or electroporation. For in vivo delivery, encapsulate CRISPR machinery in LNPs optimized for bacterial uptake [24].
- Biofilm Assay: Incubate transformed bacteria in 96-well plates for 48 hours. Stain biofilm with crystal violet and quantify via solubilization in acetic acid with OD590 measurement.
- Validation: Confirm lasR gene editing via DNA sequencing. Quantify reduction in QS-controlled virulence factors (pyocyanin, elastase) and visualize biofilm architecture using confocal laser scanning microscopy (CLSM) [44].
Quorum Sensing Inhibitors: Silencing Bacterial Communication
Quorum Sensing Inhibitors (QSIs) represent an anti-virulence strategy that attenuates bacterial pathogenicity without inducing lethal pressure, thereby potentially reducing resistance selection. These compounds interfere with the production, detection, or interpretation of acyl-homoserine lactone (AHL) signals that coordinate group behaviors [83].
1.2.1 Core Mechanisms and Applications QSIs disrupt biofilm formation and virulence by targeting key nodes in the QS network. Natural compounds like resveratrol and curcumin inhibit AHL synthase activity (e.g., LasI) and compete for receptor binding (e.g., LasR) [83]. Enzymatic QSIs, such as AHL lactonases, degrade signaling molecules directly in the extracellular environment, effectively "deafening" the bacterial population [83]. This disruption leads to a significant downregulation of virulence factors, including pyocyanin production and protease secretion, and results in poorly formed, susceptible biofilms.
A critical consideration is the potential for QSIs to indirectly influence other resistance pathways. For instance, QS inhibition in P. aeruginosa downregulates the CRISPR-Cas immune system and Type IV pilus expression, which can unexpectedly favor the evolution of CRISPR-based phage resistance by reducing phage adsorption rates [9].
1.2.2 Experimental Protocol: Evaluating QSI Efficacy Against Biofilms
- Objective: To quantify the efficacy of a natural QSI (e.g., resveratrol) in inhibiting P. aeruginosa biofilm formation and virulence.
- Materials:
- P. aeruginosa PA14 wild-type strain
- Resveratrol stock solution (in DMSO)
- AHL biosensor strains (e.g., Chromobacterium violaceum for violacein inhibition assay)
- Microtiter plates, spectrophotometer
- Methodology:
- Autoinducer Bioassay: Co-culture P. aeruginosa with sub-inhibitory concentrations of resveratrol. Filter the supernatant and apply to an AHL biosensor lawn to visualize and quantify reduction in AHL production via zone-of-inhibition assays [83].
- Virulence Factor Quantification: Grow P. aeruginosa with and without resveratrol. Measure pyocyanin concentration in chloroform-HCl extracts via OD520 and quantify protease activity using azocasein hydrolysis assays.
- Biofilm Analysis: Culture bacteria in the presence of resveratrol in flow cells or microplates. Quantify biofilm biomass and analyze 3D architecture using COMSTAT software on CLSM images.
Phage Therapy: Evolutionary Biological Control
Bacteriophage (phage) therapy utilizes viruses to specifically infect and lyse bacterial pathogens. Its efficacy is often limited by narrow host ranges and the rapid evolution of bacterial resistance, which can be mitigated through phage adaptive evolution and exploitation of bacterial fitness trade-offs [82].
1.3.1 Core Mechanisms and Applications Phages infect bacteria by attaching to specific surface receptors (e.g., Type IV pili, LPS), leading to bacterial lysis and biofilm penetration. Some phages produce depolymerases that degrade the extracellular polymeric substance (EPS) matrix of biofilms, enhancing antibiotic penetration [82].
A key advanced strategy is adaptive evolution (e.g., the Appelmans protocol), where phages are serially passaged through a diverse population of bacterial strains, including resistant mutants. This directs the evolution of phages with expanded host ranges, often through mutations in Receptor-Binding Proteins (RBPs), enabling them to overcome surface resistance mutations [82]. Furthermore, bacteria that evolve phage resistance through surface receptor modification often incur fitness trade-offs, such as restored antibiotic susceptibility or reduced virulence, creating opportunities for combination therapies [82].
1.3.2 Experimental Protocol: Adaptive Evolution of Phages to Counter Resistance
- Objective: To generate evolved phages capable of infecting phage-resistant P. aeruginosa isolates.
- Materials:
- Ancestor phage (e.g., DMS3vir)
- P. aeruginosa PA14 (sensitive and isogenic resistant mutants)
- M9 media, soft agar, filtration units (0.22 µm)
- Methodology:
- Evolution Setup: Initiate co-cultures of the ancestral phage and a mixed bacterial population (containing sensitive and resistant isolates) in liquid medium.
- Serial Passaging: At 24-hour intervals, filter the culture supernatant and use it to infect a fresh bacterial culture, repeatedly cycling the phage population.
- Plaque Assay Screening: After 10-15 cycles, plate phage isolates on lawns of initially resistant bacterial strains. Isolate phages that form clear plaques, indicating regained infectivity.
- Characterization: Sequence the tail fiber protein genes of evolved phages to identify mutations responsible for host range expansion. Test the infectivity of evolved phages against a panel of clinical isolates to determine the breadth of host range expansion [82].
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Anti-Biofilm and Anti-Virulence Research
| Reagent / Tool | Function/Application | Example Use-Case |
|---|---|---|
| dCas9-KRAB CRISPRi System | Targeted gene repression without DNA cleavage; for studying essential genes. | Reversible silencing of QS master regulator lasR to assess virulence attenuation [25]. |
| Lipid Nanoparticles (LNPs) | In vivo delivery of CRISPR payloads (RNP/mRNA); enhanced biofilm penetration. | Delivery of Cas9-gRNA complexes targeting ecaA gene in E. coli biofilms [24]. |
| AHL Biosensor Strains | Detection and quantification of AHL-based quorum sensing signals. | Visual confirmation of QSI activity by monitoring violacein pigment reduction in C. violaceum [83]. |
| Baicalein | A specific small-molecule inhibitor of the LasI/RhlI QS systems in P. aeruginosa. | Experimental dissection of QS influence on phage receptor (Type IV pilus) availability and CRISPR-Cas immunity [9]. |
| Engineered Phage Libraries | Pre-adapted phage collections with expanded host ranges against common resistant pathogens. | Rapid screening for effective phage candidates against multidrug-resistant (MDR) clinical isolates, bypassing de novo evolution [82]. |
Integrated Pathway and Workflow Visualizations
CRISPR-QSI Crosstalk in Pseudomonas aeruginosa
Integrated Experimental Workflow for Technology Evaluation
The fight against antimicrobial resistance necessitates a new arsenal of precision tools. CRISPR-based technologies, phage therapy, and quorum sensing inhibitors each present a powerful, distinct mechanism of action. The future of antimicrobial therapy lies not in selecting a single winner but in designing intelligent combination therapies that leverage the unique strengths of each platform. For instance, using QSIs to attenuate virulence and suppress CRISPR-mediated phage immunity, followed by administration of evolved phages to lyse bacterial cells, could be a highly effective strategy. Simultaneously, CRISPR-based precision antimicrobials could be deployed to selectively eradicate resistance genes from complex microbial communities, preserving the beneficial microbiome. Overcoming delivery challenges, particularly for CRISPR components, through advanced nanoparticle systems is critical for clinical translation. As these technologies mature, they will collectively form a versatile, powerful, and evolution-resistant framework for addressing the global crisis of antibiotic-resistant infections.
The persistent challenge of biofilm-associated infections necessitates advanced research tools that can transition effectively from controlled laboratory settings to complex physiological environments. CRISPR-Cas technology has emerged as a transformative approach for precisely dissecting the molecular mechanisms underlying bacterial social behaviors, particularly quorum sensing (QS) and biofilm formation [25] [20]. This technical guide outlines validated methodologies for employing CRISPR-based systems across a spectrum of experimental models, from initial in vitro biofilm characterization to sophisticated in vivo infection systems, providing researchers with a comprehensive framework for mechanistic investigation and therapeutic development.
The protective matrix of biofilms, comprising extracellular polymeric substances (EPS), confers remarkable tolerance to conventional antimicrobials, with biofilm-embedded bacteria exhibiting up to 1000-fold greater resistance compared to their planktonic counterparts [11]. This resilience is orchestrated by complex genetic networks regulating bacterial surface adhesion, microcolony formation, EPS production, and maturation—processes tightly controlled by QS systems [25]. CRISPR-Cas systems offer unprecedented precision in manipulating these pathways, enabling researchers to move beyond correlation to direct causal validation of gene function in biofilm biology [44] [20].
CRISPR-Cas Mechanisms in Biofilm Research
CRISPR-Cas systems function as adaptive immune mechanisms in prokaryotes but have been repurposed as programmable molecular tools for genetic manipulation [20]. These systems consist of two key components: the Cas nuclease, which introduces double-strand breaks in DNA, and a guide RNA (gRNA) that directs Cas9 to specific genomic sequences through complementary base pairing [11]. In biofilm research, this technology enables targeted disruption of genes involved in QS, EPS production, adhesion, and virulence regulation.
The applications of CRISPR-Cas in biofilm studies extend beyond simple gene knockout to include:
- CRISPR interference (CRISPRi): Utilizing catalytically dead Cas9 (dCas9) to block transcription without permanent genetic alteration, allowing reversible gene silencing of QS regulators like LasI and RhlR in Pseudomonas aeruginosa [25]
- CRISPR activation (CRISPRa): Employing modified dCas9 fused to transcriptional activators to enhance expression of biofilm suppressor genes
- Precision antimicrobials: Designing sequence-specific antimicrobials that selectively target pathogenicity genes while preserving commensal microbiota [25]
- Functional genomics: Systematically dissecting complex genetic networks controlling biofilm development through multiplexed gRNA libraries [25]
The selection of appropriate CRISPR tools depends on experimental objectives, whether requiring permanent genetic modification (Cas9 nucleases) or transient modulation (CRISPRi/a), with careful consideration of delivery mechanisms and specificity controls to ensure meaningful results in complex models.
In Vitro Biofilm Models and Validation Methodologies
Static Biofilm Formation Assays
Protocol: Microtiter Plate Biofilm Formation with Isogenic Mutants
Bacterial Strain Preparation:
- Generate knockout mutants using CRISPR-Cas9 with homology-directed repair (HDR) templates
- For A. baumannii cas3 deletion mutants: Design gRNAs flanking the cas3 gene and co-transform with 500bp homologous arms [14]
- Culture validated mutants in appropriate medium (e.g., LB for A. baumannii) to mid-exponential phase (OD600 = 0.6)
Biofilm Setup:
- Dilute bacterial cultures 1:100 in fresh medium containing essential nutrients for biofilm formation
- Dispense 200μL aliquots into sterile 96-well polystyrene plates
- Incubate statically at 37°C for 24-48 hours
Biofilm Quantification:
- Carefully remove planktonic cells by inverting plates and washing twice with phosphate-buffered saline (PBS)
- Fix adherent cells with 200μL of 99% methanol for 15 minutes
- Air dry wells completely, then stain with 200μL of 0.1% crystal violet for 15 minutes
- Gently rinse excess stain under running tap water and air dry
- Destain with 200μL of 33% glacial acetic acid with shaking for 30 minutes
- Measure OD570 of solubilized crystal violet using plate reader
Validation Data: In A. baumannii, deletion of type I-Fa cas3 significantly reduced biofilm formation by approximately 70% compared to wild-type strains, while complementation restored biofilm formation to near-wild-type levels [14].
Flow Cell Systems for Biofilm Architecture Analysis
Protocol: Confocal Laser Scanning Microscopy (CLSM) of 3D Biofilm Structures
Biofilm Growth Conditions:
- Assemble flow cells with appropriate surface materials (e.g., glass, silicone, or plastic)
- Inoculate with bacterial suspensions (OD600 = 0.1) and allow initial attachment for 1 hour without flow
- Initiate medium flow at rate of 0.2mm/s to create low shear stress environment
- Maintain continuous flow for 48-72 hours with defined medium
Biofilm Staining:
- Gently introduce fluorescent stains via injection port
- For EPS visualization: Use Alexa Fluor 647-conjugated dextran (10μg/mL) with 30-minute incubation [14]
- For bacterial cells: Counterstain with SYTO9 green fluorescent nucleic acid stain (5μM) for 15 minutes
- Restore flow for 15 minutes to remove unbound dye
Image Acquisition and Analysis:
- Acquire z-stack images at 1μm intervals using 20x or 40x objectives
- Set laser powers and gain to avoid signal saturation
- Analyze minimum of 5 random fields per sample
- Quantify biofilm parameters: biomass, thickness, surface coverage, and roughness coefficient
Validation Data: CLSM analysis demonstrated that A. baumannii Δcas3 mutants formed significantly thinner biofilms with reduced EPS matrix compared to wild-type strains, indicating Cas3's role in maintaining proper biofilm architecture [14].
High-Throughput Genetic Screening in Biofilms
Protocol: CRISPR Library Screening for Biofilm-Associated Genes
Library Design and Delivery:
- Construct pooled gRNA library targeting 100-500 genes potentially involved in biofilm regulation
- Clone gRNA library into appropriate CRISPR plasmid backbone (e.g., lentiviral vector)
- Transfer library to target bacteria via conjugation or electroporation at high coverage (>500x)
Selection and Enrichment:
- Infect flow cell systems with CRISPR library-containing bacteria
- Allow biofilm development for 72 hours under continuous flow
- Collect biofilm and planktonic fractions separately
- Isplicate genomic DNA from both populations
Sequencing and Hit Identification:
- Amplify gRNA regions by PCR with barcoded primers
- Sequence on Illumina platform to minimum depth of 10 million reads
- Calculate enrichment/depletion scores comparing biofilm vs. planktonic fractions
- Validate hits using individual mutant strains in secondary assays
Table 1: Quantitative Assessment of CRISPR-Mediated Biofilm Modulation in Pathogenic Bacteria
| Bacterial Species | Target Gene | CRISPR Approach | Biofilm Reduction | Experimental Model |
|---|---|---|---|---|
| Acinetobacter baumannii | cas3 (Type I-Fa) | Gene knockout | ~70% reduction [14] | Microtiter plate, CLSM |
| Escherichia coli | Quorum sensing & adhesion genes | CRISPR-Cas9 HDR | ~2.5 log reduction [25] | Urinary catheter model |
| Pseudomonas aeruginosa | lasI/rhlR QS system | CRISPRi (dCas9) | ~65% inhibition [25] | Flow cell system |
| Klebsiella pneumoniae | 1330, htpx, yjgH | CRISPR-Cas9 knockout | Significant reduction in biofilm & virulence [84] | A549 cell adhesion assay |
Advanced In Vivo Infection Models
Galleria mellonella Infection Model
Protocol: Virulence Assessment in Wax Moth Larvae
Bacterial Preparation:
- Grow wild-type and CRISPR-modified bacteria to mid-log phase
- Harvest cells by centrifugation and wash twice with phosphate-buffered saline (PBS)
- Adjust bacterial density to 1×10^7 CFU/mL in PBS
- Prepare serial dilutions for infection (typically 1×10^5 to 1×10^6 CFU/larva)
Larval Infection and Monitoring:
- Select healthy Galleria mellonella larvae (250-350mg in weight)
- Disinfect prolegs with 70% ethanol before injection
- Inject 10μL bacterial suspension into hemocoel via last proleg using microsyringe
- Include PBS-injected controls for background mortality
- Incubate injected larvae at 37°C in petri dishes
- Monitor survival every 12 hours for 3-5 days, recording death as lack of movement to stimulus
Post-Mortem Analysis:
- Sacrifice surviving larvae at endpoint for bacterial burden assessment
- Homogenize individual larvae in 1mL PBS
- Plate serial dilutions on selective media for CFU enumeration
- Compare bacterial loads between wild-type and mutant strains
Validation Data: In A. baumannii infection models, larvae infected with the Δcas3 mutant showed significantly higher survival rates (50% at 96 hours) compared to those infected with wild-type strains (100% mortality within 24 hours), demonstrating Cas3's critical role in pathogenicity [14].
Murine Bacteremia and Tissue Colonization Models
Protocol: Systemic Infection for Virulence Factor Validation
Animal Preparation and Ethical Considerations:
- Use 6-8 week old, sex-matched mice (e.g., C57BL/6 or BALB/c)
- Acclimate animals for 1 week prior to experimentation
- Randomize into experimental groups (n=8-10 per group)
- Obtain institutional IACUC approval following 3Rs principles
Infection Procedure:
- Prepare bacterial inoculum as described for Galleria model
- Inject 100μL containing 1×10^7 CFU via tail vein for systemic infection
- Monitor mice twice daily for clinical signs: weight loss, posture, activity, piloerection
- Establish humane endpoints (e.g., >20% weight loss, lethargy, inability to access food/water)
Sample Collection and Analysis:
- Euthanize moribund mice or at predetermined endpoints
- Aseptically collect organs (spleen, liver, lungs) for bacterial enumeration
- Homogenize tissues in PBS and plate serial dilutions for CFU counting
- Preserve tissues in formalin for histopathological examination
- Collect blood for cytokine analysis (IL-6, TNF-α, IL-1β) via ELISA
Validation Data: Mice infected with A. baumannii Δcas3 mutants showed significantly reduced bacterial loads in organs and milder lung inflammation compared to wild-type infections, confirming the role of Cas3 in virulence and pathogenicity [14].
Device-Associated Biofilm Infection Models
Protocol: Catheter-Associated Biofilm Establishment
Catheter Preparation and Implantation:
- Sterilize silicone or polyethylene catheter segments (5mm length)
- Pre-coat with host proteins (e.g., fibronectin) if studying conditioned surface attachment
- Surgically implant subcutaneously in murine dorsal region under anesthesia
- Allow 1-week recovery for foreign body capsule formation
Biofilm Infection and Monitoring:
- Directly inoculate catheters with 1×10^5 CFU of CRISPR-modified bacteria
- Monitor local infection signs: erythema, swelling, pus formation
- Track systemic manifestations: weight loss, temperature changes
Ex Vivo Biofilm Analysis:
- Explant catheters at designated timepoints (3-7 days post-infection)
- Gently rinse with PBS to remove non-adherent cells
- Process for: (1) CFU enumeration after sonication, (2) SEM visualization, or (3) CLSM with fluorescent stains
- Compare biofilm architecture and bacterial viability between strains
Table 2: Nanoparticle-Mediated CRISPR Delivery Systems for Enhanced Biofilm Penetration
| Nanoparticle Type | CRISPR Payload | Target Bacteria | Key Outcomes | Advantages for In Vivo Use |
|---|---|---|---|---|
| Liposomal nanoparticles | Cas9/sgRNA targeting resistance genes | Pseudomonas aeruginosa | >90% reduction in biofilm biomass [11] | Enhanced stability, controlled release |
| Gold nanoparticles | CRISPR-Cas9 ribonucleoproteins | Multiple pathogens | 3.5× increase in editing efficiency [11] | Excellent biocompatibility, surface functionalization |
| Polymeric nanoparticles (PEI) | CRISPR plasmids | Staphylococcus aureus | Synergistic effect with antibiotics [11] | Protection from nucleases, high loading capacity |
Integrated Analytical Approaches
Molecular Validation of CRISPR Effects
Protocol: Transcriptomic Analysis of QS and Biofilm Genes
RNA Extraction from Biofilms:
- Grow biofilms of wild-type and CRISPR-modified strains under identical conditions
- Harvest biofilm cells mechanically or enzymatically (e.g., DNase I treatment)
- Preserve RNA immediately using RNA stabilization reagents
- Extract total RNA using column-based kits with DNase treatment
- Assess RNA quality (RIN >8.0) using bioanalyzer
Gene Expression Profiling:
- Prepare cDNA libraries from high-quality RNA samples
- Perform RNA sequencing (Illumina platform) at minimum depth of 20 million reads/sample
- Analyze differential expression of QS regulators (lasR, rhlR, luxS), biofilm-related genes (pel, psl, algD), and virulence factors
- Validate key findings using RT-qPCR with reference genes
Pathway and Network Analysis:
- Map differentially expressed genes to KEGG pathways and Gene Ontology terms
- Construct gene regulatory networks using bioinformatics tools
- Identify upstream regulators and downstream effectors of targeted genes
Validation Data: Transcriptomic analysis of A. baumannii Δcas3 mutants revealed significant downregulation of virulence factors including biofilm formation-related genes and ompA, along with alterations in carbon metabolism and oxidative phosphorylation pathways [14].
Computational Integration and AI-Assisted Modeling
Protocol: Machine Learning for Predictive Modeling of Biofilm Intervention
Feature Selection and Data Compilation:
- Compile dataset from multi-omics analyses: genomics, transcriptomics, proteomics
- Include experimental parameters: biofilm biomass, thickness, dispersal rates
- Incorporate CRISPR screening results and fitness scores
- Normalize data across experiments and platforms
Model Training and Validation:
- Implement random forest or neural network architectures
- Partition data into training (80%) and validation (20%) sets
- Perform 5-fold cross-validation to prevent overfitting
- Optimize hyperparameters using grid search approaches
Therapeutic Target Prioritization:
- Apply trained models to rank potential CRISPR targets by predicted efficacy
- Consider essentiality, conservation, and accessibility in scoring
- Validate top predictions in secondary assays
- Iteratively refine models with new experimental data
Validation Data: Machine learning models applied to pneumonia Klebsiella resistance prediction demonstrated >86% accuracy for 11 antimicrobial drugs with AUC values exceeding 0.9, highlighting the power of computational approaches in biofilm research [84].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for CRISPR-Biofilm Investigations
| Reagent Category | Specific Examples | Research Application | Technical Considerations |
|---|---|---|---|
| CRISPR Delivery Systems | Liposomal Cas9 formulations, Gold nanoparticle carriers, Engineered phages | Efficient delivery of CRISPR components through biofilm matrix [11] | Optimize for target bacteria, minimize cytotoxicity |
| Biofilm Staining Reagents | SYTO9, Alexa Fluor-dextran conjugates, Crystal violet | Visualization and quantification of biofilm biomass and architecture [14] | Validate staining specificity, optimize dye concentrations |
| QS Signal Molecule Analogs | AHL analogs, AI-2 precursors, Quorum quenching enzymes | Modulation of bacterial communication pathways [84] | Determine effective concentrations, monitor stability |
| Gene Expression Tools | dCas9-VP64 (CRISPRa), dCas9-KRAB (CRISPRi), RNA-seq kits | Precise control of biofilm gene expression without DNA cleavage [25] | Optimize gRNA designs for regulatory elements |
| Animal Model Reagents | Galleria mellonella larvae, Murine catheter materials, Tissue homogenizers | Validation of biofilm interventions in complex host environments [14] | Establish ethical endpoints, standardize infection doses |
Visualizing Experimental Workflows and Signaling Pathways
Integrated CRISPR-Biofilm Validation Pipeline
CRISPR Modulation of Quorum Sensing Pathways
In Vivo Biofilm Infection Model Workflow
The emergence of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-Cas systems as precision tools in antibacterial research represents a paradigm shift in tackling biofilm-mediated infections [11] [44]. Within the broader thesis context of CRISPR's role in bacterial quorum sensing and biofilm formation research, this whitepaper addresses the critical translational challenge of long-term safety and stability assessment. While CRISPR technologies show remarkable efficacy in disrupting quorum sensing pathways and biofilm-associated antibiotic resistance genes [11] [78], their path to clinical application requires rigorous evaluation of genotoxicity and immune responses.
Biofilms, structured microbial communities embedded in extracellular polymeric substances, contribute significantly to antimicrobial resistance by creating protective niches that limit drug penetration and enhance horizontal gene transfer [11] [55]. CRISPR-based interventions targeting these complex communities must demonstrate not only efficacy but also minimal off-target effects and acceptable immune profiles. This technical guide provides researchers, scientists, and drug development professionals with comprehensive methodologies for assessing these critical safety parameters, enabling the responsible advancement of CRISPR-based anti-biofilm therapies toward clinical application.
CRISPR in Biofilm Research: Technical Foundations
CRISPR-Cas Systems: From Bacterial Immunity to Therapeutic Applications
CRISPR-Cas systems function as adaptive immune mechanisms in prokaryotes, providing sequence-specific defense against invasive genetic elements [20]. These systems consist of CRISPR arrays containing short DNA repeats interspersed with spacer sequences acquired from previous invaders, coupled with Cas genes encoding effector proteins [20]. The system operates through three distinct stages: adaptation (spacer acquisition from foreign DNA), expression (CRISPR RNA processing), and interference (target cleavage guided by crRNA) [20].
The classification of CRISPR-Cas systems into two classes (Class 1 and Class 2) and six types (I-VI) reflects their evolutionary diversity and functional specialization [20]. Class 2 systems, particularly Type II featuring the Cas9 nuclease, have been extensively repurposed for biomedical applications due to their simplicity and programmability [20]. In biofilm research, CRISPR systems have been engineered both to study biofilm formation mechanisms and to develop novel antimicrobial strategies targeting biofilm-associated infections [44] [78].
Molecular Interplay Between CRISPR and Biofilm Pathways
The relationship between native CRISPR-Cas systems and bacterial virulence is complex and context-dependent. In Acinetobacter baumannii, Cas3 of the type I-Fa CRISPR-Cas system upregulates biofilm formation and virulence by modulating expression of virulence factors such as outer membrane protein A (OmpA) and metabolic pathways including carbon metabolism and oxidative phosphorylation [14]. Deletion of cas3 significantly reduces biofilm formation, adhesion, invasion, and pathogenicity in murine models [14].
Conversely, CRISPR-based interventions can be designed to target essential biofilm processes. Key strategic targets include:
- Quorum sensing networks that coordinate population-level behaviors [29]
- Genes encoding extracellular polymeric matrix components [11]
- Antibiotic resistance genes housed within biofilm communities [11] [44]
- Regulatory systems such as two-component systems and c-di-GMP signaling pathways [78]
Table 1: Key Biofilm Targets for CRISPR-Based Interventions
| Target Category | Specific Molecular Targets | Anticipated Outcome |
|---|---|---|
| Quorum Sensing | AHL synthases, transcription factors | Disrupted cell-cell communication and coordination |
| Matrix Synthesis | Polysaccharide biosynthesis operons, adhesion proteins | Reduced structural integrity and adhesion capability |
| Antibiotic Resistance | β-lactamases, efflux pump genes, resistance plasmids | Resensitization to conventional antibiotics |
| Metabolic Regulation | c-di-GMP metabolic enzymes, GacA/S two-component system | Inhibition of motile-to-sessile transition |
Methodologies for Genotoxicity Assessment
In Vitro Genotoxicity Screening Platforms
Comprehensive genotoxicity assessment begins with high-throughput in vitro screening to identify potential DNA damage responses triggered by CRISPR-based antimicrobials. The following experimental protocol provides a standardized approach for initial risk evaluation.
Protocol 3.1: High-Throughput Genotoxicity Screening
Purpose: To assess DNA damage potential of CRISPR-Cas formulations in human cell lines and bacterial biofilms.
Materials:
- Human epithelial cell lines (HEK293, A549)
- Bacterial biofilm models (Pseudomonas aeruginosa, Acinetobacter baumannii)
- CRISPR-Cas9 formulations (nanoparticle-encapsulated or free)
- γH2AX immunofluorescence staining kit
- Comet assay reagents
- SOS response reporter strain in target bacteria
Procedure:
- Cell Exposure: Treat human cell monolayers with CRISPR formulations at concentrations ranging from 0.1-100 μg/mL for 24-72 hours.
- Immunofluorescence Staining: Fix cells and stain for γH2AX foci, a sensitive marker of DNA double-strand breaks.
- Comet Assay: Perform alkaline comet assay to detect single and double-strand DNA breaks.
- SOS Response Evaluation: Transform target bacteria with SOS-responsive promoter fused to reporter gene; measure activation after CRISPR exposure.
- Quantitative Analysis: Image and quantify γH2AX foci per nucleus; measure comet tail moments; quantify reporter gene expression.
Data Interpretation: Compare results to positive controls (known genotoxins) and negative controls (untreated cells). A dose-dependent increase in DNA damage markers indicates potential genotoxicity concerns requiring further investigation.
Off-Target Editing Analysis in Complex Biofilm Communities
The heterogeneous nature of biofilms presents unique challenges for assessing CRISPR specificity. The following methodology enables comprehensive evaluation of off-target effects in biologically relevant systems.
Protocol 3.2: Off-Target Analysis in Multi-Species Biofilms
Purpose: To identify and quantify unintended genetic modifications in target and non-target species within biofilm consortia.
Materials:
- Defined multi-species biofilm community including target pathogen and commensal strains
- CRISPR-Cas formulation with known on-target site
- Whole-genome sequencing services
- Bioinformatic tools for CRISPR off-target prediction (Cas-OFFinder, GUIDE-seq)
- Quantitative PCR reagents
- Fluorescence in situ hybridization (FISH) probes
Procedure:
- Biofilm Establishment: Grow multi-species biofilms in flow cells or microtiter plates for 48-72 hours.
- CRISPR Treatment: Apply CRISPR formulations at therapeutically relevant concentrations.
- Genomic DNA Extraction: Harvest biofilm biomass and extract high-quality genomic DNA from total community and individual species.
- Whole-Genome Sequencing: Perform deep sequencing (≥50x coverage) of pre- and post-treatment samples.
- Bioinformatic Analysis:
- Map sequencing reads to reference genomes
- Identify insertions/deletions (indels) at predicted off-target sites
- Characterize novel mutations using variant calling pipelines
- Validation: Design PCR assays for putative off-target sites; confirm by Sanger sequencing.
Data Interpretation: Off-target rates should be <0.1% of on-target activity for clinical consideration. Species-specific analysis should confirm minimal impact on non-target commensals.
Diagram 1: Comprehensive Genotoxicity Assessment Workflow
Quantitative Genotoxicity Endpoints
Standardized quantitative metrics enable comparison across different CRISPR platforms and formulations. The following table summarizes key parameters for genotoxicity assessment.
Table 2: Quantitative Genotoxicity Endpoints for CRISPR Safety Assessment
| Assessment Method | Measured Parameter | Acceptance Criteria | Clinical Relevance |
|---|---|---|---|
| γH2AX Foci Count | DNA double-strand breaks | <5 foci/nucleus (vs. control) | Predictive of chromosomal instability |
| Comet Assay | Tail moment (DNA fragmentation) | ≤2x negative control | Detection of single/double strand breaks |
| SOS Response Activation | Reporter gene expression | <2-fold induction | Bacterial stress response to DNA damage |
| Off-Target Mutation Rate | Indels at predicted sites | <0.1% of on-target activity | Editing specificity in complex communities |
| Chromosomal Aberrations | Micronuclei formation | <2% of cells (vs. baseline) | Cytogenetic damage assessment |
Immune Response Profiling
Innate Immune Activation Assays
CRISPR components can trigger pattern recognition receptors, initiating inflammatory cascades that may compromise therapeutic efficacy or cause tissue damage. The following protocol details methodology for comprehensive innate immune response assessment.
Protocol 4.1: Innate Immune Response Profiling
Purpose: To characterize innate immune recognition of CRISPR-Cas formulations and associated nanocarriers.
Materials:
- Human peripheral blood mononuclear cells (PBMCs) from healthy donors
- THP-1 monocyte cell line differentiated to macrophages
- TLR-transfected HEK293 reporter cells
- Cytokine multiplex assay (TNF-α, IL-6, IL-1β, IFN-α/β)
- Western blot reagents for NF-κB and IRF3 signaling
- Flow cytometry with surface marker antibodies (CD14, CD86, HLA-DR)
Procedure:
- Cell Stimulation:
- Treat PBMCs and macrophages with CRISPR formulations (0.1-10 μg/mL) for 6-24 hours
- Include controls: LPS (TLR4 agonist), imiquimod (TLR7 agonist), transfection reagent alone
- Cytokine Profiling: Collect supernatants; measure 12-plex cytokine panel using Luminex technology
- Signaling Pathway Analysis: Prepare cell lysates; perform Western blot for phospho-NF-κB, IRF3, and STAT1
- Surface Marker Expression: Analyze activation markers (CD86, HLA-DR) on monocytes/macrophages by flow cytometry
- TLR-Specific Activation: Treat TLR reporter cells; measure luciferase activity after 18 hours
Data Interpretation: Clinically acceptable formulations should demonstrate <50% of the inflammatory response elicited by positive controls. TLR activation should be minimal, with cytokine levels below established safety thresholds.
Adaptive Immune Monitoring
Pre-existing or therapy-induced adaptive immunity against CRISPR components presents significant challenges for repeated administration. The following methodology enables comprehensive assessment of humoral and cellular immune responses.
Protocol 4.2: Adaptive Immune Response Evaluation
Purpose: To detect and quantify Cas protein-specific antibodies and T-cell responses.
Materials:
- Serum samples from pre- and post-treatment timepoints
- Recombinant Cas proteins (Cas9, Cas3, etc.)
- ELISpot kits for IFN-γ and IL-4
- Peptide libraries spanning Cas protein sequences
- Flow cytometry with tetramer staining capabilities
Procedure:
- Antibody Detection:
- Develop ELISA using recombinant Cas proteins as capture antigen
- Test serial serum dilutions; establish titers for IgG, IgM, and IgA
- Include positive controls (known positive sera) and negative controls (pre-immune sera)
- T-cell ELISpot:
- Isolate PBMCs from blood samples
- Stimulate with Cas protein peptide libraries (15-mer overlapping peptides)
- Detect IFN-γ and IL-4 secreting cells; quantify spot-forming units
- Tetramer Staining:
- Design MHC-matched tetramers for immunodominant Cas epitopes
- Stain PBMCs; analyze by flow cytometry for antigen-specific CD4+ and CD8+ T-cells
- Memory B-cell Assessment:
- Isolate B-cells; culture with stimulatory cytokines
- Measure Cas-specific antibody secretion by ELISA
Data Interpretation: Pre-existing immunity (seroprevalence) varies by population. Therapy-induced responses should be monitored for correlation with reduced efficacy upon repeated administration.
Diagram 2: Comprehensive Immune Response Profiling Strategy
Immunological Safety Parameters
Standardized immunological parameters enable cross-study comparisons and safety benchmarking. The following table outlines key metrics for comprehensive immune safety assessment.
Table 3: Immune Response Parameters for CRISPR Safety Assessment
| Immune Parameter | Assessment Method | Safety Threshold | Clinical Implications |
|---|---|---|---|
| Inflammatory Cytokines | Multiplex assay (TNF-α, IL-6, IL-1β) | <2x baseline levels | Acute inflammatory potential |
| Type I Interferons | IFN-α/β ELISA or bioassay | <100 pg/mL | Antiviral state induction |
| TLR Activation | Reporter cell assays | <20% of positive control | Nucleic acid sensor engagement |
| Anti-Cas Antibodies | Antigen-specific ELISA | Titer <1:100 (pre-existing) | Neutralizing antibody risk |
| Cas-Specific T-cells | ELISpot (IFN-γ) | <50 SFC/10^6 PBMCs | Cellular immune memory |
| Complement Activation | C3a, C5a measurement | Within normal range | Acute inflammatory potential |
The Scientist's Toolkit: Essential Research Reagents
Successful safety assessment requires carefully selected reagents and controls. The following table details essential materials for comprehensive genotoxicity and immune response evaluation.
Table 4: Essential Research Reagents for Safety Assessment
| Reagent Category | Specific Examples | Function in Safety Assessment | Key Considerations |
|---|---|---|---|
| CRISPR Formulations | Nanoparticle-encapsulated Cas9-gRNA [11], CRISPR-Cas9 hybrids [44] | Test articles for safety evaluation | Include various delivery platforms (liposomal, gold nanoparticles) |
| Control Materials | Empty nanoparticles, catalytically dead Cas9, scrambled gRNA | Control for non-specific effects | Essential for distinguishing CRISPR-specific effects |
| Detection Antibodies | Anti-γH2AX, phospho-NF-κB, CD86, HLA-DR | Immune cell activation and DNA damage markers | Validate for specific applications (flow cytometry, Western blot) |
| Cytokine Assays | Luminex multiplex panels, ELISpot kits | Quantify inflammatory responses | Include both pro-inflammatory and regulatory cytokines |
| Reporter Systems | TLR reporter cells, SOS response strains | Pathway-specific activation assessment | Ensure relevance to human biology |
| Bioinformatic Tools | Cas-OFFinder, GUIDE-seq analysis pipelines | Off-target prediction and validation | Regularly update with latest genome annotations |
Integrated Analysis and Risk Assessment
Correlation of Safety Parameters
A comprehensive safety assessment requires integration of genotoxicity and immunogenicity data to establish structure-activity relationships and identify potential risk mitigation strategies. Key correlation analyses should include:
- Dose-response relationships between CRISPR delivery efficiency and DNA damage markers
- Association between nanocarrier properties (charge, size, composition) and immune activation
- Temporal analysis of acute immune responses versus persistent immunological memory
- Species-specific differences in genotoxicity endpoints to inform translational relevance
Advanced statistical approaches, including multivariate analysis and machine learning algorithms, can identify potential safety signatures predictive of adverse outcomes in later-stage development.
Risk Mitigation Strategies
Based on emerging safety data, several risk mitigation approaches show promise for enhancing the safety profile of CRISPR-based anti-biofilm therapies:
- Delivery system optimization: Lipid nanoparticles with tunable immunogenicity profiles [11]
- Cas enzyme engineering: High-fidelity variants with reduced off-target potential [20]
- Regulatory element targeting: Focus on non-coding RNA and virulence factors rather than essential genes [78]
- Dosing strategy optimization: Pulsed administration to minimize immune activation while maintaining efficacy
- Biodistribution control: Localized delivery to biofilm sites to minimize systemic exposure
The continuous iteration between efficacy assessment and safety profiling represents a crucial feedback loop for responsible therapeutic development. As CRISPR-based anti-biofilm strategies advance toward clinical application, this integrated safety assessment framework provides the necessary foundation for balancing therapeutic innovation with patient safety.
Conclusion
The dynamic interplay between CRISPR-Cas systems and quorum sensing represents a pivotal regulatory node controlling bacterial biofilm formation and virulence. Targeting this axis with precision CRISPR tools offers a paradigm shift from broad-spectrum antimicrobials to highly specific, genetically-defined therapies. The integration of advanced delivery platforms, particularly nanoparticles, is crucial for translating this potential into clinically viable treatments that can penetrate resilient biofilm matrices. Future research must prioritize the refinement of high-fidelity systems to minimize off-target effects, comprehensive in vivo safety profiling, and the development of strategies that preempt bacterial resistance. Overcoming these challenges will pave the way for CRISPR-based interventions to effectively combat biofilm-driven infections, thereby addressing one of the most pressing issues in the global antimicrobial resistance crisis.
References
- 1. Quorum Sensing Controls Adaptive Immunity through the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5179492/]
- 2. Bacterial Quorum Sensing: Its Role in Virulence and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3543102/]
- 3. 有助于绿脓杆菌基于CRISPR免疫系统的噬菌体抗性进化 [https://news.bioon.com/article/8708e1380489.html]
- 4. Relationship Between Quorum Sensing and Secretion ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2019.01100/full]
- 5. Quorum sensing: How bacteria can coordinate activity and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3526984/]
- 6. Plant-Derived Inhibitors of AHL-Mediated Quorum Sensing ... [https://www.mdpi.com/1422-0067/20/22/5588]
- 7. Mechanisms regulating the CRISPR-Cas systems [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2023.1060337/full]
- 8. Controlling and enhancing CRISPR systems - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8101458/]
- 9. The effect of Quorum sensing inhibitors on the evolution ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8319334/]
- 10. CRISPR-CAS9 Against Antibiotic Resistance-A Review [https://pubmed.ncbi.nlm.nih.gov/38702575/]
- 11. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366227/]
- 12. CRISPR-Cas systems: role in cellular processes beyond ... [https://link.springer.com/article/10.1007/s12223-022-00993-2]
- 13. An updated evolutionary classification of CRISPR–Cas ... [https://www.nature.com/articles/s41564-025-02180-8]
- 14. Cas3 of type I-Fa CRISPR-Cas system upregulates ... [https://www.nature.com/articles/s42003-025-08124-6]
- 15. - CRISPR systems target endogenous genes to impact bacterial... Cas [https://link.springer.com/article/10.1186/s43556-022-00084-1]
- 16. of type I-Fa Cas - 3 system upregulates... | CRISPR Square Cas Research [https://www.researchsquare.com/article/rs-4658723/v1]
- 17. BaeR and H-NS control CRISPR-Cas-mediated immunity and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12625773/]
- 18. Quorum Sensing Inhibits Type III‐A CRISPR‐Cas System ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12462963/]
- 19. Quorum Sensing Inhibits Type III-A CRISPR-Cas System ... [https://pubmed.ncbi.nlm.nih.gov/40642869/]
- 20. CRISPR-Cas Systems: Bridging Bacterial Immunity and ... [https://www.mdpi.com/2673-8007/5/4/118]
- 21. CRISPR-Cas: biology, mechanisms and relevance - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5052741/]
- 22. CRISPR-cas3 of Salmonella Upregulates Bacterial Biofilm ... [https://www.mdpi.com/2076-0817/9/1/53]
- 23. CRISPR-epigenetic crosstalk: From bidirectional regulation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12595283/]
- 24. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-025-04323-4]
- 25. CRISPR–Cas systems as emerging tools for precision ... [https://www.sciencedirect.com/science/article/abs/pii/S0963996925021416]
- 26. The Role of CRISPR-Cas Systems in Virulence ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3957734/]
- 27. The virulence regulator CovR boosts CRISPR-Cas9 ... [https://www.nature.com/articles/s41467-025-60871-6]
- 28. The effect of Quorum sensing inhibitors on the evolution of ... [https://www.nature.com/articles/s41396-021-00946-6]
- 29. Biofilm, resistance, and quorum sensing: The triple threat in ... [https://www.sciencedirect.com/science/article/pii/S2950194625003462]
- 30. CRISPRa and CRISPRi [https://www.synthego.com/guide/crispr-methods/crispri-crispra/]
- 31. CRISPR-Cas-mediated transcriptional modulation [https://pmc.ncbi.nlm.nih.gov/articles/PMC10362391/]
- 32. CRISPRa and CRISPRi: Why You Need These Powerful ... [https://bitesizebio.com/47575/crispra-and-crispri/]
- 33. CRISPR interference-mediated silencing of pro ... [https://www.sciencedirect.com/science/article/abs/pii/S1567576925012470]
- 34. CRISPR GENome and epigenome engineering improves loss ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12272254/]
- 35. CMN Weekly (26 September 2025) [https://crisprmedicinenews.com/news/cmn-weekly-26-september-2025-your-weekly-crispr-medicine-news/]
- 36. The role of RNA regulators, quorum sensing and c‐di‐GMP in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10240345/]
- 37. CRISPR Interference (CRISPRi) Inhibition of luxS Gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5445563/]
- 38. In silico sgRNA tool design for CRISPR control of quorum ... [https://www.sciencedirect.com/science/article/pii/S2352304218300564]
- 39. Understanding Quorum-Sensing and Biofilm Forming in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11641729/]
- 40. Small RNAs and their role in biofilm formation - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC3752386/]
- 41. CRISPR gRNA Design Tool [https://www.atum.bio/catalog/vectors/grna-design]
- 42. Innovative approaches to combat antibiotic resistance: integrating... [https://pubmed.ncbi.nlm.nih.gov/40830872/]
- 43. -Cas System and its Role in CRISPR - Quorum ... | SpringerLink Sensing [https://link.springer.com/chapter/10.1007/978-981-99-8529-6_29]
- 44. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm- ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2024.1408569/full]
- 45. Intelligent nanotherapeutic strategies for the delivery of ... [https://www.sciencedirect.com/science/article/pii/S2211383522005226]
- 46. CRISPR's efficiency triples with DNA-wrapped nanoparticles [https://news.northwestern.edu/stories/2025/09/crisprs-efficiency-triples-with-dna-wrapped-nanoparticles]
- 47. Biofilms: structure, resistance mechanism, emerging control ... [https://pubs.rsc.org/en/content/articlehtml/2025/pm/d5pm00094g]
- 48. Harnessing bacterial immunity: CRISPR-Cas system as a ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1588446/full]
- 49. Novel Strategy to Combat Antibiotic Resistance: A Sight ... [https://www.mdpi.com/1999-4923/13/3/352]
- 50. CRISPR-Cas Systems in the Fight Against Antimicrobial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11608035/]
- 51. CRISPR-Cas-Based Antimicrobials: Design, Challenges, ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10353011/]
- 52. Measuring Antimicrobial Efficacy against Biofilms: a Meta- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6496104/]
- 53. Role of CRISPR-Cas systems and anti-CRISPR proteins in ... [https://www.sciencedirect.com/science/article/pii/S2405844024107232]
- 54. The Novel Quantitative Assay for Measuring the Antibiofilm ... [https://www.mdpi.com/2076-3417/10/20/7343]
- 55. Biofilms and multidrug resistance: an emerging crisis ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1625356/full]
- 56. BLOG: Selecting the Right Gene Editing Off-Target Assay [https://seqwell.com/guide-to-selecting-right-gene-editing-off-target-assay/]
- 57. Off-target effects in CRISPR-Cas genome editing for ... [https://www.sciencedirect.com/science/article/pii/S2162253125001908]
- 58. A versatile CRISPR/Cas9 system off-target prediction tool ... [https://www.nature.com/articles/s42003-025-08275-6]
- 59. Improved CRISPR/Cas9 off-target prediction with DNABERT ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12611124/]
- 60. Improving CRISPR-Cas nuclease specificity using ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3988262/]
- 61. CRISPRi-Mediated Gene Silencing in Biofilm Cycle and Quorum ... [https://link.springer.com/chapter/10.1007/978-981-99-8529-6_6]
- 62. Frontiers | CRISPR Interference (CRISPRi) Inhibition of luxS Gene... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2017.00214/full]
- 63. 中科院韩雪祥团队总结用于递送CRISPR基因编辑组件的脂质 ... [https://bydrug.pharmcube.com/news/detail/66383feb3c018a232fb92f1c662d3ba1]
- 64. 综述:纳米颗粒载体:CRISPR/Cas9基因编辑精准化的新时代 [https://www.ebiotrade.com/newsf/2025-5/20250528142307823.htm]
- 65. 综述:探索噬菌体调控肠道菌群失调:微生物组疗法的新前沿 [https://www.ebiotrade.com/newsf/2025-8/20250828003321972.htm]
- 66. Reduction of biofilm of Escherichia coli by targeting... formation [https://pubmed.ncbi.nlm.nih.gov/37271098/]
- 67. 细菌生物膜形成、耐药性及创新防治策略的全面解析与展望 [https://www.forwardpathway.com/270820]
- 68. 系统总结基于CRISPR的基因编辑工具及其递送载体 [https://finance.sina.cn/stock/med/2023-11-05/detail-imztphuw1878792.d.html?from=wap]
- 69. Quantum biological insights into CRISPR-Cas9 sgRNA ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10602897/]
- 70. Classy CRISPR: Differences between Class 1 and Class 2 ... [https://www.synthego.com/blog/crispr-systems/]
- 71. CrisprVi: a software for visualizing and analyzing CRISPR ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-022-04716-9]
- 72. Bacteriophages in Pseudomonas aeruginosa evade the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12174588/]
- 73. Structural and biochemical insights into the mechanism of ... [https://pubmed.ncbi.nlm.nih.gov/39541974/]
- 74. controls the Pseudomonas aeruginosa Quorum -Cas... sensing CRISPR [https://pubmed.ncbi.nlm.nih.gov/27849583/]
- 75. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11257871/]
- 76. A novel dual-staining method for cost-effective visualization ... [https://www.nature.com/articles/s41598-024-80644-3]
- 77. A novel multiplex fluorescent-labeling method for the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11218471/]
- 78. CRISPR interference to interrogate genes that control ... [https://www.nature.com/articles/s41598-019-52400-5]
- 79. CRISPR/Cas9 triumph over biofilm-associated antibiotic ... [https://pubmed.ncbi.nlm.nih.gov/39035353/]
- 80. The Battle Against Biofilms: A Focus on Novel Antimicrobial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11852117/]
- 81. Antimicrobial proteins from oyster hemolymph improve the ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0312305]
- 82. Phage Therapy as a Novel Alternative to Antibiotics Through ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12562026/]
- 83. Using quorum as a novel approach to address... sensing inhibitors [https://www.news-medical.net/news/20250225/Using-quorum-sensing-inhibitors-as-a-novel-approach-to-address-antibiotic-resistance.aspx]
- 84. 杨启文教授团队多项研究入选口头报告,多维覆盖微生物与慢病 [https://www.iidf.com.cn/html/2025-4-21/20250421031900.html]