CRISPR-Cas13: Precision RNA Targeting for Disrupting Biofilm Metabolic Pathways
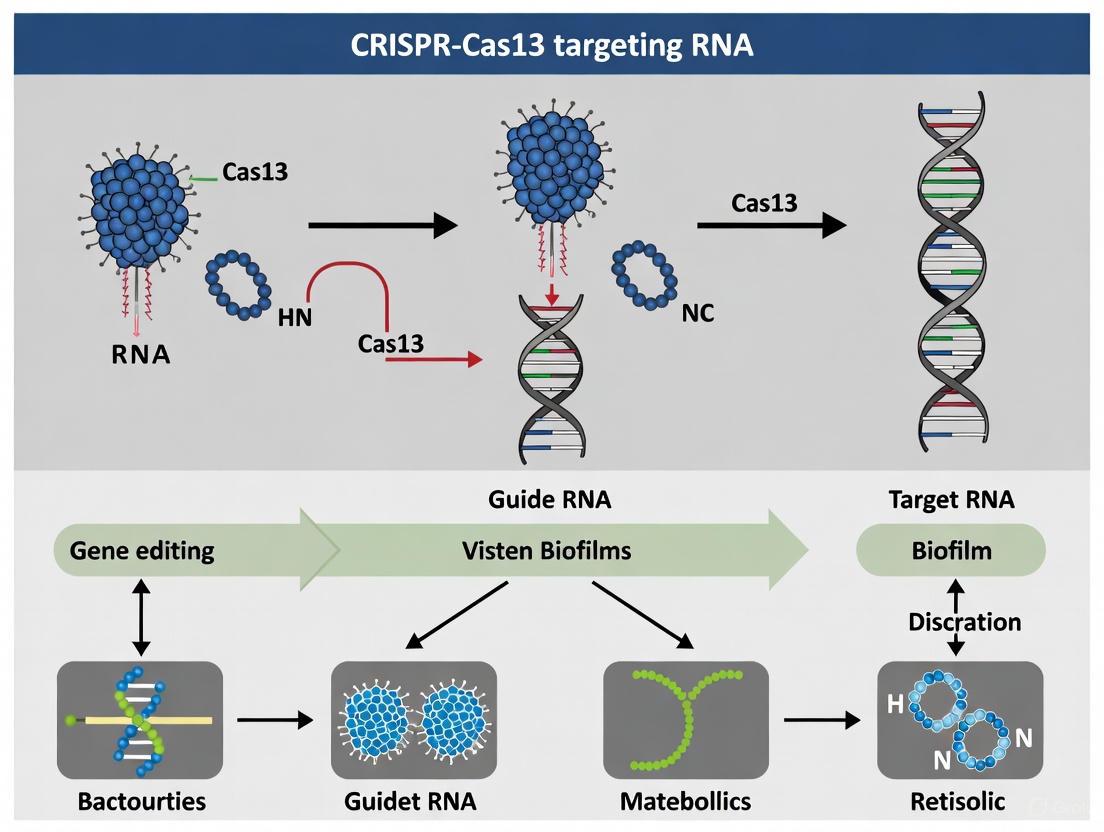
This article explores the transformative potential of the RNA-targeting CRISPR-Cas13 system as a precision tool for disrupting biofilm-associated metabolic pathways.
CRISPR-Cas13: Precision RNA Targeting for Disrupting Biofilm Metabolic Pathways
Abstract
This article explores the transformative potential of the RNA-targeting CRISPR-Cas13 system as a precision tool for disrupting biofilm-associated metabolic pathways. We provide a foundational overview of Cas13's unique mechanism as an RNA-guided ribonuclease, distinct from DNA-targeting CRISPR systems. The content details methodological strategies for applying Cas13 to silence key genes involved in quorum sensing, extracellular polymeric substance (EPS) production, and stress response in bacterial biofilms. We critically evaluate optimization techniques to enhance delivery and efficacy, including nanoparticle carriers and guide RNA modifications, while addressing challenges such as collateral activity and bacterial delivery. A comparative analysis validates Cas13's performance against other RNA-targeting methods like RNAi, highlighting its superior specificity and applicability for developing next-generation, sequence-specific antimicrobials to combat resistant, biofilm-based infections.
The RNA-Guided Ribonuclease: Understanding CRISPR-Cas13's Mechanism for Biofilm Intervention
The CRISPR-Cas (Clustered Regularly Interspaced Short Palindromic Repeats and CRISPR-associated proteins) adaptive immune system in prokaryotes has yielded revolutionary tools for precision molecular biology. Among these, Cas9 and Cas13 represent distinct classes of programmable nucleases with fundamentally different targeting preferences and mechanistic principles [1]. Cas9 is an RNA-guided DNA endonuclease that has become the cornerstone of genome engineering, while Cas13 is an RNA-guided RNase that enables precise manipulation of transcriptomes [2] [3]. Understanding their differences is crucial for selecting the appropriate tool for specific research applications, particularly in emerging fields such as biofilm metabolic pathway research where precise temporal control over gene expression is often required.
This application note details the fundamental distinctions between these two systems, providing structured comparisons, experimental protocols, and specific guidance for their application in studying bacterial biofilm metabolism.
Fundamental Mechanisms and Structural Comparisons
Core Functional Distinctions
The most fundamental distinction lies in their substrate specificity: Cas9 targets DNA, while Cas13 targets RNA [4] [3]. This difference dictates their cellular roles, applications, and safety profiles.
Cas9 Mechanism: Cas9 creates double-strand breaks (DSBs) in DNA target sequences [5]. After forming an R-loop with the target DNA, its HNH nuclease domain cleaves the complementary strand, while the RuvC-like domain cleaves the non-target strand, resulting primarily in blunt-ended breaks [1]. This activity permanently alters the genetic code, making it ideal for stable gene knockout strategies.
Cas13 Mechanism: Cas13 exhibits single-stranded RNA (ssRNA) cleavage activity via two conserved Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains [6] [2]. Upon recognition and binding to its target RNA sequence, Cas13 becomes activated and cleaves both the target RNA (cis-cleavage) and surrounding non-specific RNA molecules (trans- or collateral cleavage) [6] [7]. This activity enables transient modulation of gene expression without permanent genomic changes.
Table 1: Fundamental Characteristics of Cas9 and Cas13 Systems
| Feature | Cas9 | Cas13 |
|---|---|---|
| Primary Target | Double-stranded DNA | Single-stranded RNA |
| Nuclease Domains | HNH, RuvC-like | Two HEPN domains |
| Cleavage Products | Blunt-ended double-strand breaks (Cas9) or staggered ends (Cas12a) | Fragmented RNA molecules |
| Collateral Activity | No | Yes - non-specific RNase upon activation |
| Natural Biological Role | Adaptive immunity against DNA phages & plasmids | Adaptive immunity against RNA phages |
| Representative Subtypes | SpCas9, SaCas9 | Cas13a, Cas13b, Cas13d, Cas13X/Y |
Molecular Architecture and Guide RNA Requirements
Structurally, both proteins form ribonucleoprotein complexes but differ significantly in their architecture and guide RNA requirements.
Cas9 exhibits a bilobed architecture consisting of recognition (REC) and nuclease (NUC) lobes, recognizing a protospacer adjacent motif (PAM) in the target DNA (typically 5'-NGG-3' for SpCas9) [5] [1]. It uses a single guide RNA (sgRNA) engineered by fusing CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA) [1].
Cas13 also possesses a bilobed architecture with REC and NUC lobes, but its NUC lobe contains the HEPN domains responsible for RNA cleavage [6] [7]. Cas13 recognizes target sequences flanked by protospacer flanking sites (PFS), rather than PAM sequences, with preferences varying by subtype (e.g., Cas13a prefers 3' A, U, or C motifs) [3]. It typically requires only a crRNA for guidance, without the need for tracrRNA [6].
Quantitative Comparison of Molecular Properties
Table 2: Comparative Molecular Properties for Experimental Design
| Property | Cas9 | Cas13 | Experimental Implication |
|---|---|---|---|
| Protein Size | ~1000-1600 amino acids | ~800-1200 amino acids (Cas13bt3: ~800 aa) | AAV packaging capacity favors smaller variants like Cas13bt3, Cas13X/Y [7] |
| Editing Permanence | Permanent genomic changes | Transient, reversible effects | Cas9 for stable knockouts, Cas13 for transient knockdowns or dynamic studies |
| Off-Target Concerns | DNA off-target edits at similar sites | RNA off-target effects via collateral cleavage | Cas13 collateral activity requires engineered variants for precise work [7] |
| Target Specificity | Sensitive to single-nucleotide mismatches | Tolerates some mismatches, especially at 5' end | Cas9 better for SNP discrimination; Cas13 more flexible in mismatch tolerance |
| Multiplexing Capacity | Limited by multiple gRNA expression | High due to minimal crRNA structure | Cas13 superior for targeting multiple transcripts simultaneously |
| Therapeutic Delivery | Challenges with AAV packaging | More compact variants available for AAV | Cas13 advantageous for viral delivery approaches [4] [7] |
Application to Biofilm Metabolic Pathways Research
The study of biofilm metabolic pathways presents unique challenges that can be addressed by strategic selection of CRISPR tools. Biofilms are structured microbial communities encased in extracellular polymeric substances, exhibiting complex metabolic networks and gradients of metabolic activity that change throughout biofilm development [8].
Cas9 Applications in biofilm research include:
- Creating stable knockout mutants of key metabolic enzymes to study their necessity in biofilm formation and maintenance.
- Introducing point mutations in regulatory genes to dissect metabolic regulation networks.
- Generating reporter strains by inserting fluorescent protein genes downstream of metabolic promoters.
Cas13 Applications are particularly valuable for:
- Transient knockdown of essential metabolic genes without permanent genetic alterations, allowing study of genes essential for viability.
- Multiplexed targeting of multiple metabolic pathway components simultaneously to understand network redundancy and robustness.
- Dynamic studies of metabolic shifts during biofilm development from attachment to maturation and dispersal.
- Precise modulation of quorum sensing signaling molecules that regulate metabolic cooperation in biofilms.
Experimental Protocols
Protocol 1: Cas9-Mediated Gene Knockout in Biofilm-Forming Bacteria
Objective: To create stable gene knockouts in metabolic genes to assess their role in biofilm formation.
Materials:
- pCas9 plasmid system with inducible expression
- pTargetF plasmid for sgRNA expression or synthetic sgRNA
- HR donor template (for homology-directed repair if needed)
- Appropriate bacterial strain(s)
- Antibiotics for selection
- Biofilm culturing vessels (e.g., microtiter plates, flow cells)
- Crystal violet or other biofilm staining materials
Procedure:
- sgRNA Design: Design sgRNAs targeting the metabolic gene of interest using computational tools (e.g., CRISPOR). Select targets with high on-target and low off-target scores.
- Transformation: Introduce the Cas9 and sgRNA constructs into the target bacterium via electroporation or chemical transformation.
- Selection and Screening: Plate transformed cells on selective media. Screen individual colonies by colony PCR and sequencing to verify gene editing.
- Phenotypic Analysis:
- Grow biofilm cultures of wild-type and mutant strains under standardized conditions.
- Quantify biofilm biomass using crystal violet staining or similar method.
- Assess metabolic activity using assays such as ATP quantification or tetrazolium reduction.
- Analyze metabolic profiles via LC-MS or GC-MS if available.
Troubleshooting:
- Low editing efficiency: Optimize sgRNA design, use different PAM sites, or increase Cas9 expression.
- No viable colonies: Target gene may be essential; consider partial knockdown with Cas13 instead.
- Off-target effects: Validate key phenotypes with complementary approaches.
Protocol 2: Cas13-Mediated Transcriptional Knockdown for Metabolic Studies
Objective: To achieve transient knockdown of metabolic gene expression for studying essential pathways in biofilms.
Materials:
- Cas13d (CasRx) expression plasmid or purified protein
- crRNA expression plasmid or synthetic crRNA
- Appropriate delivery system (electroporation, nanoparticles, conjugative plasmids)
- qRT-PCR reagents for validation
- Metabolic activity assays (e.g., resazurin reduction, substrate utilization)
- RNA sequencing reagents (optional)
Procedure:
- crRNA Design: Design crRNAs targeting the mRNA of interest, focusing on accessible regions. Include multiple crRNAs for enhanced efficacy.
- Delivery:
- For plasmid-based delivery: Co-transform Cas13 and crRNA expression plasmids.
- For RNP delivery: Complex purified Cas13 protein with synthetic crRNA and deliver via electroporation.
- Knockdown Validation:
- Extract RNA 24-48 hours post-delivery.
- Perform qRT-PCR to quantify target mRNA reduction.
- Assess protein level reduction if antibodies are available.
- Metabolic Phenotyping:
- Monitor growth curves and substrate utilization patterns.
- Assess biofilm formation capacity under different nutrient conditions.
- Measure metabolic flux using labeled substrates if available.
- Multi-timepoint Analysis: Repeat analyses at 24-hour intervals to track duration of knockdown effects.
Troubleshooting:
- Low knockdown efficiency: Test multiple crRNAs targeting different regions of the transcript.
- Cellular toxicity: Use lower Cas13 expression levels or engineer collateral-deficient variants.
- Short knockdown duration: Consider stable integration or repeated delivery for longer studies.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for CRISPR-Cas Experiments in Biofilm Research
| Reagent Category | Specific Examples | Function & Application Notes |
|---|---|---|
| Cas Expression Plasmids | pCas9, pCas13d, compact variants (Cas13X/Y) | Source of nuclease activity; choose inducible systems for toxic targets |
| Guide RNA Cloning Systems | pTargetF, crRNA expression backbones | Customizable guides for specific targets; consider multiplexed arrays for Cas13 |
| Delivery Tools | Electroporators, lipid nanoparticles, conjugative plasmids | Introduction of CRISPR components; method depends on bacterial strain |
| Selection Markers | Antibiotic resistance genes, fluorescence reporters | Enrichment for successfully transformed/edited cells |
| Screening Reagents | PCR kits, sequencing primers, restriction enzymes | Validation of editing events or knockdown efficiency |
| Biofilm Assay Materials | Crystal violet, microtiter plates, flow cells, confocal imaging dishes | Standardized assessment of biofilm phenotypes |
| Metabolic Assays | Tetrazolium salts, ATP kits, pH sensors, substrate utilization panels | Functional assessment of metabolic consequences |
Strategic Selection Guide for Biofilm Research
Choosing between Cas9 and Cas13 requires careful consideration of the specific research question:
Select Cas9 when:
- Studying non-essential genes where permanent knockout is acceptable
- Creating stable mutant strains for long-term studies
- Precise DNA editing is required (point mutations, insertions)
- Working with well-characterized genetic systems
Select Cas13 when:
- Targeting essential genes that cannot be knocked out
- Studying metabolic transitions or dynamic processes
- Multiplexed targeting of pathway components is advantageous
- Transient effects are desirable to avoid compensatory adaptations
- Working with genetically intractable strains where stable transformation is difficult
For comprehensive biofilm metabolic studies, a combined approach often yields the deepest insights: using Cas9 to create stable mutations in regulatory elements and Cas13 for fine-tuning expression of metabolic enzymes to map flux distributions.
Visualization of CRISPR-Cas Mechanisms in Biofilm Context
This application note provides the foundational knowledge and practical methodologies required to effectively leverage both Cas9 and Cas13 systems in biofilm metabolic research. The complementary strengths of these technologies enable researchers to address complex biological questions about metabolic regulation in structured microbial communities with unprecedented precision.
CRISPR-Cas13 systems provide adaptive immunity in prokaryotes through RNA-guided RNA targeting. Distinct from DNA-targeting Cas9 and Cas12 proteins, Cas13 effectors are single-subunit RNA-guided ribonucleases (RNases) that recognize and cleave single-stranded RNA (ssRNA) targets. This unique mechanism centers on two key functional characteristics: RNA-guided target recognition and HEPN domain-mediated RNase activity. When deployed against biofilm-forming pathogens, this mechanism enables precise targeting of metabolic pathway mRNAs, offering a novel approach for disrupting biofilm integrity and persistence [6] [2].
The Cas13 effector complex undergoes significant conformational changes upon target RNA binding, activating its catalytic HEPN domains. This activation triggers not only sequence-specific cleavage of the target RNA but also collateral RNase activity that degrades non-specific nearby RNA molecules. This dual functionality makes Cas13 particularly valuable for both fundamental research on biofilm regulation and diagnostic applications for pathogen detection [6] [9].
Structural Organization and Functional Domains
Cas13 proteins share a conserved bilobed architecture consisting of Recognition (REC) and Nuclease (NUC) lobes:
- REC Lobe: Composed of N-Terminal (NTD) and Helical-1 domains, primarily responsible for crRNA recognition and binding.
- NUC Lobe: Contains Helical-2, Helical-3 (linker), and two higher eukaryotes and prokaryotes nucleotide-binding (HEPN) domains that facilitate target RNA accommodation and cleavage [6].
The HEPN domains, present in pairs within all Cas13 orthologs, contain characteristic RxxxxH motifs that form the catalytic core for RNase activity. These domains are conserved across diverse Cas13 subtypes including VI-A, VI-B, VI-C, and VI-D systems [6] [2].
Figure 1: Structural organization of Cas13 effector protein showing REC and NUC lobes with their functional domains. HEPN domains contain the catalytic RxxxxH motifs essential for RNase activity.
RNA-Guided Target Recognition and Cleavage Mechanism
crRNA Biogenesis and Effector Complex Formation
The Cas13 mechanism initiates with precursor CRISPR RNA (pre-crRNA) processing, where Cas13 itself cleaves within direct repeat sequences to generate mature crRNAs. In type VI-A systems, this processing generates a 2',3'-cyclic phosphate on the 5'-flank product and a 5'-OH on the mature crRNA [6].
Mature crRNAs remain bound to Cas13, forming the surveillance effector complex. The crRNA architecture consists of:
- 5'-handle: Stem-loop structure buried in the cleft between NTD and Helical-1 domains
- Spacer region: Binds complementary target RNA in sequence-specific manner
- 3'-flank: Stabilized by interactions with HEPN-2 domain [6]
Target Recognition and Cleavage Activation
Upon encountering complementary target RNA, the spacer region of crRNA forms base pairs, triggering conformational changes that activate Cas13's RNase capability. The catalytic mechanism involves:
- Dual HEPN Domain Coordination: Both HEPN domains contribute residues to form a single RNase active site
- Metal Ion Dependence: Target RNA cleavage requires Mg²⁺ ions, while pre-crRNA processing is Mg²⁺-independent
- Collateral Cleavage: Activated Cas13 non-specifically degrades nearby RNA molecules, providing amplification for diagnostic applications [6] [2]
Figure 2: Sequential mechanism of Cas13 RNA-guided target recognition and cleavage, showing progression from effector complex formation to collateral RNA degradation.
Quantitative Analysis of Cas13 Subtypes and Characteristics
Table 1: Comparison of Cas13 subtypes and their functional characteristics relevant to biofilm research
| Subtype | Size (aa) | crRNA Features | Processing Activity | Applications in Biofilm Research |
|---|---|---|---|---|
| VI-A (Cas13a) | 1000-1200 | Spacer at 3' end, requires 5' flank processing | Mg²⁺-independent pre-crRNA processing | Gene knockdown in biofilm metabolic pathways |
| VI-B (Cas13b) | 1000-1200 | Spacer at 5' end, distinct direct repeats | Variable processing mechanisms | Multiplexed targeting of quorum-sensing genes |
| VI-C (Cas13c) | 1000-1200 | Conserved stem-loop architecture | Dependent on conserved catalytic residues | Pathogen detection in biofilm samples |
| VI-D (Cas13d) | 775-800 | Compact guide architecture | High-fidelity processing | Engineering of probiotic anti-biofilm strains |
| Cas13X/Y | 775-800 | Minimal structural requirements | Efficient processing in compact form | Nanoparticle delivery for biofilm penetration |
Table 2: HEPN domain catalytic residues across Cas13 subtypes
| Cas13 Subtype | HEPN-1 Motif | HEPN-2 Motif | Catalytic Mechanism | Metal Ion Requirement |
|---|---|---|---|---|
| VI-A (LshCas13a) | R...H | R...H | General acid-base catalysis | Mg²⁺-dependent for target RNA |
| VI-B (PguCas13b) | R...H | R...H | Similar RNase mechanism | Mg²⁺-dependent for target RNA |
| VI-C | R...H | R...H | Conserved catalytic core | Mg²⁺-dependent for target RNA |
| VI-D (RfxCas13d) | R...H | R...H | Compact active site | Mg²⁺-dependent for target RNA |
| Cas13e | R...H | R...H | Intermediate evolutionary traits | Mg²⁺-dependent for target RNA |
Research Reagent Solutions for Biofilm Metabolic Pathway Targeting
Table 3: Essential research reagents for implementing CRISPR-Cas13 in biofilm metabolic pathways research
| Reagent Category | Specific Examples | Function in Experiment | Biofilm Research Application |
|---|---|---|---|
| Cas13 Effectors | LwaCas13a, PspCas13b, RfxCas13d | RNA-guided RNase execution | Target mRNA degradation in biofilm matrix |
| crRNA Design Tools | CRISPR-RT, Cas13design | Spacer sequence optimization | Targeting quorum-sensing transcripts |
| Delivery Systems | Liposomal nanoparticles, Gold nanoparticle carriers | Cellular delivery of RNP complexes | Enhanced biofilm penetration [11] |
| Detection Reporters | Fluorescent RNA reporters, Quenched RNA substrates | Monitoring collateral activity | Real-time detection of pathogen viability |
| Control Elements | Catalytically dead Cas13 (dCas13), Scrambled crRNAs | Specificity validation | Differentiation from antibiotic effects |
| Expression Systems | T7 polymerase-based vectors, Constitutive promoters | Heterologous Cas13 expression | Engineering probiotic anti-biofilm strains |
Experimental Protocol: Targeting Biofilm Metabolic Pathways with CRISPR-Cas13
crRNA Design and Synthesis for Metabolic Gene Knockdown
Principle: Design crRNAs complementary to mRNA targets encoding essential biofilm metabolic enzymes, quorum-sensing regulators, or stress response factors.
Procedure:
- Target Selection: Identify accessible regions within target mRNAs using RNA accessibility prediction tools (e.g., CRISPR-RT)
- Spacer Design: Design 28-30 nt spacers with complementary to target sequence
- Direct Repeat Addition: Append appropriate direct repeat sequence for your Cas13 subtype (e.g., 5'-GAUUUAGACUACCCCAAAAACGAAGGGGACUAAAAC-3' for LwaCas13a)
- Specificity Verification: BLAST spacer against host genome to minimize off-target effects
- Synthesis: Chemically synthesize crRNA using commercial RNA synthesis services
Critical Parameters:
- Avoid stretches of ≥4 identical nucleotides
- Ensure G/C content between 40-60%
- Verify absence of self-complementarity that could impair Cas13 binding
Cas13-crRNA Ribonucleoprotein (RNP) Complex Assembly
Reagents:
- Purified recombinant Cas13 protein (commercial sources available)
- Synthetic crRNA (resuspended in nuclease-free water)
- Assembly buffer (20 mM HEPES pH 7.5, 150 mM KCl, 1 mM DTT, 5% glycerol)
Procedure:
- Denaturation: Heat crRNA to 65°C for 5 minutes then immediately place on ice
- Complex Formation: Mix Cas13 protein and crRNA in 1:1.2 molar ratio in assembly buffer
- Incubation: Incubate mixture at 25°C for 30 minutes to allow RNP formation
- Quality Assessment: Analyze complex formation using native PAGE or EMSA
- Storage: Aliquot and store at -80°C for up to 3 months
Delivery to Biofilm-Forming Microbes Using Nanoparticle Carriers
Principle: Lipid-based nanoparticles enhance penetration through biofilm extracellular polymeric substances (EPS) and improve cellular uptake.
Procedure:
- Nanoparticle Preparation: Formulate RNP complexes with cationic lipids (e.g., DOTAP, DOPE) using microfluidics-based mixing
- Characterization: Measure particle size (target 80-120 nm) and zeta potential (>+20 mV) using dynamic light scattering
- Biofilm Treatment: Apply nanoparticle formulation to mature biofilms (typically 72-hour growth)
- Incubation: Allow 4-6 hours for nanoparticle penetration and cellular uptake
- Assessment: Evaluate gene knockdown efficiency using RT-qPCR and functional assays
Optimization Notes:
- Liposomal Cas13 formulations demonstrated >90% reduction in P. aeruginosa biofilm biomass [11]
- Gold nanoparticle carriers showed 3.5-fold increase in editing efficiency compared to non-carrier systems [11]
Validation of Target Knockdown and Phenotypic Effects
Molecular Validation:
- RNA Extraction: Harvest biofilm cells 24-48 hours post-treatment using mechanical disruption and RNA stabilization
- RT-qPCR Analysis: Quantify target mRNA levels relative to housekeeping genes
- Collateral Activity Assessment: Monitor non-target transcript levels to evaluate specificity
Functional Assays:
- Biofilm Biomass Quantification: Crystal violet staining or confocal microscopy analysis
- Metabolic Activity: Resazurin reduction assays or ATP quantification
- Antibiotic Sensitivity Testing: Evaluate synergy between Cas13 treatment and conventional antibiotics
Application Notes for Biofilm Metabolic Pathway Research
When applying CRISPR-Cas13 to investigate biofilm metabolic pathways, consider these critical factors:
Delivery Optimization: Biofilm EPS matrices significantly impede macromolecular delivery. Nanoparticle formulations must be optimized for size, surface charge, and stability to achieve effective penetration. Combining Cas13 RNPs with EPS-degrading enzymes (e.g., DNase I, dispersin B) can enhance delivery efficiency.
Target Selection Strategy: Prioritize mRNAs encoding:
- Quorum-sensing master regulators (e.g., lasR, rhIR in P. aeruginosa)
- Central carbon metabolism enzymes
- Stress response transcription factors
- Efflux pump components
- Polysaccharide synthesis enzymes
Temporal Considerations: Metabolic pathway disruption requires careful timing relative to biofilm developmental stage. Interventions during early attachment phase may prevent maturation, while targeting established biofilms requires combination approaches.
Specificity Controls: Always include:
- Catalytically dead Cas13 (dCas13) with same crRNA
- Non-targeting crRNA controls
- Multiple crRNAs against same target gene
- Off-target transcript monitoring
The RNA-guided cleavage and HEPN domain RNase activity of CRISPR-Cas13 provides a powerful, programmable tool for dissecting metabolic pathways in biofilm research. By enabling precise degradation of target mRNAs, this technology facilitates functional genomics studies and potential therapeutic interventions against persistent biofilm-associated infections.
The CRISPR-Cas13 system represents a groundbreaking RNA-targeting platform derived from bacterial adaptive immune systems. As a programmable RNA-guided RNase, Cas13 has emerged as a powerful tool for transcriptome engineering, enabling precise knockdown of targeted RNA molecules. In the context of biofilm research, where conventional antimicrobial therapies often fail against the protective extracellular polymeric substance (EPS) matrix, Cas13 technology offers unprecedented opportunities for precision intervention in biofilm metabolic pathways [8] [12]. Unlike DNA-targeting CRISPR systems, Cas13 operates at the transcriptional level, allowing transient and reversible modulation of gene expression without permanent genomic changes—a particularly valuable characteristic for studying essential metabolic pathways in biofilm communities [2].
Cas13 proteins share fundamental characteristics, including two conserved Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains that form the catalytic site for RNA cleavage. Upon recognition of its target RNA through complementary CRISPR RNA (crRNA), Cas13 undergoes conformational changes that activate its RNase capability, cleaving both the target RNA and exhibiting collateral activity against nearby non-target RNAs [13] [14]. However, despite these shared mechanisms, different Cas13 orthologs exhibit substantial variation in size, crRNA requirements, efficiency, and specificity, making the selection of appropriate orthologs crucial for specific research applications, particularly in complex biofilm environments [15] [2].
Comparative Analysis of Cas13 Orthologs
Structural and Functional Characteristics
The diversity of Cas13 orthologs stems from their evolutionary adaptation across different bacterial species. Each ortholog possesses distinct structural features that influence its functionality and applicability in biofilm research.
Cas13a was the first characterized subtype of the Cas13 family, with LwaCas13a from Leptotrichia wadei demonstrating robust RNA-targeting activity [15] [13]. This ortholog typically requires a protospacer flanking site (PFS) with a preference for adenine or uracil bases adjacent to the target sequence. With over 1,000 amino acids, Cas13a is one of the larger orthologs and contains HEPN domains positioned at both the center and C-terminus of its linear structure [15] [14]. Its crRNA architecture features a 28nt direct repeat (DR) with a 5' handle and a 3' spacer, which influences its targeting specificity.
Cas13d, particularly RfxCas13d from Ruminococcus flavefaciens, has gained significant research attention due to its compact size (approximately 190-300 amino acids) and high efficiency in eukaryotic cells [15] [16]. Unlike earlier Cas13 nucleases, RfxCas13d does not impose strict PFS constraints, providing greater targeting flexibility [15]. Structurally, Cas13d's HEPN domains are located at both the center and C-terminus, similar to Cas13a, but its minimal size facilitates easier delivery—a significant advantage for biofilm applications where penetration through EPS matrices is challenging [8] [16].
Cas13x and Cas13y represent more recently discovered subtypes characterized by their exceptionally compact sizes. Cas13x.1 is approximately 200 amino acids smaller than RfxCas13d, making it one of the most miniature Cas13 variants identified [15] [16]. These orthologs feature HEPN domains situated at the extreme N-terminus and C-terminus of the linear protein, with DR sequences positioned at the 5' end in an orientation contrasting with other subtypes [15]. Their minimal size enables more straightforward packaging into delivery vectors, offering significant potential for biofilm therapeutic applications where size constraints are critical.
Performance Comparison in Biological Systems
Recent systematic evaluations of Cas13 orthologs in plant systems provide valuable quantitative insights into their relative performances, with implications for biofilm research applications. The table below summarizes the key performance metrics of various Cas13 orthologs based on empirical studies:
Table 1: Comparative Performance of Cas13 Orthologs
| Ortholog | Subtype | Size (aa) | Editing Efficiency | Key Characteristics | Biofilm Research Suitability |
|---|---|---|---|---|---|
| LwaCas13a | VI-A | ~1000-1200 | Moderate | Requires PFS preference; first characterized | Moderate - larger size may limit delivery |
| PbuCas13b | VI-B | ~1000-1200 | High | Superior to LwaCas13a in mammalian cells | High - efficient but delivery challenging |
| RfxCas13d | VI-D | 190-300 | 58-80% | No PFS constraints; versatile | Very High - compact size enhances delivery |
| Cas13x.1 | VI-X | ~775-800 | 58-80% | Extremely compact; enhanced stability | Excellent - minimal size optimal for delivery |
| Cas13x.2 | VI-X | ~775-800 | 58-80% | Extremely compact; enhanced stability | Excellent - minimal size optimal for delivery |
| Cas13y.1 | VI-Y | ~790 | 58-80% | Comparable efficiency to Cas13d | Excellent - balances size and efficiency |
| Cas13y.2 | VI-Y | ~790 | 58-80% | Comparable efficiency to Cas13d | Excellent - balances size and efficiency |
A comprehensive study systematically evaluating seven Cas13 orthologs from five distinct subtypes revealed that RfxCas13d, Cas13x.1, and Cas13x.2 exhibit enhanced stability with editing efficiencies ranging from 58% to 80%, closely followed by Cas13y.1 and Cas13y.2 at similar efficiency levels [15]. Notably, both Cas13x.1 and Cas13y.1 demonstrated the ability to simultaneously degrade two endogenous transcripts using a tRNA-crRNA cassette approach, achieving editing efficiencies of up to 50%—a particularly valuable feature for targeting multiple components of biofilm metabolic pathways simultaneously [15].
The compact size of Cas13x and Cas13y orthologs provides significant advantages for delivery in biofilm environments. Their minimal dimensions enable more efficient packaging into nanoparticle delivery systems, enhancing penetration through the protective EPS matrix of biofilms [12] [16]. Furthermore, studies indicate that these newer orthologs generate minimal off-target effects, making them particularly suitable for precise manipulation of biofilm metabolic pathways without disrupting essential cellular functions [15].
Experimental Protocols for Biofilm Metabolic Pathway Targeting
crRNA Design and Validation for Metabolic Genes
Effective targeting of biofilm metabolic pathways begins with meticulous crRNA design against key enzymes or regulators. The following protocol outlines a systematic approach for crRNA design and validation:
Target Selection: Identify critical genes in biofilm metabolic pathways, such as those involved in quorum sensing (e.g., luxS), EPS production (e.g., pel, psl operons), or central carbon metabolism (e.g., ackA, ldh). Prioritize accessible regions of the target mRNA by predicting secondary structure using tools like RNAfold [17].
Spacer Design: Design 20-30nt spacers complementary to your target region. For Cas13d, use 23nt spacers; for Cas13x/Cas13y, 21-22nt spacers are optimal. Avoid stretches of ≥4 identical nucleotides and ensure GC content between 40-60% [15] [17].
Specificity Verification: BLAST spacer sequences against the host genome to minimize off-target effects. For single-nucleotide specificity, incorporate synthetic mismatches in the crRNA or extend the 3' end with a hairpin structure to enhance discrimination [13] [17].
crRNA Construction: Clone spacer sequences into appropriate expression vectors containing direct repeat sequences. For multiplexing, utilize tRNA-crRNA cassettes enabling simultaneous targeting of multiple pathway components [15].
Validation: Test crRNA efficiency by transferring Cas13-crRNA constructs into model systems and quantifying target RNA knockdown using RT-qPCR. Successful crRNAs typically achieve >60% knockdown of target transcripts [15].
Delivery Optimization for Biofilm Environments
Efficient delivery of CRISPR-Cas13 components through biofilm matrices presents unique challenges. The following protocol outlines strategies for optimizing delivery in biofilm systems:
Nanoparticle Formulation: Encapsulate Cas13 ribonucleoproteins (RNPs) or expression plasmids in lipid nanoparticles (LNPs) or gold nanoparticles (AuNPs). For LNPs, use ionizable lipids (e.g., DLin-MC3-DMA) with cholesterol, DSPC, and PEG-lipid at molar ratios of 50:38.5:10:1.5 [12].
Surface Functionalization: Enhance biofilm penetration by functionalizing nanoparticles with biofilm-penetrating peptides (e.g., KHKKHKHKHKHKHKHKHKKH-K-ε-ahx-K-LL-37). Incubate nanoparticles with 100μM peptide in PBS for 1 hour at room temperature with gentle agitation [8] [12].
Delivery Parameters: For established biofilms, apply nanoparticle formulations at 100μg/mL total lipid concentration in fresh medium. Incubate for 4-6 hours at 37°C with gentle shaking [12].
Efficiency Assessment: Measure Cas13 expression and target knockdown using fluorescence microscopy (if using fluorescent tags) and RT-qPCR. Successful delivery should achieve >50% target RNA reduction without significant cytotoxicity [15] [12].
Diagram 1: crRNA Design Workflow for Biofilm Metabolic Gene Targeting
Visualization of Cas13 Mechanisms and Workflows
Cas13 RNA Targeting Mechanism in Biofilm Cells
The molecular mechanism of Cas13-mediated RNA targeting involves a sophisticated sequence recognition and activation process. The following diagram illustrates this mechanism within the context of a biofilm-associated bacterial cell:
Diagram 2: Cas13 RNA Targeting Mechanism in Biofilm Cells
Multiplexed Targeting of Biofilm Pathways
Simultaneous targeting of multiple biofilm metabolic pathway components represents a powerful application of Cas13 technology. The following workflow illustrates the process for implementing multiplexed targeting:
Diagram 3: Workflow for Multiplexed Targeting of Biofilm Pathways
Research Reagent Solutions for Cas13 Applications
The successful implementation of Cas13-based approaches in biofilm research requires specific reagent systems optimized for this application. The following table outlines essential research reagents and their functions:
Table 2: Essential Research Reagents for Cas13 Biofilm Studies
| Reagent Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Cas13 Expression Plasmids | pC013-RfxCas13d, pC013-Cas13x.1 | Expresses Cas13 ortholog in target cells | Select based on size constraints; Cas13x.1 for stringent size limitations |
| crRNA Cloning Vectors | pC013-sgRNA, ptRNA-crRNA | Expresses crRNA guides | tRNA-crRNA vectors enable multiplexed targeting |
| Nanoparticle Delivery Systems | Lipid Nanoparticles (LNPs), Gold Nanoparticles (AuNPs) | Enhances delivery through biofilm matrix | Functionalize with biofilm-penetrating peptides |
| Fluorescent Reporters | BROKEN GREEN, MS2-MCP tagging systems | Visualizes delivery and target engagement | Enables quantification of efficiency |
| Target Validation Tools | RT-qPCR primers, RNA-seq libraries | Confirms target knockdown | Essential for validating metabolic pathway disruption |
| Biofilm Assessment Kits | Crystal violet, SYTO 9/propidium iodide | Quantifies biofilm biomass and viability | Correlates gene knockdown with phenotypic effects |
These reagent systems provide the foundational tools for implementing Cas13-based approaches in biofilm metabolic pathway research. When selecting Cas13 expression systems, consider the specific ortholog characteristics outlined in Table 1, balancing size constraints with efficiency requirements [15] [16]. For delivery systems, prioritize functionalized nanoparticles that enhance penetration through the complex EPS matrix of biofilms [8] [12]. Validation approaches should include both molecular confirmation of target knockdown and phenotypic assessment of biofilm metabolic consequences.
The continuing diversification of Cas13 orthologs presents expanding opportunities for precision targeting of biofilm metabolic pathways. The compact dimensions and high efficiency of more recently characterized orthologs like Cas13x and Cas13y, combined with advanced delivery strategies, offer powerful approaches for intervening in biofilm resilience mechanisms at the transcriptional level. As these tools continue to evolve, they promise to significantly advance our ability to precisely manipulate biofilm physiology for both basic research and therapeutic applications.
Biofilms are structured microbial communities embedded in extracellular polymeric substances (EPS) that pose a significant threat in clinical and industrial settings due to their extreme tolerance to antimicrobial treatments [8]. The metabolic pathways within these biofilms are tightly regulated by complex RNA-based networks, making them prime targets for precision disruption. Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-Cas13 systems have emerged as transformative tools for targeting these RNA networks, offering programmable, sequence-specific degradation of target transcripts without permanent genomic alterations [8] [18]. This Application Note provides detailed protocols for identifying key RNA targets within biofilm metabolic pathways and implementing CRISPR-Cas13 for their precise disruption, framed within the broader thesis of advancing RNA-targeting antimicrobial strategies.
Key RNA Targets in Biofilm Metabolic Pathways
Critical metabolic functions essential for biofilm formation and maintenance are orchestrated by specific mRNA transcripts. Disruption of these key RNAs can effectively compromise biofilm integrity. The table below summarizes primary RNA targets, their associated pathways, and the phenotypic consequences of their disruption.
Table 1: Key RNA Targets for Biofilm Disruption
| Target mRNA | Encoded Protein/Function | Associated Biofilm Pathway | Disruption Phenotype |
|---|---|---|---|
| epsA-E operon mRNA | Exopolysaccharide (EPS) biosynthesis enzymes | EPS Matrix Production | Inhibited biofilm maturation, weakened structural integrity [8] |
| luxS mRNA | Autoinducer-2 synthesis enzyme | Quorum Sensing | Disrupted cell-cell communication, reduced virulence and coordination [8] |
| csgA mRNA | Curli fiber major subunit (E. coli) | Adhesion & Initial Attachment | Impaired surface colonization, reduced biofilm biomass [8] |
| algC mRNA | Alginate biosynthesis enzyme (P. aeruginosa) | EPS Production & Stress Response | Increased antibiotic susceptibility, defective matrix formation [8] |
| icdA mRNA | Isocitrate dehydrogenase | Central Carbon Metabolism (TCA Cycle) | Reduced metabolic activity, increased persister cell susceptibility [11] |
Experimental Protocol: CRISPR-Cas13-Mediated RNA Targeting
This section provides a step-by-step methodology for designing and implementing a CRISPR-Cas13 system to disrupt a selected RNA target, using the epsA mRNA as a model.
Protocol 1: crRNA Design and In Vitro Transcription
Objective: To design and synthesize target-specific crRNAs for Cas13a (e.g., from Leptotrichia wadei).
Materials:
- Template DNA oligonucleotides (with T7 promoter sequence and crRNA scaffold)
- T7 RNA Polymerase Kit
- DNase I (RNase-free)
- RNA Cleanup Kit
Methodology:
- Target Selection: Identify a 22-28 nt target sequence within the epsA mRNA transcript using the rule: 3' of a protospacer flanking sequence (PFS) for LwCas13a, typically an "A" or "U" nucleotide.
- crRNA Template Design: Design a DNA oligonucleotide template as follows:
5'-TAATACGACTCACTATA-GGG-[YOUR 22-28 NT TARGET SEQUENCE]-GTTTAAGAGCTAATGCTGGAAAAC-3'Where the T7 promoter is underlined, and the standard LwCas13a crRNA direct repeat is in bold. - In Vitro Transcription (IVT):
- Set up the IVT reaction using the T7 RNA Polymerase Kit as per manufacturer's instructions.
- Incubate at 37°C for 4-6 hours.
- DNase Treatment and Purification:
- Add 1 µL of DNase I to the reaction and incubate for 15 minutes at 37°C.
- Purify the crRNA using an RNA Cleanup Kit. Elute in nuclease-free water.
- Quantify the crRNA concentration using a spectrophotometer and validate integrity via denaturing urea-PAGE.
Protocol 2: Delivery via Lipid Nanoparticles and Biofilm Assay
Objective: To package the Cas13a ribonucleoprotein (RNP) complex and evaluate its efficacy against a pre-formed biofilm.
Materials:
- Recombinant LwCas13a protein
- Cationic lipid nanoparticles (LNPs) [11]
- 96-well polystyrene microtiter plates
- SYTO 9 and propidium iodide (for live/dead staining)
- Crystal violet stain
Methodology:
- RNP Complex Formation: Pre-complex the purified LwCas13a protein (50 nM final concentration) with the in vitro-transcribed epsA-targeting crRNA (75 nM final concentration) in nuclease-free buffer. Incubate at 25°C for 15 minutes.
- Nanoparticle Formulation:
- Formulate the RNP complex into cationic LNPs according to the manufacturer's protocol. This encapsulates the RNP to facilitate delivery into bacterial cells within the biofilm [11].
- A negative control RNP complex with a non-targeting crRNA must be prepared in parallel.
- Biofilm Treatment and Analysis:
- Grow a Staphylococcus epidermidis biofilm in a 96-well plate for 24-48 hours.
- Carefully remove the spent medium and add 100 µL of the LNP-RNP formulation to the pre-formed biofilm. Incubate for 24 hours.
- Viability Assessment: Stain the biofilm with SYTO 9/propidium iodide and visualize via confocal laser scanning microscopy (CLSM) to assess bacterial cell viability.
- Biomass Quantification: Fix parallel biofilm samples with methanol, stain with 0.1% crystal violet for 15 minutes, wash, solubilize the dye in 30% acetic acid, and measure the absorbance at 595 nm.
Workflow Visualization
Diagram 1: CRISPR-Cas13 Biofilm Targeting Workflow.
Diagram 2: Key RNA Targets in Biofilm Metabolic Pathways.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CRISPR-Cas13 Biofilm Experiments
| Reagent / Material | Function / Role | Example Specification / Notes |
|---|---|---|
| Recombinant LwCas13a Protein | RNA-targeting effector nuclease; binds crRNA and cleaves target mRNA. | ≥90% purity, nuclease-free storage buffer. Commercial source or purified in-house from E. coli. |
| crRNA In Vitro Transcription Kit | Synthesis of target-specific guide RNAs. | Must include T7 RNA Polymerase, NTPs, and RNase inhibitors. |
| Cationic Lipid Nanoparticles (LNPs) | Delivery vector for RNP complexes into biofilm-embedded bacteria. | Enables efficient penetration of the EPS matrix [11]. |
| Confocal Laser Scanning Microscope (CLSM) | High-resolution 3D imaging of biofilm architecture and viability post-treatment. | Used with live/dead fluorescent stains (e.g., SYTO 9/propidium iodide). |
| 96-Well Polystyrene Microtiter Plates | Standardized substrate for growing biofilms for high-throughput assays. | Compatible with absorbance readers for crystal violet quantification. |
Advantages of RNA-Level Targeting for Transient and Reversible Metabolic Control
Targeting cellular processes at the RNA level represents a transformative approach for achieving transient and reversible metabolic control. Unlike DNA-level interventions that create permanent genetic alterations, RNA-targeting technologies enable precise, dose-dependent, and temporary modulation of gene expression. This is particularly advantageous for manipulating metabolic pathways in biofilm research, where fine-tuned, adaptive interventions are required to dissect complex metabolic networks without inducing irreversible phenotypic changes. The emergence of programmable RNA-targeting platforms, especially CRISPR-Cas13 systems, provides researchers with unprecedented tools for post-transcriptional gene regulation, offering unique benefits for metabolic pathway engineering and functional genomics studies in bacterial biofilms [2] [19].
RNA-level targeting operates through multiple mechanisms including transcript degradation, translation inhibition, and splicing modulation. This flexibility allows researchers to design interventions that range from complete gene knockdown to subtle fine-tuning of expression levels. For metabolic engineering applications, this precision enables the selective redirection of metabolic fluxes without compromising essential cellular functions, making it ideal for studying the dynamic metabolic adaptations that occur during biofilm development and maintenance [20] [21].
Technological Foundations: RNA-Targeting Platforms
Comparative Analysis of RNA-Targeting Modalities
Several technological platforms enable RNA-level interventions, each with distinct mechanisms and applications. The table below summarizes the key RNA-targeting modalities relevant to metabolic control in biofilm research:
Table 1: RNA-Targeting Platforms for Metabolic Control
| Technology | Mechanism of Action | Reversibility Profile | Key Advantages for Metabolic Studies |
|---|---|---|---|
| CRISPR-Cas13 | RNA-guided RNA cleavage via Cas endonuclease [2] [14] | Transient (hours to days) [19] | Programmable, multi-target capability, minimal off-target DNA effects |
| Antisense Oligonucleotides (ASOs) | Steric blocking or RNase H-mediated degradation [20] | Transient (hours to days) | Established chemistry, predictable pharmacokinetics |
| RNA Interference (RNAi) | Endogenous RISC-mediated degradation [2] | Transient (days) | High specificity, well-characterized delivery |
| RNA Editing (ADAR/APOBEC) | Nucleotide conversion (A-to-I, C-to-U) [22] [21] | Transient (dependent on transcript turnover) | Precise single-base modifications, endogenous enzyme recruitment |
CRISPR-Cas13 Systems: Structural and Functional Diversity
The CRISPR-Cas13 system represents a particularly versatile platform for RNA-level metabolic interventions. Cas13 proteins are single-component, RNA-guided RNases that target single-stranded RNA molecules through complementary CRISPR RNA (crRNA) spacers [14] [6]. Unlike DNA-targeting Cas9 systems, Cas13 operates exclusively at the RNA level, eliminating concerns about permanent genomic alterations while maintaining high programmability and specificity.
Cas13 effectors share a conserved bilobed architecture consisting of recognition (REC) and nuclease (NUC) lobes. The NUC lobe contains two Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains that confer RNase activity [14] [6]. Upon target RNA recognition and binding, these domains form a composite catalytic site that cleaves both the target RNA (cis-cleavage) and non-specific bystander RNA molecules (collateral cleavage) [6]. This collateral activity, while requiring careful experimental control, has been harnessed for highly sensitive nucleic acid detection applications.
Table 2: Cas13 Subtypes and Their Characteristics
| Cas13 Subtype | Size (aa) | Guide RNA Features | Notable Characteristics for Metabolic Studies |
|---|---|---|---|
| Cas13a (VI-A) | ~1000-1200 [2] | Spacer at 3′ end [2] | First characterized, robust activity |
| Cas13b (VI-B) | ~1000-1200 [2] | Spacer at 5′ end [2] | Compatible with additional regulatory proteins |
| Cas13d (VI-D) | ~900 [2] | Compact guide structure [2] | High efficiency, minimal size for delivery |
| Cas13X/Y | ~775-800 [2] [14] | Optimized minimal guides | Ultra-compact size, reduced immunogenicity |
Experimental Protocols for Metabolic Pathway Targeting
Protocol 1: CRISPR-Cas13-Mediated Gene Knockdown in Biofilm Models
Objective: Targeted knockdown of metabolic genes in established bacterial biofilms to assess pathway essentiality and metabolic flux redistribution.
Materials and Reagents:
- Cas13 Protein: Recombinant LbuCas13a or RfxCas13d (commercially available)
- crRNA Design: Target-specific 28-nt spacers with minimal off-target potential
- Delivery Vehicle: Cationic lipid nanoparticles (LNPs) optimized for biofilm penetration [20]
- Biofilm Culture: Established using flow-cell systems or microtiter plates
- Validation: RNA extraction kits, qRT-PCR reagents, metabolic profiling assays
Procedure:
- crRNA Design and Preparation:
- Identify target sequences within metabolic genes of interest (e.g., glycolysis, TCA cycle, or efflux pump components)
- Design spacers with 28-nt length complementary to accessible regions of target mRNA
- Avoid regions with extensive secondary structure that may impair Cas13 binding [17]
- Synthesize crRNAs with 5′ handle sequences appropriate for selected Cas13 ortholog
Ribonucleoprotein (RNP) Complex Formation:
- Combine 2μM purified Cas13 protein with 3μM crRNA in nuclease-free buffer
- Incubate at 37°C for 15 minutes to facilitate RNP complex formation
- Verify complex formation via native gel electrophoresis if necessary
Biofilm Treatment and Delivery:
- For mature biofilms (3-5 days development), replace growth medium with fresh medium containing RNP complexes
- For LNP-mediated delivery, encapsulate RNP complexes at nitrogen:phosphate ratio of 6:1
- Apply treatment at appropriate concentration (typically 100-500nM RNP final concentration)
- Incubate under standard growth conditions for 6-48 hours depending on desired knockdown duration
Efficacy Assessment:
- Harvest biofilm cells at designated timepoints (6, 12, 24, 48 hours)
- Extract total RNA and quantify target transcript levels via qRT-PCR
- Normalize to housekeeping genes and compare to non-targeting crRNA controls
- Perform metabolic profiling via LC-MS or extracellular flux analysis to quantify pathway alterations
Troubleshooting Notes:
- Low knockdown efficiency may indicate poor target accessibility; redesign crRNAs to different regions
- Cellular toxicity may result from excessive collateral activity; titrate RNP concentration downward
- Variable biofilm penetration can be addressed by extending treatment duration or optimizing LNP formulation
Protocol 2: Multiplexed Metabolic Pathway Engineering
Objective: Simultaneous targeting of multiple pathway components to redirect metabolic flux in biofilms.
Materials and Reagents:
- Multiplex crRNA Array: Designed with direct repeats separating target-specific spacers
- Cas13 Expression System: Plasmid-based or integrated Cas13d (RfxCas13d) for sustained expression
- Delivery Platform: Conjugative plasmid or engineered phage for bacterial systems
- Analysis Tools: RNA-seq library preparation kits, metabolic flux analysis software
Procedure:
- Multiplex crRNA Array Design:
- Select 3-5 target genes within the metabolic pathway of interest
- Design individual spacers with minimal cross-reactivity (BLAST against host genome)
- Assemble spacers separated by direct repeats into a single transcriptional unit
- Clone array into appropriate expression vector with polymerase III promoter
System Delivery and Expression:
- Transform constructs into biofilm-forming strains via electroporation or conjugation
- Induce Cas13 and crRNA array expression with sub-inhibitory concentrations of inducer (e.g., 0.1-0.5mM IPTG)
- Allow expression for 4-6 hours before biofilm formation assays
Validation of Multiplex Targeting:
- Assess transcript levels for all targets simultaneously via RNA-seq
- Quantify metabolic intermediates via targeted metabolomics
- Correlative analysis of transcript reduction and metabolic flux changes
Phenotypic Screening:
- Monitor biofilm formation dynamics under varying nutrient conditions
- Assess susceptibility to antimicrobial agents that target affected pathways
- Measure metabolic flexibility through nutrient shift experiments
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for RNA-Level Metabolic Targeting
| Reagent Category | Specific Examples | Function/Application | Considerations for Biofilm Studies |
|---|---|---|---|
| Cas13 Effectors | LwaCas13a, LbuCas13a, RfxCas13d [2] [6] | RNA-guided RNase activity | Cas13d shows highest efficiency in prokaryotic systems [2] |
| Delivery Systems | Lipid Nanoparticles (LNPs) [20], Electroporation, Conjugative Plasmids | Transport of RNPs or expression constructs | LNPs enhance biofilm penetration; conjugative plasmids enable strain-specific delivery |
| crRNA Design Tools | NUPACK [17], ADAPT [17] | Predict target accessibility and guide efficiency | Avoid regions with stable secondary structure (<-10 kcal/mol ΔG) [17] |
| Activity Reporters | Quenched fluorescent RNA (e.g., FAM-UUUU-BHQ1) [17] | Quantify Cas13 activation and cleavage efficiency | Validates RNP complex functionality before biofilm experiments |
| Metabolic Probes | Seahorse XF reagents, Stable isotope-labeled precursors | Monitor metabolic flux changes | Correlate transcript knockdown with functional metabolic outcomes |
| Validation Assays | qRT-PCR, RNA-seq, Targeted Metabolomics | Confirm target engagement and efficacy | Multi-omics approach captures systems-level metabolic adaptations |
Visualization of Mechanisms and Workflows
Cas13 RNA Targeting Mechanism
Experimental Workflow for Metabolic Gene Knockdown
Advantages for Metabolic Control in Biofilm Research
Temporal Control and Reversibility
RNA-level interventions offer unparalleled temporal control compared to DNA-level modifications. The transient nature of RNA knockdown enables researchers to apply metabolic perturbations at specific stages of biofilm development—from initial attachment to maturation and dispersion—and observe recovery upon intervention withdrawal. This dynamic control is essential for understanding metabolic plasticity in biofilms, where cells constantly adapt to nutrient availability, quorum signals, and microenvironmental gradients [21] [19].
The reversibility of RNA targeting also enables study designs that would be impossible with permanent genetic knockouts. Essential metabolic genes can be temporarily suppressed to assess their contribution to biofilm maintenance without selecting for compensatory mutations that often arise in serial passage experiments with deletion mutants. This temporal precision more closely mimics natural metabolic fluctuations and therapeutic interventions, providing more clinically relevant insights for antimicrobial development.
Multi-Target Screening and Pathway Analysis
CRISPR-Cas13 systems facilitate efficient multiplexing, enabling simultaneous targeting of multiple pathway components. This capability is particularly valuable for metabolic studies, where pathway redundancy and regulatory networks often complicate single-gene analyses. Researchers can design crRNA arrays that target several enzymes within a pathway to identify rate-limiting steps or target parallel pathways to dissect metabolic network interactions [2] [6].
The quantitative nature of RNA-level knockdown (as opposed to all-or-nothing knockout effects) enables researchers to establish dose-response relationships between gene expression levels and metabolic outputs. By titrating Cas13 expression or varying crRNA concentrations, researchers can achieve partial knockdowns that reveal non-linear relationships between enzyme abundance and metabolic flux, information critical for understanding metabolic control structures in biofilm communities.
Technical Practicality and Experimental Flexibility
From a practical perspective, RNA-targeting approaches offer several advantages for biofilm research. Cas13 systems function effectively in diverse bacterial species without requiring host-specific genetic tools. The single-protein nature of Class 2 CRISPR systems simplifies delivery compared to multi-component CRISPR systems, and the RNA-targeting function avoids the ethical and safety concerns associated with DNA modification in potential clinical applications [14] [19].
The rapid onset of RNA-level effects (typically within hours) enables higher-throughput screening compared to approaches requiring chromosomal integration or selection. This experimental flexibility allows researchers to quickly iterate through multiple target hypotheses and optimize intervention strategies. Furthermore, the programmability of CRISPR-Cas13 systems means that target sequences can be redesigned and validated much more rapidly than constructing new genetic knockouts, significantly accelerating the research timeline for metabolic pathway analysis in biofilms.
From Theory to Practice: Implementing Cas13 to Silence Critical Biofilm Pathways
Designing Guide RNAs (crRNAs) for Biofilm Gene Targets (e.g., Quorum Sensing, EPS Genes)
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) systems have evolved from bacterial adaptive immune mechanisms into versatile programmable molecular tools. The Class 2 Type VI CRISPR-Cas13 system is distinguished by its RNA-guided RNA-targeting capability, functioning as a programmable ribonuclease (RNase). Unlike DNA-targeting Cas9, Cas13 targets RNA transcripts, making it particularly suitable for modulating gene expression in metabolic pathways without altering the genome [13] [23]. This application note details the design and implementation of guide RNAs (crRNAs) for targeting key biofilm genes, enabling precise investigation of biofilm metabolic pathways.
Biofilms are structured microbial communities encased in a self-produced extracellular polymeric substance (EPS) matrix. This matrix, primarily composed of polysaccharides, proteins, and extracellular DNA, provides structural integrity and protection against environmental stresses and antimicrobial agents [12] [24]. Biofilm development is orchestrated by complex regulatory networks, including quorum sensing (QS) cell-to-cell communication and the intracellular second messenger cyclic di-GMP (c-di-GMP) [25]. Targeting the genes involved in these pathways with CRISPR-Cas13 allows for systematic dissection of their roles in biofilm formation, maturation, and dispersal.
The following diagram illustrates the core biofilm formation pathway and the key RNA targets for Cas13-crRNA intervention:
Key Biofilm Gene Targets and crRNA Design Principles
Critical Biofilm Regulatory Genes
Effective crRNA design begins with the selection of appropriate target genes within biofilm regulatory networks. The table below summarizes high-value targets across different functional categories, supported by experimental evidence from literature.
Table 1: Key Biofilm Gene Targets for crRNA Design
| Gene Target | Function in Biofilm | Biological Process | Evidence/Model System |
|---|---|---|---|
| luxS | Autoinducer-2 (AI-2) synthase | Quorum Sensing | CRISPRi knockdown reduced biofilm in E. coli [26] |
| gacA | Response regulator in two-component system | Biofilm Regulation, EPS Production | CRISPRi silencing altered biofilm architecture in P. fluorescens [25] |
| alg44 | Alginate copolymerase, c-di-GMP dependent | EPS Biosynthesis | Key polysaccharide production gene in Pseudomonas [25] |
| bifA | Phosphodiesterase degrades c-di-GMP | Biofilm Dispersion, Motility | CRISPRi phenocopied knockout in P. fluorescens [25] |
| gcbA | Diguanylate cyclase produces c-di-GMP | Biofilm Formation, Attachment | High c-di-GMP promotes sessile lifestyle [25] |
crRNA Design Parameters for Cas13
Cas13 crRNAs consist of a direct repeat (DR) sequence that binds the Cas13 protein and a spacer sequence that determines target specificity through complementary base pairing with the target mRNA. The design principles outlined below are critical for ensuring high on-target efficiency and minimal off-target effects.
Spacer Sequence Selection:
- Length: Design spacer sequences of 28-30 nucleotides for optimal activity and specificity [27]. The precise length can vary slightly between Cas13 orthologs.
- Specificity: Perform BLAST analysis to ensure spacer sequence is unique to the target transcript, minimizing off-target binding.
- Accessibility: Target regions with minimal secondary RNA structure to facilitate crRNA binding. Computational tools like RNAfold can predict accessible regions.
- Mismatch Tolerance: Be aware that Cas13 exhibits tolerance for single mismatches, with spacer nucleotides in positions 15-21 being particularly sensitive to mismatches [13]. Avoid designs where off-target transcripts differ by only a single nucleotide in permissive regions.
Specificity Enhancement Strategies:
- Synthetic Mismatch: Intentionally introduce one or more mismatches in the crRNA spacer to the desired target. This design tolerates limited mismatches to the intended target but prevents binding to off-targets with additional mismatches, enabling specific detection of single nucleotide polymorphisms (SNPs) [13].
- Hairpin Extension: Extend the spacer at the 3' end with a sequence that forms a short stem-loop structure. This hairpin blocks part of the spacer sequence, preventing the crRNA from binding to off-target RNA sequences and improving specificity for SNP identification, as demonstrated in the CRISPR/Cas13a system [13].
Direct Repeat Considerations:
- The direct repeat sequence is conserved for each Cas13 ortholog and is typically 60-66 nucleotides long, forming a short hairpin structure adjacent to the spacer sequence [27].
- Ensure compatibility between the DR in your expression vector and the specific Cas13 protein being expressed (e.g., Cas13a, Cas13b, Cas13d).
Table 2: crRNA Design Parameters for Different Cas13 Orthologs
| Parameter | Cas13a | Cas13b | Cas13d (CasRx) |
|---|---|---|---|
| Spacer Length | 28 nt | 30 nt | 30 nt |
| Direct Repeat Length | ~66 nt | ~64 nt | ~60 nt |
| PFS Requirement | None in eukaryotic cells [27] | None in eukaryotic cells [27] | None |
| Reported Efficiency in Cells | High | High | Highest efficiency, minimal off-targets [27] [23] |
| Size (aa) | ~1000 | ~1100 | ~930 |
Experimental Protocol: crRNA Cloning and Biofilm Screening
crRNA Cloning into Expression Vectors
This protocol describes the insertion of custom spacer sequences into a crRNA expression plasmid via inverse PCR, adapted from established methods [26] [25].
Materials:
- crRNA expression plasmid (e.g., pgRNA plasmid [26])
- High-fidelity DNA polymerase (e.g., Q5 Hot Start)
- T4 Polynucleotide Kinase
- T4 DNA Ligase
- DpnI restriction enzyme
- Gel extraction kit
- Competent E. coli (e.g., Top10)
Procedure:
- Primer Design: Design phosphorylated primers containing the 20-30 nt target-specific spacer sequence flanked by 35 nt overhangs homologous to the crRNA scaffold in the expression plasmid.
- Inverse PCR: Set up PCR reaction using the crRNA expression plasmid as template. Cycling conditions: initial denaturation 98°C for 30 sec; 25 cycles of 98°C for 10 sec, 60°C for 20 sec, 72°C for 2-3 min (depending on plasmid size); final extension 72°C for 5 min.
- Template Digestion: Add DpnI directly to PCR product (1 μL per 50 μL reaction), incubate at 37°C for 1 hour to digest methylated template DNA.
- Purification: Gel-purify the linear PCR product using a gel extraction kit.
- Ligation: Perform blunt-end ligation using T4 DNA Ligase. Incubate at room temperature for 1 hour or overnight at 16°C.
- Transformation: Transform ligation reaction into competent E. coli cells, plate on selective media.
- Colony Screening: Pick individual colonies, perform colony PCR with verification primers, and send PCR products for Sanger sequencing to confirm correct insertion.
Biofilm Phenotyping Workflow
After verifying crRNA sequences, the following comprehensive workflow characterizes the resulting biofilm phenotypes. The process from genetic targeting to phenotypic analysis is summarized below:
Detailed Protocols for Phenotyping Assays:
A. mRNA Knockdown Validation (RT-qPCR)
- Extract total RNA from bacterial cultures using Trizol reagent [26].
- Treat RNA with DNase I to remove genomic DNA contamination.
- Synthesize cDNA using reverse transcriptase with random hexamers.
- Perform qPCR with gene-specific primers and normalize to housekeeping genes (e.g., 16S rRNA).
- Calculate knockdown efficiency using the 2^(-ΔΔCt) method.
B. Crystal Violet Biofilm Assay (Total Biomass)
- Grow bacterial cultures with induced Cas13/crRNA expression in 96-well plates for 24-48 hours.
- Carefully remove planktonic cells and gently wash adhered cells with PBS.
- Fix biofilms with 99% methanol for 15 minutes, then stain with 0.1% crystal violet for 20 minutes.
- Wash excess stain and solubilize bound dye with 33% acetic acid.
- Measure absorbance at 570 nm to quantify total biofilm biomass [26].
C. XTT Reduction Assay (Metabolic Activity)
- Prepare XTT solution (0.5 mg/mL) with menadione (1 μM) as an electron-coupling agent.
- Add XTT-menadione to washed biofilms and incubate in dark for 2-3 hours.
- Measure absorbance at 490 nm to assess metabolic activity of biofilm cells [26].
D. Confocal Laser Scanning Microscopy (CLSM)
- Grow biofilms on appropriate surfaces (e.g., glass coverslips).
- Stain with fluorescent dyes: SYTO9 for bacterial cells (green fluorescence), ConA-TRITC for exopolysaccharides (red fluorescence), or FilmTracer SYPRO Ruby for matrix proteins.
- Image using 20× or 40× water immersion objectives.
- Acquire z-stacks and reconstruct 3D architecture using Imaris or ImageJ software [25].
- Quantify biofilm parameters: biomass, thickness, roughness coefficient, and surface coverage.
Research Reagent Solutions
The table below outlines essential reagents and tools for implementing CRISPR-Cas13 based biofilm research, compiled from methodologies in the cited literature.
Table 3: Essential Research Reagents for CRISPR-Cas13 Biofilm Studies
| Reagent/Tool | Function/Application | Example Sources/References |
|---|---|---|
| dCas13 Effectors | Catalytically dead Cas13 for RNA binding without cleavage; base for fusion proteins | dCas13-GFP (RNA tracking) [13]; dCas13-deaminase (RNA editing) [13] |
| Cas13 Expression Plasmids | Mammalian, bacterial, or viral expression of Cas13 orthologs | AddGene plasmids #44249, #44251 [26] |
| crRNA Cloning Vectors | Backbone for expressing custom guide RNAs | pgRNA plasmid [26] |
| Inducible Systems | Tight control of Cas13/dCas13 expression | PtetA promoter with aTc inducer [25] |
| Fluorescent Reporters | Visualizing target RNA localization and abundance | dCas13-GFP, dCas13-mNeonGreen fusions [13] [25] |
| AAV Delivery Vectors | In vivo delivery of Cas13 components; small size of Cas13d advantageous | AAV-mediated Cas13d for in vivo models [13] |
| Biofilm Staining Kits | Matrix and cellular visualization for microscopy | FilmTracer SYPRO Ruby, Concanavalin A conjugates [25] |
Troubleshooting and Technical Considerations
Knockdown Efficiency Optimization:
- Test multiple crRNAs targeting different regions of the same transcript to identify the most effective guide.
- Verify Cas13 expression and nuclear localization if targeting nuclear RNAs.
- For in vivo applications, select the appropriate Cas13 ortholog based on size constraints and efficiency. Cas13d (CasRx) has demonstrated high efficiency and minimal off-target effects in mammalian cells [27] [23].
Specificity Controls:
- Include mismatch controls with crRNAs containing 3-5 nucleotide mismatches to the target.
- Perform RNA sequencing to assess transcriptome-wide off-target effects.
- Use multiple crRNAs against the same target to confirm phenotype consistency.
Delivery Considerations:
- For bacterial systems, utilize conjugation-compatible or electroporation-compatible plasmids.
- For eukaryotic cells and in vivo applications, consider AAV, lipid nanoparticles (LNPs), or extracellular vesicles (EVs) for efficient delivery [13].
- For biofilm-specific targeting, explore nanoparticle-based delivery systems that enhance penetration through EPS matrices [12].
CRISPR-Cas13 technology provides a powerful and specific platform for targeting RNA transcripts central to biofilm regulation. The design principles and experimental protocols outlined in this application note enable researchers to systematically dissect the roles of quorum sensing, EPS production, and signaling pathways in biofilm development and maintenance. By implementing these guidelines, scientists can generate robust, reproducible data on biofilm gene function, accelerating both basic research and therapeutic development against biofilm-associated infections.
Bacterial pathogenesis and antimicrobial resistance (AMR) are increasingly defined by the synergistic interplay of biofilm formation, antimicrobial resistance, and quorum sensing (QS) [28]. This "triple threat" enables pathogens to persist in hostile environments, evade immune defenses, and resist conventional therapies, contributing to chronic and recurrent infections. Within this framework, QS acts as a master regulator, coordinating collective bacterial behaviors such as virulence factor production, biofilm maturation, and antibiotic tolerance in response to population density [29].
The emergence of CRISPR-Cas13 technology presents a transformative approach for precision intervention in bacterial RNA-regulated pathways. As an RNA-guided RNA endonuclease, Cas13 can be programmed to target and degrade messenger RNA (mRNA) transcripts of critical QS regulators and biofilm metabolic components, offering a powerful tool for disrupting this coordinated bacterial communication [2]. This case study details the application of CRISPR-Cas13 systems to impede QS circuits and biofilm communication, providing detailed protocols and analytical frameworks for research and therapeutic development.
Technical Foundation
Quorum Sensing Mechanisms in Biofilm Formation
Quorum sensing is a complex bacterial communication system regulated by extracellular signaling molecules called autoinducers. As bacterial population density increases, autoinducer concentration rises, eventually reaching a threshold that triggers synchronized population-wide genetic expression [29]. This process is fundamental to biofilm development and maintenance.
Table 1: Primary Quorum Sensing Signaling Molecules in Gram-Negative Bacteria
| Signaling Molecule | Abbreviation | Common Bacterial Species | Primary Regulatory Role |
|---|---|---|---|
| N-(3-oxododecanoyl)-L-homoserine lactone | 3-oxo-C12-HSL | Pseudomonas aeruginosa | Virulence, biofilm maturation [29] |
| N-butanoyl-L-homoserine lactone | C4-HSL | Aeromonas hydrophila | Protease production, biofilm formation [29] |
| N-(3-oxohexanoyl)-L-homoserine lactone | 3-oxo-C6-HSL | Vibrio fischeri, Erwinia spp. | Bioluminescence, exoenzyme production [29] |
| 2-heptyl-3-hydroxy-4-quinolone | PQS | Pseudomonas aeruginosa | Stress response, virulence factor production [29] |
| Autoinducer-2 | AI-2 | Vibrio harveyi (and many others) | Interspecies communication [29] |
The LuxI/LuxR homolog system represents the canonical QS mechanism in Gram-negative bacteria. LuxI-type synthases produce acyl-homoserine lactone (AHL) autoinducers, which bind to LuxR-type transcriptional regulators upon reaching threshold concentrations, activating or repressing target genes governing biofilm formation, virulence, and antibiotic efflux systems [29].
CRISPR-Cas13 System for RNA Targeting
The CRISPR-Cas13 system (Class 2, Type VI) employs a single RNA-guided RNase protein that targets and cleaves single-stranded RNA (ssRNA) molecules. Unlike DNA-targeting Cas9, Cas13 enables transient, reversible modulation of gene expression without altering the genome, making it ideal for targeting non-essential regulatory pathways like QS [2].
Key Cas13 features relevant to biofilm disruption include:
- Programmable RNA Targeting: Cas13 uses a guide RNA (gRNA) with a spacer sequence complementary to the target mRNA, enabling precise degradation of specific QS transcripts.
- HEPN Domain Catalysis: Target RNA cleavage occurs via two Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains that exhibit Mg²⁺-dependent RNase activity.
- Collateral Activity: Some Cas13 subtypes exhibit promiscuous RNase activity upon target recognition, though engineered variants minimize this for specific transcriptional silencing [2].
Table 2: CRISPR-Cas13 Subtypes and Characteristics for Biofilm Research
| Cas13 Subtype | Size (aa) | sgRNA Spacer Position | Key Features for Biofilm Application |
|---|---|---|---|
| Cas13a | 1000-1200 | 3' end | First characterized; robust activity in prokaryotes [2] |
| Cas13b | 1000-1200 | 5' end | High specificity; multiple variants available |
| Cas13d (CasRx) | ~800 | 5' end | Compact size; high efficiency in eukaryotic and prokaryotic cells [2] |
| Cas13X/Y | 775-800 | 5' end | Smallest variants; minimal collateral damage [2] |
Application Notes: Quantitative Data on CRISPR-Cas13 Biofilm Disruption
Recent studies demonstrate the efficacy of CRISPR-Cas systems in combating biofilm-associated infections. When integrated with nanoparticle delivery platforms, these systems show enhanced penetration and editing efficiency within the complex biofilm matrix.
Table 3: Quantitative Efficacy of CRISPR-Based Anti-Biofilm Strategies
| Intervention Strategy | Target System/Bacteria | Efficacy Metric | Result | Source |
|---|---|---|---|---|
| Liposomal Cas9 formulation | Pseudomonas aeruginosa | Reduction in biofilm biomass | >90% reduction in vitro [12] | |
| CRISPR-gold nanoparticle hybrids | Antibiotic-resistant bacteria | Gene-editing efficiency | 3.5-fold increase vs. non-carrier systems [12] | |
| CRISPRi with dCas9 | Escherichia coli | Target gene repression | Up to 99.9% reduction in gene expression [8] | |
| Cas13d RNA silencing | Bacterial pathogens | mRNA knockdown | Significant transcript reduction with minimal collateral activity [2] |
The integration of CRISPR-Cas13 with nanoparticles enables co-delivery with antibiotics, producing synergistic antibacterial effects and superior biofilm disruption compared to mono-therapeutic approaches. These hybrid platforms address the dual challenge of biofilm penetration and genetic resistance by simultaneously targeting protective matrices and disrupting QS genetic circuits [12].
Experimental Protocols
Protocol 1: Designing gRNAs to Target Quorum Sensing Genes
Principle: Program CRISPR-Cas13 to silence key QS regulatory genes by designing specific gRNAs against mRNA transcripts of LuxI/LuxR homologs.
Materials:
- Target bacterial genome sequence (e.g., P. aeruginosa PAO1)
- gRNA design software (e.g., CHOPCHOP, CRISPRscan)
- Cas13d (CasRx) expression plasmid
- T4 DNA ligase and buffer
- Cloning competent E. coli cells
Procedure:
- Identify Target Sequences: Access the nucleotide sequences for QS regulatory genes (e.g., lasI, lasR, rhlI, rhlR in P. aeruginosa) from NCBI GenBank.
- Design gRNA Spacers: Using design software, identify 20-30nt protospacer sequences within the target mRNA with the following criteria:
- Avoid secondary structure regions in the target RNA
- Ensure minimal off-target potential by BLAST analysis
- Select spacers targeting the 5' region of the coding sequence for maximum knockdown efficiency
- Clone gRNA Expression Cassettes:
- Synthesize oligonucleotides corresponding to the selected spacers
- Anneal and phosphorylate oligonucleotides
- Ligate into the BsmBI-digested Cas13d gRNA expression plasmid
- Transform into cloning competent E coli, then plate on selective media
- Verify Clones: Screen colonies by colony PCR and Sanger sequencing to confirm correct gRNA insertion.
- Package with Delivery System: For nanoparticle delivery, purify plasmid DNA and complex with selected nanocarriers (see Protocol 3).
Protocol 2: Assessing Quorum Sensing Inhibition In Vitro
Principle: Quantify CRISPR-Cas13-mediated QS disruption using reporter strains and virulence factor assays.
Materials:
- Bioluminescent QS reporter strains (e.g., E. coli pSB1075 for 3-oxo-C12-HSL detection)
- Target pathogen with functional QS system (e.g., P. aeruginosa)
- Cas13d ribonucleoprotein (RNP) complexes with target-specific gRNA
- LB broth and agar
- Microtiter plates (96-well, black-walled for bioluminescence)
- Spectrofluorometer or microplate reader
Procedure:
- Prepare Bacterial Cultures:
- Grow target pathogen overnight in LB broth at 37°C with shaking
- Dilute 1:100 in fresh medium and grow to mid-log phase (OD600 ≈ 0.5)
- Treat with CRISPR-Cas13:
- Divide culture into experimental groups:
- Group 1: Untreated control
- Group 2: Non-targeting gRNA-Cas13 RNP (negative control)
- Group 3: Target-specific gRNA-Cas13 RNP
- Group 4: Traditional quorum quenching agent (positive control)
- Transfer 200µL aliquots to 96-well plates
- Divide culture into experimental groups:
- Incubate and Monitor:
- Incubate plates at 37°C with continuous shaking in plate reader
- Measure OD600 (growth) and bioluminescence/fluorescence every 30 minutes for 24h
- Assess Virulence Factor Production (endpoint assays):
- Pyocyanin quantification: Extract with chloroform and HCl, measure OD520
- Protease activity: Azocasein assay, measure OD440
- Elastase activity: Elastin-Congo red assay, measure OD495
- Analyze Data: Normalize virulence factor production to cell density, calculate percentage inhibition compared to controls.
Protocol 3: Nanoparticle-Mediated CRISPR Delivery to Biofilms
Principle: Utilize engineered nanoparticles to enhance CRISPR-Cas13 delivery and penetration through biofilm extracellular polymeric substance (EPS).
Materials:
- Cationic lipid nanoparticles (LNPs) or gold nanoparticles (AuNPs)
- Cas13d protein and in vitro transcribed gRNA
- Microfluidic mixer or sonication equipment
- Pre-formed biofilms of target pathogen
- Confocal laser scanning microscopy (CLSM) supplies
- SYTO9 and propidium iodide live/dead stains
Procedure:
- Prepare CRISPR-Cas13 RNP Complexes:
- Purify Cas13d protein via affinity chromatography
- Transcribe gRNA in vitro using T7 RNA polymerase
- Form RNP complexes by incubating Cas13d with gRNA (1:2 molar ratio) at 25°C for 15min
- Formulate Nanoparticle-CRISPR Complexes:
- For LNPs: Use microfluidic mixer to combine cationic lipids, cholesterol, PEG-lipid, and RNP complexes in aqueous buffer
- For AuNPs: Conjugate RNP complexes to citrate-stabilized AuNPs via electrostatic adsorption or thiol chemistry
- Characterize size and zeta potential using dynamic light scattering
- Treat Pre-formed Biofilms:
- Grow 48h biofilms of target pathogen on suitable substrate (e.g., peg lids, glass coverslips)
- Gently wash with PBS to remove planktonic cells
- Add nanoparticle-CRISPR formulations to biofilm, incubate 2-4h at 37°C
- Include untreated and empty nanoparticle controls
- Assess Biofilm Viability and Structure:
- Live/dead staining: Incubate with SYTO9/propidium iodide, visualize by CLSM
- Biomass quantification: Analyze CLSM z-stacks with image analysis software (e.g., COMSTAT)
- Metabolic activity: Resazurin reduction assay, measure fluorescence (Ex560/Em590)
- Evaluate Gene Knockdown Efficiency:
- Harvest biofilm cells, extract total RNA
- Perform RT-qPCR for target QS genes (e.g., lasI, lasR)
- Normalize to housekeeping genes, calculate fold reduction vs. controls
Pathway Visualization and Experimental Workflows
Diagram 1: CRISPR-Cas13 Mediated Disruption of Quorum Sensing in Biofilm Formation
Diagram 2: Experimental Workflow for CRISPR-Cas13 Mediated QS Disruption
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Research Reagents for CRISPR-Cas13 Biofilm Studies
| Reagent / Material | Supplier Examples | Function in QS Disruption Research |
|---|---|---|
| Cas13d (CasRx) Protein | Thermo Fisher, Sigma-Aldrich, Abcam | RNA-guided RNase effector for targeted mRNA degradation |
| gRNA In Vitro Transcription Kit | NEB, Thermo Fisher, Takara | Production of specific guide RNAs for Cas13 programming |
| Cationic Lipid Nanoparticles | Precision NanoSystems, Avanti Polar Lipids | Enhanced delivery of CRISPR components through biofilm EPS |
| Gold Nanoparticles (20nm) | nanoComposix, Sigma-Aldrich | Conjugation platform for RNP complex delivery |
| QS Reporter Strains | ATCC, DSMZ | Bioluminescent/fluorescent detection of QS activity |
| P. aeruginosa PAO1 Wild Type | ATCC 15692, DSM 22644 | Model organism for Gram-negative QS studies |
| Microfluidic Mixer | Dolomite, Precision NanoSystems | Reproducible nanoparticle formulation |
| Confocal Laser Scanning Microscope | Zeiss, Leica, Nikon | High-resolution 3D biofilm imaging and analysis |
| RT-qPCR Reagents | Bio-Rad, Qiagen, Thermo Fisher | Quantification of QS gene expression knockdown |
The extracellular polymeric substance (EPS) matrix is a critical determinant of biofilm integrity, conferring remarkable resistance to antimicrobial treatments and environmental stresses. This matrix, constituting 75-90% of the biofilm's dry mass, is a complex amalgam of polysaccharides, proteins, nucleic acids, and lipids that forms a protective scaffold around microbial cells [30]. The resilient nature of biofilms stems primarily from this EPS architecture, which limits antibiotic penetration, facilitates nutrient cycling, and protects embedded cells from host immune responses [30] [11].
CRISPR-Cas13 technology presents a transformative approach for precision targeting of biofilm stability through RNA-level interference with EPS biosynthesis pathways. Unlike DNA-editing CRISPR systems, Cas13 targets messenger RNA (mRNA) transcripts, allowing for transient knockdown of essential EPS genes without permanent genomic alterations [8] [31]. This RNA-targeting capability enables precise disruption of the metabolic pathways responsible for building and maintaining the EPS matrix, potentially weakening the structural foundation that makes biofilms so recalcitrant to conventional treatments.
EPS Composition and Key Biosynthesis Targets
Architectural Components of the EPS Matrix
The biofilm EPS matrix is a sophisticated polymeric network whose composition varies between bacterial species but typically includes three primary components:
- Exopolysaccharides: Heterogeneous sugar polymers (e.g., Pel, Psl, and alginate in Pseudomonas aeruginosa) that form the structural backbone of the matrix through van der Waals interactions, hydrogen bonding, and electrostatic forces [30]. These polysaccharides act as molecular adhesives for surface attachment and provide mechanical stability to the biofilm architecture.
- Extracellular proteins: Secreted proteins, cell surface adhesins, and protein subunits of appendages (pili, flagella) that interact with polysaccharides to stabilize the matrix architecture [30]. These proteins also include enzymes for matrix degradation and remodeling during biofilm development.
- Extracellular DNA (eDNA): Nucleic acids released from lysed cells that contribute to matrix viscosity, structural integrity, and horizontal gene transfer between biofilm-embedded cells [30].
Essential EPS Biosynthesis Gene Clusters
Genetic studies across multiple bacterial species have identified dedicated gene clusters responsible for EPS production and biofilm formation. The experimental disruption of these clusters provides compelling evidence for their essential role in biofilm integrity:
Table 1: Experimentally Validated EPS Gene Clusters in Model Bacteria
| Bacterial Species | Gene Cluster | Gene Count | Function | Phenotype After Disruption |
|---|---|---|---|---|
| Paenibacillus polymyxa WLY78 | pep-2 | 17 | EPS biosynthesis | ~70% reduction in biofilm formation; enhanced motility; colony morphology changes [32] |
| Bacillus subtilis | epsA-O | 15 | Exopolysaccharide synthesis | Complete loss of structured biofilm; impaired colony wrinkling [33] [32] |
| Lactobacillus paraplantarum | eps | 18 | Heteropolysaccharide production | Reduced matrix production and surface adhesion [32] |
The critical finding from these genetic studies is that targeted disruption of specific EPS biosynthesis genes, particularly those within the identified clusters, consistently produces structurally compromised biofilms with enhanced susceptibility to antimicrobial agents and physical removal [32].
CRISPR-Cas13 Mechanism for RNA Targeting
Molecular Basis of Cas13 Function
CRISPR-Cas13 systems belong to Class 2 Type VI CRISPR systems, characterized by a single effector protein that mediates RNA-guided RNA targeting. The mechanism involves:
- Guide RNA Complex Formation: A CRISPR RNA (crRNA) containing a programmable spacer sequence complexes with the Cas13 protein to form the ribonucleoprotein effector complex [10] [31].
- Target Recognition: The crRNA spacer directs Cas13 to complementary single-stranded RNA (ssRNA) targets through base-pairing interactions, without requiring a protospacer-adjacent motif (PAM) [34].
- Collateral Cleavage Activation: Upon target recognition, Cas13 undergoes conformational activation and exhibits non-specific RNase activity (trans-cleavage) that degrades both target and collateral RNA molecules in the immediate vicinity [34].
The recent discovery that Cas13 evolved from AbiF toxin-antitoxin systems explains its unique RNA-targeting mechanism and collateral cleavage activity, distinguishing it from DNA-targeting CRISPR effectors [10].
Experimental Workflow for EPS Gene Knockdown
The following diagram illustrates the complete experimental workflow for implementing CRISPR-Cas13 to target EPS biosynthesis genes in bacterial biofilms:
Application Notes: CRISPR-Cas13 Implementation for EPS Targeting
Protocol 1: Target Selection and crRNA Design
Objective: Identify optimal EPS biosynthesis gene targets and design corresponding crRNAs for Cas13-mediated knockdown.
Table 2: EPS Gene Target Selection Criteria and Design Parameters
| Parameter | Considerations | Recommendations |
|---|---|---|
| Target Gene Selection | Essentiality for EPS production; redundancy within pathway; conservation across strains | Prioritize non-redundant, pathway-specific genes (e.g., glycosyltransferases, polysaccharide export proteins) [32] |
| crRNA Spacer Design | Target accessibility; secondary structure; GC content (40-60%); length (20-30 nt) | Design 3-5 crRNAs per target with spacer sequences complementary to different regions of the mRNA [31] |
| Specificity Validation | Off-target potential; cross-hybridization with non-target genes | Perform BLAST analysis against host genome; utilize computational tools (Cas13design, CHOPCHOP) [8] |
| Control Designs | Non-targeting crRNAs; essential gene targets for viability assessment | Include crRNAs with scrambled sequences and target housekeeping genes (e.g., rRNA, metabolic enzymes) [33] |
Procedure:
- Identify Target Genes: Select 2-3 critical EPS biosynthesis genes from validated clusters (e.g., pep genes from Paenibacillus, eps genes from Bacillus) using transcriptomic data or literature evidence [32].
- Design crRNA Spacers: For each target gene, design 28-nt spacer sequences targeting regions within the first 500 bp of the coding sequence, avoiding stable secondary structures (ΔG > -5 kcal/mol).
- Validate Specificity: Confirm spacer uniqueness by alignment against the host genome using BLASTN with an E-value cutoff of 0.1.
- Synthesize crRNA Arrays: Clone validated spacer sequences into a crRNA expression vector under a strong, constitutive promoter (U6 for mammalian systems, J23119 for bacterial systems).
Protocol 2: Delivery System Optimization for Biofilm Penetration
Objective: Efficiently deliver CRISPR-Cas13 components through the EPS barrier to target cells within mature biofilms.
Table 3: Delivery Platforms for CRISPR-Cas13 in Biofilm Environments
| Delivery Platform | Mechanism | Efficiency in Biofilms | Applications |
|---|---|---|---|
| Nanoparticle Carriers | Enhanced penetration through EPS; protected cargo delivery; controlled release | Liposomal Cas13: >90% biofilm biomass reduction [11]; Gold nanoparticles: 3.5× editing efficiency [11] | Mature, thick biofilms; in vivo applications |
| Phagemid Vectors | Bacterial virus-mediated transduction; natural biofilm penetration | Variable; species-dependent transduction efficiency | Species-specific delivery; laboratory models |
| Conjugative Plasmids | Bacterial mating; self-transmissible plasmids | Moderate; limited to susceptible strains | Multi-species biofilms; engineered probiotics |
| Chemical Transformation | Permeabilization with cations or polymers; electroporation | Low; primarily for planktonic cells | Preliminary testing; laboratory-adapted strains |
Procedure:
- Prepare Cas13-crRNA Complexes:
- For in vitro assembled complexes: Express and purify LwaCas13a protein, then combine with synthetic crRNA (1:2 molar ratio) in reaction buffer.
- For in vivo expression: Co-transform bacterial cells with Cas13 expression plasmid and crRNA expression vector.
Formulate Nanocarriers:
- Lipid nanoparticles: Mix Cas13-crRNA complexes with cationic lipids (DOTMA, DOPE) using microfluidic mixing at 1:3 weight ratio.
- Gold nanoparticles: Conjugate thiol-modified crRNA to 15-nm gold nanoparticles, then adsorb Cas13 protein onto the complex.
Apply to Biofilms:
- Grow biofilms for 48-72 hours until mature architecture develops.
- Treat with delivery formulations containing 100-500 nM Cas13-crRNA complexes in fresh medium.
- Incubate for 24-48 hours with gentle agitation (50 rpm) to maintain biofilm integrity while allowing penetration.
Protocol 3: Functional Assessment of EPS Disruption
Objective: Quantitatively evaluate the structural and functional consequences of EPS gene knockdown on biofilm integrity.
Procedure:
- Quantify Target mRNA Knockdown:
- Extract total RNA from treated and control biofilms using mechanical disruption (bead beating) and RNA stabilization reagents.
- Perform reverse transcription quantitative PCR (RT-qPCR) with gene-specific primers for targeted EPS genes.
- Calculate knockdown efficiency using the 2^(-ΔΔCt) method normalized to housekeeping genes (e.g., rpoB, gyrA).
Measure EPS Matrix Components:
- Polysaccharide Quantification: Islect EPS by high-speed centrifugation (13,000 × g, 30 min) and measure total carbohydrate content using phenol-sulfuric acid assay with glucose standards [32].
- Protein Content: Determine EPS-associated proteins by Bradford assay of the same EPS extracts.
- eDNA Measurement: Extract eDNA from EPS supernatants using CTAB method and quantify by fluorometry with PicoGreen dye.
Assess Structural Integrity:
- Biomass Quantification: Stain biofilms with crystal violet (0.1% w/v), elute with acetic acid (30% v/v), and measure absorbance at 590 nm.
- Confocal Microscopy: Image GFP-expressing biofilms before and after treatment, quantifying thickness, biovolume, and surface coverage using COMSTAT analysis.
- Mechanical Stability: Apply calibrated shear stress (0-500 mPa) in flow cells and measure biofilm detachment by effluent turbidity.
Evaluate Enhanced Susceptibility:
- Treat Cas13-targeted and control biofilms with sub-inhibitory concentrations of conventional antibiotics (e.g., tobramycin, ciprofloxacin).
- Determine minimum biofilm eradication concentration (MBEC) using the Calgary biofilm device.
- Assess bacterial viability within treated biofilms using LIVE/DEAD BacLight staining and colony-forming unit (CFU) enumeration.
Research Reagent Solutions
Table 4: Essential Research Reagents for CRISPR-Cas13 Biofilm Studies
| Reagent Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Cas13 Orthologues | LwaCas13a (Leptotrichia wadei); PspCas13b (Prevotella sp.); RfxCas13d (Ruminococcus flavefaciens) | RNA-guided RNA cleavage; collateral RNase activity | LwaCas13a shows highest efficiency in bacterial systems; RfxCas13d offers compact size for delivery [31] |
| Delivery Materials | Cationic liposomes (DOTMA/DOPE); Gold nanoparticles (15-20 nm); Polymeric nanoparticles (PLGA) | Protect nucleic acids; enhance biofilm penetration; facilitate cellular uptake | Lipid nanoparticles optimal for laboratory strains; gold nanoparticles provide conjugation flexibility [11] |
| Biofilm Assay Kits | Crystal violet staining kits; LIVE/DEAD BacLight bacterial viability kits; Microtiter plate biofilm kits | Quantify biofilm biomass; assess bacterial viability; high-throughput screening | Standardize across experiments; enable comparative analysis between studies [30] |
| EPS Extraction Reagents | Cetyltrimethylammonium bromide (CTAB); Ethylenediaminetetraacetic acid (EDTA); Cation exchange resins (Dowex) | Precipitate polysaccharides; chelate divalent cations; isolate EPS components | CTAB method effective for polysaccharide-rich matrices; EDTA helps disrupt ionic interactions [35] |
| Analytical Standards | Monosaccharide standards; DNA quantification standards; Protein assay standards | Calibrate analytical measurements; normalize cross-experiment data | Use mixture of glucose, galactose, mannose for polysaccharide analysis [35] |
Targeting EPS biosynthesis genes with CRISPR-Cas13 represents a promising precision approach for compromising biofilm structural integrity. The methodology outlined enables researchers to systematically validate EPS gene function, assess matrix disruption, and quantify consequent increases in antimicrobial susceptibility. Future developments should focus on optimizing delivery efficiency in complex multispecies biofilms, exploring multiplexed targeting of redundant EPS pathways, and integrating Cas13 with conventional antibiofilm strategies for synergistic effects. As Cas13 engineering advances with higher-fidelity variants and improved guide RNA designs, this RNA-targeting approach may ultimately translate into effective therapeutic interventions against persistent biofilm-associated infections.
Application Note: CRISPR-Cas13 for Targeted Gene Knockdown in Biofilms
Antibiotic tolerance in bacterial biofilms presents a major challenge in treating persistent infections. This tolerance is driven by two key mechanisms: the acquisition of heritable antibiotic resistance genes and the presence of dormant persister cells. Conventional antibiotics often fail against biofilms due to reduced penetration through the extracellular polymeric substance (EPS) and the metabolic heterogeneity of embedded bacterial cells [12] [36]. Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) systems offer a transformative, programmable approach for precision antimicrobial therapy [8]. Unlike DNA-cutting Cas nucleases (e.g., Cas9), the RNA-targeting Cas13 system provides a powerful tool for targeted gene knockdown without permanent genomic alteration. This application note details the use of CRISPR-Cas13 to combat antibiotic tolerance by specifically knocking down resistance and persister cell genes within bacterial biofilms, framing this approach within metabolic pathway research.
Mechanism of Action: Cas13 RNA Targeting
The Cas13 protein (e.g., Cas13a) uses a guide RNA (gRNA) to bind and cleave complementary single-stranded RNA sequences target mRNA transcripts. When deployed against bacterial biofilms, this mechanism can be harnessed to:
- Disrupt Antibiotic Resistance: Target and degrade mRNA transcripts of specific antibiotic resistance genes (e.g., beta-lactamases, efflux pump components), resensitizing bacteria to conventional antibiotics [8] [18].
- Eliminate Persister Cells: Knock down key genes involved in toxin-antitoxin systems and stress response (e.g., RelA, components of the (p)ppGpp-mediated stringent response), preventing entry into or promoting exit from the dormant persister state [37] [36].
- Impair Biofilm Integrity: Target transcripts of genes critical for quorum sensing (QS) and EPS production, thereby weakening the biofilm structure and enhancing antibiotic penetration [8].
This application induces a transient and programmable bactericidal or bacteriostatic effect, selectively targeting pathogenic bacteria while sparing the surrounding microbiota—a significant advantage over broad-spectrum antibiotics [8].
Experimental Protocols
Protocol 1: Design and In Vitro Validation of gRNAs for Cas13
Objective
To design and validate gRNAs that efficiently direct Cas13 to cleave target mRNA sequences associated with antibiotic resistance and persistence.
Materials
- Oligonucleotides for gRNA template synthesis.
- In vitro Transcription Kit (e.g., HiScribe T7 Quick High Yield Kit) for gRNA production.
- Purified Cas13 Protein (commercially available or purified in-house).
- Synthetic Target RNA Fragment containing the gene of interest.
- Agarose Gel Electrophoresis system for analyzing cleavage products.
Procedure
- gRNA Design: Using sequence analysis software (e.g., CCTop, Cas13design), identify 28-30 nt target sites within the mRNA of the resistance or persister gene. Avoid off-target sites by performing a BLAST search against the host bacterium's genome.
- gRNA Synthesis: Synthesize DNA oligonucleotides encoding the gRNA scaffold and target sequence. Transcribe gRNAs in vitro, then purify them.
- In Vitro Cleavage Assay:
- In a 20 µL reaction tube, combine:
- Purified Cas13 protein (100 nM final concentration)
- gRNA (120 nM final concentration)
- Target RNA substrate (50 nM final concentration)
- 1X Cas13 reaction buffer
- Incubate at 37°C for 60 minutes.
- Stop the reaction with RNA loading dye.
- Analyze the products on a 2% agarose gel. Successful cleavage is indicated by the disappearance of the full-length target RNA band and the appearance of smaller fragments.
- In a 20 µL reaction tube, combine:
Protocol 2: Delivery of CRISPR-Cas13 System to Biofilms Using Lipid-Based Nanoparticles
Objective
To efficiently deliver the Cas13 ribonucleoprotein (RNP) complex into bacterial cells within a mature biofilm using lipid nanoparticles (LNPs).
Materials
- Cas13 RNP Complex: Pre-complexed purified Cas13 protein and synthesized gRNA.
- Lipid Nanoparticles: Commercially available transfection reagents (e.g., Lipofectamine) or custom-formulated LNPs.
- Mature Bacterial Biofilm: Grown for 48-72 hours in a suitable medium (e.g., LB) in a 96-well plate or on a relevant surface (e.g., silicone, plastic) [8].
- Confocal Laser Scanning Microscopy (CLSM) setup for biofilm imaging.
- Viability Stains: SYTO 9 and propidium iodide (for Live/Dead staining).
Procedure
- RNP Complex Formation: Incubate Cas13 protein with a 1.2:1 molar ratio of gRNA at room temperature for 15 minutes to form the RNP complex.
- LNP Formulation: Mix the Cas13 RNP complex with the LNP reagent according to the manufacturer's protocol. For custom LNPs, use microfluidics to encapsulate the RNP at a specific nitrogen-to-phosphate ratio.
- Biofilm Treatment:
- Gently wash mature biofilms twice with PBS or a suitable buffer to remove non-adherent cells.
- Apply the LNP-RNP formulation to the biofilm. For a 96-well plate, a 100 µL volume per well is typical.
- Incubate under conditions optimal for the target bacterium (e.g., 37°C for 4-6 hours).
- Efficacy Assessment:
- Biomass Quantification: Use crystal violet staining to measure total biofilm biomass post-treatment.
- Viability Assay: Perform Live/Dead staining and analyze with CLSM. Calculate the percentage of dead (red) cells versus live (green) cells.
- Gene Knockdown Confirmation: Extract total RNA from treated and control biofilms. Perform RT-qPCR to quantify the reduction in the target mRNA level.
Protocol 3: Combinatorial Treatment with CRISPR-Cas13 and Antibiotics
Objective
To evaluate the synergistic effect of Cas13-mediated gene knockdown and conventional antibiotics on biofilm eradication.
Materials
- Cas13 RNP-LNP Formulation (from Protocol 2).
- Relevant Antibiotic (e.g., ampicillin for a β-lactamase-targeting approach).
- 96-well Microtiter Plates with pre-grown biofilms.
- Microplate Reader for absorbance measurement.
Procedure
- Pre-treatment: Treat mature biofilms with the Cas13 RNP-LNP formulation as described in Protocol 2.
- Antibiotic Challenge: After the 4-6 hour incubation with RNP-LNPs, gently wash the biofilms and add fresh medium containing a sub-inhibitory concentration (e.g., 0.5 to 2 µg/mL for ampicillin) or a standard MIC-level concentration of the antibiotic.
- Incubation and Analysis:
- Incubate the plates for an additional 16-24 hours.
- Assess biofilm viability and biomass using the methods described in Protocol 2.
- Compare the results to biofilms treated with RNP-LNPs alone, antibiotics alone, and an untreated control. Synergy is indicated by a significantly greater reduction in viability or biomass in the combination treatment group.
Table 1: Summary of Key Quantitative Findings from CRISPR-Based Biofilm Control Studies
| Parameter | Value / Result | Experimental Context | Source |
|---|---|---|---|
| Biofilm Biomass Reduction | >90% | Liposomal Cas9 formulation against P. aeruginosa biofilm in vitro | [12] |
| Gene-Editing Efficiency | 3.5-fold increase | CRISPR-gold nanoparticle hybrids vs. non-carrier systems | [12] |
| Antibiotic Tolerance | Up to 1000x greater | Biofilm-associated bacteria vs. planktonic cells | [12] |
| Global Economic Burden | ~$324 billion/year | Biofilm-related losses in the global agrifood sector | [8] |
| U.S. Foodborne Illness Cost | ~$17.6 billion/year | Annual cost of foodborne illnesses | [8] |
Table 2: Key Research Reagent Solutions for CRISPR-Cas13 Anti-Biofilm Experiments
| Reagent / Material | Function / Purpose | Example / Notes |
|---|---|---|
| Cas13 Protein | RNA-targeting nuclease; effector molecule | Purified Cas13a (LwaCas13a) or Cas13b |
| Guide RNA (gRNA) | Programmable RNA component for target specificity | Chemically synthesized or in vitro transcribed |
| Lipid Nanoparticles (LNPs) | Delivery vector for RNP complexes into biofilm cells | Enhances cellular uptake and protects RNPs [12] |
| Gold Nanoparticles | Alternative delivery vector for CRISPR components | Can increase editing efficiency [12] |
| Dispersin B | Adjunct enzyme; degrades biofilm polysaccharide matrix | Enhances RNP penetration [36] |
| DNase I | Adjunct enzyme; degrades extracellular DNA (eDNA) in biofilm matrix | Disrupts biofilm structure [36] |
| SYTO 9/Propidium Iodide | Fluorescent viability stains for confocal microscopy | Differentiates live (green) and dead (red) cells |
Signaling Pathways and Experimental Workflows
Diagram: CRISPR-Cas13 Mechanism for Disrupting Biofilm Homeostasis
Diagram: LNP-Mediated CRISPR-Cas13 Delivery and Workflow
Application Notes
The targeted delivery of CRISPR-Cas13 systems into bacteria represents a pivotal challenge in advancing RNA-level interventions against biofilm metabolic pathways. Two primary delivery strategies—phage-based vectors and nanoparticle systems—have demonstrated significant promise for facilitating bacterial entry, each with distinct advantages and applications for research and therapeutic development.
Phage-based vectors leverage the natural ability of bacteriophages to infect specific bacterial hosts, providing a highly targeted delivery mechanism. Recent advances have enabled the engineering of phage λ to deliver complex CRISPR-guided transposases (DART system) or Cas13a machinery directly to bacterial cells, achieving editing efficiencies surpassing 50% in mixed bacterial communities [38] [39]. This approach offers exceptional species-specificity, making it particularly valuable for precision editing within complex microbial consortia without disrupting non-target species. The incorporation of Cas13a-based counterselection in phage engineering further enhances the precision of these delivery platforms [38].
Nanoparticle systems provide an alternative delivery strategy that can be optimized for different bacterial membrane properties. Lipid nanoparticles (LNPs), previously successful in eukaryotic nucleic acid delivery, have been adapted for bacterial applications through systematic screening of lipid compositions. Research has identified specific LNP formulations (LNP 496 and LNP 470) that efficiently deliver plasmid DNA into Escherichia coli when combined with membrane-weakening agents like polymyxin B [40]. These nano-carriers demonstrate versatility in encapsulating various nucleic acid payloads, including Cas13a/gRNA expression vectors, and have shown efficacy in controlling bacterial infections in animal models [40].
For biofilm research, both delivery systems must overcome the physical and physiological barriers presented by the extracellular polymeric substance (EPS) matrix. Nanoparticles, particularly metal-based formulations, have demonstrated inherent anti-biofilm properties by penetrating the EPS and interfering with quorum-sensing and adhesion-related genes [41]. The integration of CRISPR-Cas13 with nanoparticle-based delivery creates a synergistic strategy that simultaneously targets biofilm metabolic pathways at the RNA level while disrupting the structural integrity of the biofilm community [11].
Table 1: Performance Comparison of Delivery Systems for Bacterial CRISPR-Cas13 Delivery
| Delivery System | Reported Efficiency | Key Advantages | Payload Capacity | Optimal Use Cases |
|---|---|---|---|---|
| Phage λ with DART System | >50% editing efficiency in mixed communities [38] | High species specificity; self-replicating delivery vehicle | Large inserts (>10 kb demonstrated) [38] | Complex microbiome editing; in situ manipulations |
| Engineered CAPs (SNIPR001) | 1-6 log10 CFU reduction [42] | Target diversity; reduced phage-tolerant survivors | CRISPR arrays with Cas effectors [42] | Targeted E. coli reduction; biofilm disruption |
| Lipid Nanoparticles (LNP 496/470) | Efficient plasmid delivery with polymyxin B [40] | Modular composition; proven clinical safety profile | Cas13a/gRNA expression vectors [40] | Gram-negative pathogens; animal model systems |
| Metal/Metal Oxide Nanoparticles | Up to 90% biofilm reduction [11] | Intrinsic antibacterial properties; EPS penetration | Various nucleic acid types [41] | Biofilm-related infections; multidrug-resistant strains |
The strategic selection between phage and nanoparticle delivery platforms depends on multiple factors, including target bacterial species, community complexity, payload requirements, and the specific biofilm metabolic pathways under investigation. Phage-based systems offer unparalleled specificity for well-characterized bacterial targets, while nanoparticle platforms provide more flexible, broad-spectrum delivery capabilities that can be tuned through adjustments to their physicochemical properties [40] [42].
The evaluation of delivery platform efficacy requires careful consideration of multiple quantitative parameters. The following tables consolidate key performance metrics from recent studies to enable direct comparison and informed selection for research applications.
Table 2: Quantitative Efficacy Metrics for Bacterial Delivery Systems
| Delivery Platform | Bacterial Model | Editing/Killing Efficiency | Payload Delivered | Key Experimental Conditions |
|---|---|---|---|---|
| λ-DART Phage [38] | Escherichia coli in mixed community | >50% population editing | CRISPR-guided transposase (DART) | Non-lysogenic phage; MOI optimization |
| CRISPR-Cas-armed Phages (SNIPR001) [42] | Diverse E. coli strains | 3.5 log10 CFU ml-1 reduction | Type I-E CRISPR-Cas system with PbolA promoter | Biofilm growth conditions; bolA promoter |
| LNP 496/470 with Polymyxin B [40] | Escherichia coli BW25113 | Efficient plasmid delivery demonstrated | Cas13a/gRNA expression vector | 10-25 mol% DOTAP; polymyxin B co-treatment |
| Gold Nanoparticle-CRISPR Hybrids [11] | Pseudomonas aeruginosa biofilm | 3.5× editing efficiency increase; >90% biomass reduction | CRISPR-Cas9 components | Co-delivery with antibiotics; biofilm penetration |
Table 3: Nanoparticle Formulations for Anti-Biofilm Applications
| Nanoparticle Type | Target Biofilm-Related Genes | Reported Anti-Biofilm Efficacy | Proposed Mechanism of Action |
|---|---|---|---|
| Silver (Ag) NPs [41] | rhlI-rhlR, lasR, lecA (P. aeruginosa) | Significant disruption of mature biofilms | Quorum sensing interference; EPS penetration |
| Zinc Oxide (ZnO) NPs [41] | icaADBC, eno, ebps (S. aureus) | Inhibition of biofilm formation and maturation | ROS generation; membrane disruption |
| Lipid-Based NPs [40] [11] | N/A (CRISPR delivery vehicle) | >90% P. aeruginosa biofilm biomass reduction [11] | Enhanced cellular uptake; controlled release |
| Titanium Dioxide (TiO2) NPs [41] | gtfs, gbps, relA (S. mutans) | Effective against dental plaque biofilms | Photocatalytic activity; matrix degradation |
Experimental Protocols
Protocol 1: Engineering Phage λ for CRISPR-Cas13a Delivery
Principle: This protocol describes the modification of phage λ using homologous recombination coupled with Cas13a-based counterselection to create phage vectors capable of delivering Cas13a machinery to bacterial cells [38].
Materials:
- Phage λ stock (wild-type)
- Escherichia coli donor and recipient strains
- Cas13a expression plasmid
- Homologous recombination templates with desired inserts
- Luria-Bertani (LB) broth and agar
- Antibiotics for selection (as required)
- PCR reagents for verification
Procedure:
- Design of Homology Arms: Prepare homology templates containing the Cas13a expression cassette flanked by 500-1000 bp sequences homologous to the target insertion site in the phage λ genome.
- Electroporation: Introduce the Cas13a expression plasmid and homology templates into E. coli donor strains containing the phage λ genome via electroporation.
- Induction of Recombination: Induce homologous recombination using temperature shift or chemical inducers appropriate for the specific phage λ system.
- Cas13a Counterselection: Apply Cas13a-based counterselection to eliminate unmodified phages by exploiting Cas13a's RNA-targeting capability to specifically cleave RNA in cells infected with wild-type phages [38].
- Plaque Isolation and Verification: Iscrete individual plaques and verify successful recombination events through PCR screening and sequencing of the modified regions.
- Phage Amplification: Amplify verified recombinant phages in appropriate E. coli host strains and purify using polyethylene glycol precipitation or cesium chloride gradient centrifugation.
- Titer Determination: Determine phage titers using standard plaque assay methods and store at 4°C with appropriate stabilizers.
Technical Notes:
- The Cas13a counterselection step is crucial for efficient isolation of recombinant phages without the need for additional selectable markers.
- For large insertions (>10 kb), consider sequential modifications to maintain phage viability.
- Validate Cas13a functionality in the final phage construct through RNA targeting assays in the intended bacterial host.
Protocol 2: LNP Formulation for Cas13a/gRNA Delivery to Bacteria
Principle: This protocol outlines the preparation and optimization of lipid nanoparticles for delivering Cas13a/gRNA expression vectors to bacterial cells, with particular efficacy against Gram-negative pathogens when combined with membrane-weakening agents [40].
Materials:
- Cationic lipid DOTAP (1,2-dioleoyl-3-trimethylammonium-propane)
- Helper lipids (DOPE, cholesterol, DSPC)
- PEG-lipid (DMG-PEG2000)
- Cas13a/gRNA expression vector (purified)
- Ethanol and citrate buffer (pH 4.0)
- Polymyxin B nonapeptide
- Microfluidic mixer (NanoAssemblr or similar)
- Dialysis membranes (MWCO 100 kDa)
Procedure:
- Lipid Stock Preparation: Prepare individual lipid stock solutions in ethanol at 10-50 mg/mL concentrations. For a standard formulation, use DOTAP at 10-25 mol% based on screening results showing optimal delivery in this range [40].
- Aqueous Phase Preparation: Dilute the Cas13a/gRNA expression vector in citrate buffer (pH 4.0) to a final concentration of 50-100 μg/mL.
- Nanoparticle Formation: Use a microfluidic mixer to combine the lipid mixture (in ethanol) with the aqueous nucleic acid solution at a typical flow rate ratio of 3:1 (aqueous:organic) and total flow rate of 12 mL/min.
- Buffer Exchange: Immediately dilute the formed LNPs with PBS (pH 7.4) and dialyze against a large volume of PBS for 24 hours at 4°C to remove ethanol and exchange the buffer.
- Characterization: Measure particle size and zeta potential using dynamic light scattering. Determine encapsulation efficiency using quantitative PCR or fluorescence-based assays.
- Bacterial Treatment: Combine LNP formulations with sub-inhibitory concentrations of polymyxin B (0.1-1 μg/mL) to weaken bacterial membranes and enhance LNP entry [40].
- Application: Incubate bacteria with LNP-polymyxin B combinations for 2-4 hours before assessing Cas13a delivery efficiency via RNA targeting assays.
Technical Notes:
- The cationic lipid DOTAP is critical for bacterial delivery and should constitute 10-25 mol% of the total lipid composition [40].
- Maintain a nitrogen-to-phosphate (N/P) ratio between 4:1 and 8:1 for optimal nucleic acid encapsulation and delivery efficiency.
- Polymyxin B concentration should be optimized for each bacterial strain to achieve membrane permeabilization without significant bactericidal effects.
Visualization Diagrams
The Scientist's Toolkit
Table 4: Essential Research Reagent Solutions for CRISPR-Cas13 Bacterial Delivery
| Reagent/Material | Function/Application | Example Specifications | Key Considerations |
|---|---|---|---|
| Phage λ Engineering System | Template for bacteriophage-based delivery | λ cI857 Sam7 variant for controlled infection [38] | Amber-suppressor host required for propagation |
| Cas13a Expression Constructs | RNA-targeting effector module | LwaCas13a or RfxCas13d for minimal size and high activity [2] | Consider collateral RNAse activity in experimental design |
| Lipid Nanoparticle Components | Nucleic acid encapsulation and delivery | DOTAP (10-25 mol%), DOPE, cholesterol, DMG-PEG2000 [40] | Cationic lipid content crucial for bacterial entry |
| Membrane Permeabilizers | Enhance delivery efficiency to Gram-negative bacteria | Polymyxin B nonapeptide (0.1-1 μg/mL) [40] | Use sub-inhibitory concentrations to avoid killing |
| Bacterial Biofilm Models | Experimental systems for testing delivery efficacy | Multi-species communities or monospecies biofilms [11] [41] | Consider flow conditions for mature biofilm formation |
| Promoter Systems for Bacterial Expression | Drive Cas13a expression in target bacteria | PbolA for biofilm conditions [42] | Stress-inducible promoters may enhance specificity |
| Metal/Metal Oxide Nanoparticles | Alternative delivery vehicles with intrinsic anti-biofilm properties | Ag, ZnO, or TiO2 NPs (10-100 nm) [41] | Size and surface charge critical for biofilm penetration |
| Quantitative Assessment Tools | Measure delivery efficiency and functional outcomes | qRT-PCR for target RNA degradation; confocal microscopy for biofilm architecture [11] | Combine multiple readouts for comprehensive assessment |
Enhancing Efficacy and Specificity: Overcoming Hurdles in Cas13 Application
Addressing Delivery Inefficiency in Dense Biofilm Matrices
The extracellular polymeric substance (EPS) matrix of bacterial biofilms presents a formidable barrier to therapeutic delivery, rendering conventional antimicrobial treatments increasingly ineffective [43]. This dense, gel-like matrix is composed of a complex network of exopolysaccharides, proteins, extracellular DNA (eDNA), and lipids that encase microbial communities, providing exceptional protection against environmental stresses and antimicrobial agents [43] [44]. The biofilm matrix acts as a molecular sieve, significantly reducing the diffusion and penetration of therapeutic molecules through multiple mechanisms including adsorption to matrix components, enzymatic degradation, and reduced influx due to decreased membrane permeability [44]. This protective function contributes substantially to the estimated $5 trillion USD annual economic impact of biofilms globally across healthcare, industrial, and environmental sectors [43] [45].
Within the specific context of CRISPR-Cas13 delivery for targeting biofilm metabolic pathways, these barriers become particularly problematic. The Cas13 system's ribonucleoprotein (RNP) complexes and guide RNAs are vulnerable to enzymatic degradation and physical entrapment within the biofilm matrix [13] [46]. Furthermore, the heterogeneous metabolic activity and reduced growth rates of biofilm-embedded bacteria diminish the effectiveness of RNA-targeting strategies that require cellular uptake and activity [44]. Addressing these delivery inefficiencies is therefore paramount for advancing CRISPR-Cas13 applications in biofilm research and therapeutic development.
Quantitative Analysis of Biofilm Barrier Properties
Recent investigations have quantified the composition and resilience of biofilm matrices, providing critical baseline data for developing delivery strategies. As shown in Table 1, the distribution of major matrix components creates a multi-faceted barrier system with distinct functional properties.
Table 1: Quantitative Analysis of S. aureus Biofilm Matrix Components and Response to Matrix-Targeting Agents
| Matrix Component | Function in Biofilm Integrity | Mean Occupied Area in Control Biofilms (%) | Reduction with TXA Treatment (%) | Primary Barrier Mechanism |
|---|---|---|---|---|
| Extracellular proteins [47] | Structural stability, enzyme activity | 17.58 ± 1.22 | 99.2 ± 0.1 | Molecular adsorption, enzymatic degradation |
| α-extracellular polysaccharides [47] | Architecture, adhesion, water retention | 16.34 ± 4.71 | 89.7 ± 0.3 | Size exclusion, charge interactions |
| α-β-N-acetylglucosamine [47] | Rigidity, surface attachment | 16.77 ± 1.36 | 96.6 ± 0.1 | Hydrophobic interactions, matrix density |
| Bacterial DNA [47] | Genetic information, structural support | 16.55 ± 13.42 | 90.3 ± 0.4 | Charge-mediated binding, network formation |
| Extracellular DNA (eDNA) [47] | Matrix stability, cation chelation, horizontal gene transfer | 12.43 ± 6.23 | 99.5 ± 0.1 | Viscosity increase, neutrophil trap |
The data demonstrate that effective matrix disruption requires a multi-targeted approach, as no single component dominates the barrier function. The exceptional reduction of all matrix components with tranexamic acid (TXA) treatment (89.7-99.5%) highlights the potential of matrix-disrupting agents to enhance therapeutic delivery [47]. This quantitative framework provides essential metrics for evaluating the efficacy of delivery-enhancing strategies for CRISPR-Cas13 systems in biofilm applications.
CRISPR-Cas13 Delivery Workflow for Biofilm Penetration
The following workflow integrates matrix disruption with optimized CRISPR-Cas13 delivery for effective targeting of biofilm metabolic pathways.
Experimental Protocols for Enhanced Delivery
Protocol 1: Biofilm Matrix Disruption Pre-treatment
Objective: To reduce physical and chemical barriers within the biofilm matrix prior to CRISPR-Cas13 application.
Materials:
- Tranexamic acid (TXA) stock solution (100 mg/mL in sterile water)
- Biofilm-dispersing enzymes (dispersin B, DNase I, glycoside hydrolases)
- Quorum sensing inhibitors (QSIs; e.g., 5-hydroxymethylfurfural, piper betle extract)
- 24-well polystyrene plates with established biofilms
- Phosphate-buffered saline (PBS), pH 7.4
Procedure:
- Grow biofilms for 24-48 hours using appropriate bacterial strains and growth media under conditions promoting robust matrix production [47] [48].
- Carefully aspirate planktonic cells and rinse biofilm gently with PBS to remove loosely attached cells.
- Prepare matrix disruption cocktail:
- Apply 1 mL disruption cocktail per well and incubate for 2-4 hours at 37°C with gentle agitation (50-100 rpm).
- Remove disruption cocktail and rinse briefly with PBS to terminate reaction.
- Proceed immediately to CRISPR-Cas13 application (Protocol 3).
Validation: Assess disruption efficacy via confocal microscopy with component-specific staining (Sypro Ruby for proteins, ConA for polysaccharides, TOTO-1 for eDNA) and quantify occupied area reduction using image analysis software [47].
Protocol 2: Guide RNA Modification for Enhanced Stability
Objective: To engineer chemically modified crRNAs with improved stability and longevity in the biofilm microenvironment.
Materials:
- Synthetic crRNA with target-specific spacer sequence
- 2'-O-methyl ribose modifications
- 3'-end stabilization chemistries (inverted dT, C3 spacers)
- RNAse-free water and buffers
- Purification columns for modified RNA
Procedure:
- Design crRNA spacer sequences complementary to target RNA in biofilm metabolic pathways (e.g., quorum sensing regulators, central metabolism enzymes) [13].
- Incorporate chemical modifications during synthetic crRNA production:
- Purify modified crRNAs using standard RNA cleanup protocols.
- Validate modification efficacy through:
- Nuclease protection assays comparing modified vs. unmodified crRNA half-life
- Thermal stability measurements
- Functional assessment in ribonucleoprotein (RNP) complexes with Cas13
Optimization Notes: Co-delivery of modified crRNAs with recombinant Cas13 enzymes in RNP complexes enhances protection from degradation, leading to significantly improved knockdown efficiency (60-65% with modified vs. 40-45% with unmodified crRNAs) [46].
Protocol 3: CRISPR-Cas13 Formulation and Application
Objective: To formulate CRISPR-Cas13 for optimal penetration and activity within pre-treated biofilms.
Materials:
- Purified Cas13d protein (or other Cas13 variants)
- Chemically modified crRNAs (from Protocol 2)
- Lipid nanoparticles (LNPs) or extracellular vesicle (EV) delivery systems
- Penetration enhancers (e.g., chitosan, ethylene-diamine-tetraacetic acid)
- Appropriate delivery buffer
Procedure:
- Prepare Cas13 RNP complexes:
- Combine Cas13 protein and modified crRNA at 3:1 molar ratio in delivery buffer
- Incubate at room temperature for 15-30 minutes to allow complex formation
- Formulate delivery vehicle (select one approach):
- LNP formulation: Encapsulate RNP complexes using microfluidic mixing with ionizable lipids, phospholipids, cholesterol, and PEG-lipid [13]
- EV loading: Electroporation or sonication of pre-formed extracellular vesicles with RNP complexes [13]
- Nebulizer-compatible nanoformulations: Prepare stable RNA complexes for aerosol delivery [13]
- Add penetration enhancers to final formulation:
- Chitosan (0.1-0.5% w/v) for mucosal adhesion
- EDTA (0.5-2 mM) to chelate matrix-stabilizing cations
- Apply 500 µL formulated CRISPR-Cas13 to pre-treated biofilms in 24-well plates.
- Incubate for 4-24 hours at relevant temperature (e.g., 37°C for human pathogens).
- Assess delivery efficacy via:
- Fluorescence in situ hybridization for target RNA reduction
- Quantitative PCR for transcript levels
- Metabolic activity assays (e.g., resazurin reduction)
- Phenotypic assessment of biofilm dispersal or antibiotic susceptibility
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for CRISPR-Cas13 Delivery in Biofilm Research
| Reagent/Category | Specific Examples | Function in Delivery Protocol | Key References |
|---|---|---|---|
| Matrix Disrupting Agents | Tranexamic acid (10 mg/mL), Dispersin B, DNase I | Degrades specific EPS components; increases porosity and diffusion | [47] [43] |
| Quorum Sensing Inhibitors | 5-hydroxymethylfurfural, Piper betle extract, synthetic QS peptides | Disrupts cell-cell signaling and matrix production regulation | [43] |
| CRISPR-Cas13 Systems | Cas13d RNP complexes, AAV-Cas13d constructs | RNA-targeting effector machinery; catalytic and binding functions | [49] [13] |
| Modified Guide RNAs | 3'-end 2'-O-methylated crRNAs, hairpin-extended spacers | Enhanced nuclease resistance; improved specificity and longevity | [46] [13] |
| Delivery Vehicles | Lipid nanoparticles (LNPs), Extracellular vesicles (EVs), Adeno-associated viruses (AAVs) | Protects CRISPR components; facilitates cellular uptake and penetration | [13] |
| Penetration Enhancers | Chitosan, EDTA, Triton-X 100 | Temporarily disrupts matrix integrity; improves diffusion kinetics | [48] [44] |
Metabolic Pathway Targeting Strategies
The strategic application of CRISPR-Cas13 to biofilm metabolic pathways requires careful selection of targets based on their regulatory importance and accessibility. The following diagram illustrates key targeting nodes within biofilm metabolic networks.
Critical target pathways include:
- Quorum sensing networks: Targeting autoinducer synthases (e.g., lasI, rhlI in Pseudomonas) and response regulators to disrupt cell-cell communication and coordinated behaviors [43] [44].
- Central carbon metabolism: Knocking down key glycolytic and TCA cycle enzymes to reduce energy production necessary for matrix synthesis and maintenance [49].
- Stress response systems: Modulating oxidative stress defense mechanisms that promote biofilm resilience under antimicrobial pressure.
- Toxin-antitoxin modules: Targeting persister cell formation by regulating toxin expression (e.g., MazF, RelE) that induces dormancy [44].
- EPS biosynthesis enzymes: Directly disrupting matrix production by knocking down polysaccharide synthesis and export machinery.
The multiplexing capability of CRISPR-Cas13 systems enables simultaneous targeting of multiple pathway components, potentially overcoming redundant regulatory networks in mature biofilms [49] [13].
Concluding Remarks
The integration of matrix-disruption strategies with advanced CRISPR-Cas13 delivery systems represents a promising approach to overcome the fundamental challenge of delivery inefficiency in dense biofilm matrices. The protocols and reagents outlined herein provide a framework for researchers to develop targeted interventions against biofilm-specific metabolic pathways, with potential applications ranging from biomedical device infections to industrial biofilm control. As CRISPR-Cas13 technology continues to evolve with improved specificity, novel delivery vehicles, and expanded editing capabilities, its utility in biofilm research and therapeutic development will undoubtedly increase. Future directions should focus on in vivo validation of these approaches, development of species-specific delivery systems, and exploration of combinatorial strategies that integrate RNA targeting with conventional antimicrobial agents for enhanced efficacy against persistent biofilm-associated infections.
Mitigating Collateral RNase Activity and Off-Target Effects
The CRISPR-Cas13 system has emerged as a powerful tool for programmable RNA targeting in biofilm research, yet its application is constrained by two significant challenges: collateral RNase activity and off-target effects. Unlike DNA-targeting CRISPR systems, Cas13 exhibits indiscriminate RNase activity upon target recognition, leading to widespread degradation of non-target RNAs [2]. This collateral effect, combined with sequence-based off-targeting, can confound experimental results in biofilm metabolic pathway studies, potentially triggering cellular stress responses and compromising viability [50] [51]. This application note details evidence-based strategies to mitigate these challenges, enabling more precise interrogation of biofilm RNA networks.
Understanding the Mechanisms
Collateral RNase Activity
Cas13 proteins belong to the Class 2, Type VI CRISPR systems and function as RNA-guided RNA endonucleases [2]. They possess two distinct RNase activities: one for processing its own guide RNA and another for cleaving target RNA, both mediated by Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains [2]. Upon recognition and binding to a target RNA sequence, Cas13 undergoes a conformational change that activates a non-specific RNase, leading to widespread degradation of nearby bystander RNA molecules [51].
The extent of collateral damage is directly correlated with target RNA abundance. Targeting highly expressed transcripts can result in global transcriptome destruction, with studies demonstrating up to 46% reduction in total RNA levels, ribosomal RNA fragmentation, and significantly compromised RNA integrity [51]. This non-specific RNA degradation can lead to chromatin collapse, reduced DNA replication, and impaired cell proliferation, presenting substantial challenges for metabolic pathway studies in biofilms [51].
Off-Target Effects
CRISPR-Cas13 off-target effects primarily occur through two mechanisms:
- Sequence-based off-targeting: Cas13 can tolerate mismatches between the guide RNA and target sequence, particularly outside the seed region, leading to binding and cleavage of partially complementary RNAs [52]
- Guide-independent effects: The activated collateral RNase degrades RNAs regardless of sequence similarity to the intended target [51]
These effects are particularly problematic in biofilm metabolic pathway research, where precise modulation of specific pathway components is essential for accurate functional analysis.
Quantitative Assessment of Effects
Table 1: Quantifying Collateral Activity and Off-Target Effects of RfxCas13d in Human Cells
| Target Condition | Target Abundance | Transcriptome Reduction | Cell Proliferation Impact | Mitochondrial RNA Impact |
|---|---|---|---|---|
| High-abundance target | ~100% of GAPDH | 46% median reduction | Strong inhibition (65-80% reduction) | Minimal (14% median reduction) |
| Medium-abundance target | ~10% of GAPDH | 15% median reduction | Moderate inhibition | Minimal |
| Low-abundance target | ~1% of GAPDH | 8% median reduction | No significant impact | Minimal |
| Non-targeting control | N/A | No significant reduction | No impact | No impact |
Source: Adapted from Kelley et al. (2022) and Zhang et al. (2023) [50] [51]
Table 2: Comparison of Cas13 Subtypes and Their Characteristics
| Cas13 Subtype | Size (amino acids) | Guide RNA Placement | Knockdown Efficiency | Reported Collateral Activity |
|---|---|---|---|---|
| Cas13a | 1000-1200 | 3' end | Moderate | Strong in vitro, variable in cells |
| Cas13b | 1000-1200 | 5' end | Moderate | Strong in vitro, variable in cells |
| Cas13d (CasRx) | 1000-1200 | Variable | High | Strong in mammalian cells |
| Cas13X/Y | 775-800 | Variable | Comparable to Cas13d | Not fully characterized |
Source: Adapted from Abudayyeh et al. (2016), Konermann et al. (2018), and Xu et al. (2021) [2]
Mitigation Strategies and Protocols
Target Selection and Expression Considerations
Protocol: Assessment of Target Suitability
- Determine target abundance: Quantify expression levels of intended target RNA via RNA-seq or qPCR prior to experiment design
- Establish expression threshold: Avoid targeting transcripts exceeding 10% of GAPDH expression level when possible [51]
- Implement controls: Include both non-targeting guides and guides targeting low-abundance RNAs as negative controls
- Utilize spike-in controls: Add exogenous RNA (e.g., from mouse cells) to enable normalization and detection of global transcriptome effects [51]
Rationale: Target abundance strongly correlates with collateral activity severity. Targeting low-abundance transcripts (<1% of GAPDH) minimizes non-specific effects while maintaining efficient on-target knockdown [51].
Engineering Solutions
Protocol: Implementation of GENO (Guide RNA Excision and Negative Autoregulation) System
- Vector design: Clone RfxCas13d expression cassette with negative autoregulatory features
- Guide RNA engineering: Incorporate self-targeting guides that limit Cas13 expression after initial target engagement
- Delivery optimization: Utilize AAV-compatible vectors for efficient delivery while maintaining regulation [50]
Materials:
- GENO-compatible RfxCas13d plasmid
- Tissue-specific promoter (as appropriate for biofilm model)
- Self-targeting guide sequences
- AAV packaging system (if needed)
Procedure:
- Clone RfxCas13d under control of inducible promoter
- Incorporate self-targeting guide RNA expression cassette
- Transfect biofilm model system and induce expression
- Monitor Cas13 levels over time to confirm autoregulation
- Validate target engagement and assess collateral effects [50]
Troubleshooting: If autoregulation insufficient, modify guide RNA efficacy or incorporate additional self-targeting guides. Monitor cell viability throughout experiment.
Computational and Guide Design Approaches
Protocol: Machine Learning-Guided Guide Design Using DeepCas13
- Input preparation: Compile candidate guide RNA sequences (20-27 nucleotides) targeting desired RNA
- Secondary structure prediction: Generate RNA folding predictions for each guide
- Model application: Input guide sequences and structures into DeepCas13 platform
- Guide selection: Prioritize guides with high predicted on-target efficiency and low off-target potential [52]
Access: DeepCas13 is freely accessible at http://deepcas13.weililab.org/
Key considerations:
- Select guides with minimal similarity to non-target transcripts, especially in seed region
- Avoid regions with stable secondary structure that may impede binding
- Consider tiling multiple guides across target for improved efficacy [52]
Experimental Design Considerations
Protocol: Optimization of Expression and Delivery Conditions
Moderate Cas13 expression:
- Utilize low copy number vectors
- Implement tightly regulated promoters
- Incorporate weak ribosome binding sites [2]
Limit exposure duration:
- Employ transient transfection rather than stable expression
- Use inducible systems with short induction windows
- Monitor expression kinetics to determine optimal harvest time
Validation controls:
- Include siRNA knockdown controls to distinguish specific from collateral effects
- Implement multiple guide RNAs per target to control for guide-specific artifacts
- Utilize orthogonal validation methods (e.g., RNA FISH, qPCR) [51]
The Scientist's Toolkit
Table 3: Essential Research Reagents for Cas13 Studies in Biofilm Research
| Reagent/Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| Cas13 Effectors | RfxCas13d (CasRx), PspCas13b, PguCas13b | RNA-targeting CRISPR effector | Cas13d shows high efficiency; Cas13b useful for imaging |
| Guide RNA Cloning Systems | pCRISPR-Cas13b/d backbones (Addgene) | Express sequence-specific guide RNAs | Ensure proper scaffold compatibility with chosen Cas13 |
| Computational Tools | DeepCas13, CRISPOR | Guide design and efficiency prediction | DeepCas13 incorporates secondary structure predictions |
| Control Plasmids | Non-targeting guides, GFP-targeting guides | Experimental controls | Essential for distinguishing specific vs. collateral effects |
| Detection Reagents | Spike-in RNA controls, RNA integrity assays | Monitor collateral activity | Spike-ins enable normalization for global transcript effects |
| Delivery Systems | AAV vectors, lipofectamine, nanocarriers | Introduce CRISPR components | AAV compatible with GENO system; consider transient delivery |
| Validation Tools | RNA-FISH probes, qPCR assays | Confirm on-target effects | Orthogonal validation crucial for interpreting results |
Application to Biofilm Metabolic Pathways
When applying Cas13 to biofilm metabolic pathway research, several specific considerations apply:
- Pathway redundancy: Target multiple pathway components simultaneously to overcome metabolic redundancy
- Temporal effects: Consider timing of intervention relative to biofilm development stage
- Spatial considerations: Account for metabolic heterogeneity within biofilm structures
Protocol: Biofilm-Specific Application
- Identify key metabolic nodes: Pinpoint pathway components with minimal redundancy
- Design guide repertoire: Create 4-6 high-scoring guides per target using DeepCas13
- Establish dose response: Titrate Cas13 expression to minimum effective level
- Monitor metabolic output: Measure pathway-specific metabolites in addition to transcript levels
- Validate with orthogonal approach: Confirm findings with chemical inhibitors or RNAi where possible
Mitigating collateral RNase activity and off-target effects is essential for employing CRISPR-Cas13 in biofilm metabolic pathway research. The integrated strategies presented here—including careful target selection, engineered regulation systems, computational guide design, and appropriate experimental controls—enable more precise and interpretable results. As Cas13 engineering advances, emerging solutions such as high-fidelity variants with reduced collateral activity and improved computational prediction tools will further enhance the precision of RNA-targeting applications in complex biofilm systems. Implementation of these mitigation strategies will enable researchers to harness the power of CRISPR-Cas13 while minimizing confounding effects, ultimately advancing our understanding of biofilm metabolic pathways.
Workflow for Mitigating Cas13 Effects - This diagram outlines the key steps for minimizing collateral activity and off-target effects in biofilm research applications.
GENO System vs Standard Cas13 - Comparison of the negative autoregulation system (GENO) that limits collateral damage versus conventional Cas13 expression.
Utilizing Chemically Modified Guide RNAs to Enhance Stability and Knockdown Efficiency
CRISPR-Cas13 has emerged as a powerful RNA-targeting platform for modulating gene expression outcomes in research and therapeutic development. Unlike DNA-editing CRISPR systems, Cas13 targets RNA transcripts, making it particularly valuable for investigating transient metabolic pathways in bacterial biofilms and for therapeutic applications where permanent genomic changes are undesirable [53] [54]. However, the efficacy of this technology is limited by the rapid degradation of natural guide RNAs by endogenous RNA nucleases in biological systems, leading to transient knockdown effects [46]. Recent advances demonstrate that strategic chemical modifications to CRISPR RNAs (crRNAs) can significantly enhance their stability and knockdown efficiency, opening new possibilities for targeting essential metabolic pathways in biofilm-forming bacteria [53] [54].
In the context of biofilm research, CRISPR-Cas13 enables precise dissection of complex regulatory networks controlling extracellular polymeric substance (EPS) production, quorum sensing, and stress adaptation mechanisms [8]. The implementation of chemically modified guides addresses critical delivery challenges in EPS-rich biofilm matrices, where nuclease activity and physical barriers traditionally limit intervention efficacy. This protocol details the application of stabilized crRNAs for investigating biofilm metabolic pathways, providing researchers with tools for sustained transcript knockdown in complex microbial communities [8] [11].
Chemical Modification Strategies for Cas13 Guide RNAs
Modification Patterns and Positions
The strategic placement of chemical modifications on Cas13 crRNAs is critical for enhancing nuclease resistance while maintaining guide activity. Research indicates that modifications at the 3' end of synthesized crRNAs significantly boost transcript knockdown compared to unmodified crRNAs, with particularly strong effects when modifications are placed within the spacer sequence [46]. Specific modification patterns validated in human cells include:
- 2'-O-methylation (2'-O-Me): Particularly effective at the 3' end bases
- Phosphorothioate (PS) linkages: Backbone modifications that enhance nuclease resistance
- 2'-O-methyl-3'-phosphorothioate (MS): Combined sugar and backbone modifications
- 2'-fluoro (2'-F): Ribose modifications that stabilize RNA structure
Notably, modifications at the 5' end, whether alone or in combination with 3' changes, do not consistently improve knockdown efficiency, indicating positional specificity in modification efficacy [53] [54]. The optimal configuration identified through systematic screening incorporates 2'-O-methylation at three terminal bases on the 3' end of the spacer sequence.
Quantitative Comparison of Modification Efficacy
Table 1: Comparison of crRNA Modification Effects on Knockdown Efficiency
| Modification Type | Position | Knockdown Efficiency | Half-Life Extension | Optimal Use Cases |
|---|---|---|---|---|
| 2'-O-methylation (2'-O-Me) | 3' end spacer | 60-65% | ~2-fold | Standard biofilm applications |
| Unmodified crRNA | N/A | 40-45% | Baseline | Control comparisons |
| Phosphorothioate (PS) | Terminal linkages | 55-60% | ~1.8-fold | High-nuclease environments |
| 2'-fluoro (2'-F) | Internal bases | 50-55% | ~1.5-fold | Specialized applications |
| Combined 2'-O-Me + PS | 3' end + backbone | 65-70% | ~2.5-fold | Challenging biofilm matrices |
The data in Table 1 summarizes performance metrics for different modification strategies, demonstrating that 3'-modified crRNAs can improve knockdown efficiency by 15-25 percentage points compared to unmodified guides [53]. This enhancement is particularly valuable in biofilm systems where efficient transcript targeting is essential for dissecting redundant metabolic pathways.
Experimental Protocol for Modified Guide RNA Implementation
Ribonucleoprotein (RNP) Complex Assembly
The delivery of Cas13 RNP complexes with pre-assembled modified crRNAs represents the most efficient approach for achieving robust transcript knockdown in biofilm systems.
Materials Required:
- Chemically synthesized modified crRNAs (3' 2'-O-methylated)
- Recombinant Cas13 protein (LwaCas13a, PspCas13b, or RfxCas13d)
- Nuclease-free buffer (20 mM HEPES, 150 mM KCl, pH 7.5)
- RNase inhibitor
- Target bacterial strains and biofilm growth media
Procedure:
- Complex Formation:
- Resuspend modified crRNA in nuclease-free buffer to 100 μM stock concentration
- Combine 2 μg recombinant Cas13 protein with 5 μL crRNA stock (molar ratio 1:2.5)
- Add 1 μL RNase inhibitor (40 U/μL)
- Incubate at 25°C for 15 minutes to form RNP complexes
Biofilm Treatment:
- Grow biofilms to mid-exponential phase (OD600 = 0.4-0.6) in appropriate media
- For suspended biofilms: Add RNP complexes directly to culture medium at 1:100 dilution
- For surface-adhered biofilms: Apply RNP complexes in fresh medium replacement
- Incubate at optimal growth temperature for 4-24 hours depending on application
Validation and Analysis:
- Harvest cells at appropriate time points for transcript analysis
- Extract total RNA using standard methodologies
- Quantify target transcript levels via RT-qPCR
- Assess functional consequences through metabolic assays or phenotypic readouts
The RNP delivery approach protects crRNAs from degradation and enables immediate activity upon cellular entry, resulting in more rapid and sustained knockdown compared to plasmid-based expression systems [53] [54].
Evaluation of Knockdown Efficiency in Biofilm Models
Protocol for Biofilm Metabolic Pathway Targeting:
Strain Preparation:
- Culture biofilm-forming bacteria (e.g., Pseudomonas aeruginosa, Escherichia coli) to early exponential phase
- Standardize inoculum to 10^6 CFU/mL in appropriate biofilm-promoting media
Biofilm Establishment:
- Transfer 1 mL bacterial suspension to biofilm-compatible surfaces (e.g., peg lids, silicone coupons)
- Incubate for 24 hours to establish mature biofilms with EPS production
CRISPR-Cas13 Intervention:
- Prepare RNP complexes with crRNAs targeting key metabolic pathway genes:
- Quorum sensing regulators (e.g., lasI, rhlI)
- EPS biosynthesis genes (e.g., psl, pel, alg)
- Central metabolic enzymes
- Apply RNP complexes in fresh media replacement
- Include controls with unmodified crRNAs and non-targeting guides
- Prepare RNP complexes with crRNAs targeting key metabolic pathway genes:
Assessment Time Course:
- Monitor transcript knockdown at 2, 6, 12, and 24 hours post-treatment
- Evaluate biofilm biomass through crystal violet staining at 24 and 48 hours
- Assess metabolic activity via resazurin reduction assays
- For persistent biofilms, consider secondary RNP application at 24 hours
This protocol enables systematic evaluation of how targeting specific transcripts affects biofilm development, maintenance, and metabolic activity, providing insights into pathway essentiality and potential therapeutic targets [8].
Research Reagent Solutions
Table 2: Essential Reagents for Modified Guide RNA Experiments
| Reagent/Catalog | Supplier Examples | Function/Application | Usage Notes |
|---|---|---|---|
| Chemically Modified crRNAs | Synthego, IDT, Dharmacon | Target-specific guide RNA with enhanced stability | Specify 3' 2'-O-methyl modifications |
| Recombinant Cas13 Proteins | BioVision, ABM, GenScript | RNA-targeting effector protein | LwaCas13a for general use; RfxCas13d for compact size |
| RNase Inhibitor | NEB, Thermo Fisher | Protect RNA components during handling | Essential for RNP complex assembly |
| Biofilm Matrix Reagents | Sigma-Aldrich, Millipore | Crystal violet, resazurin for assessment | Standardized biofilm quantification |
| Bacterial Strain Libraries | ATCC, BEI Resources | Relevant biofilm-forming pathogens | P. aeruginosa, E. coli, S. aureus |
| Nucleic Acid Extraction Kits | Qiagen, Zymo Research | RNA isolation from biofilm matrices | Include DNase treatment steps |
Workflow and Pathway Diagrams
Applications in Biofilm Metabolic Pathway Research
The implementation of chemically modified guide RNAs for CRISPR-Cas13 enables previously challenging investigations into biofilm metabolic pathways. The enhanced stability of modified crRNAs supports extended observation windows necessary for understanding dynamic processes in biofilm development and maintenance [8] [11].
Key Research Applications:
Functional Analysis of Essential Genes:
- Target transcripts encoding enzymes in central metabolic pathways
- Assess essentiality through metabolic flux analysis during knockdown
- Identify vulnerable nodes for potential therapeutic intervention
Quorum Sensing Network Dissection:
- Sequentially target components of cell-cell communication systems
- Map regulatory hierarchies through temporal knockdown studies
- Identify compensatory mechanisms and network redundancies
EPS Matrix Engineering:
- Modulate expression of biosynthetic enzymes for polysaccharide components
- Investigate structure-function relationships in matrix composition
- Explore potential for targeted matrix disruption
The quantitative nature of CRISPR-Cas13 targeting with modified guides enables dose-response studies where transcript reduction levels can be correlated with phenotypic effects. This precision is particularly valuable for investigating metabolic pathways that may exhibit threshold effects or complex regulation [8] [53].
Troubleshooting and Optimization Guidelines
Common Challenges and Solutions:
Insufficient Knockdown Efficiency:
- Verify crRNA modification quality through mass spectrometry
- Optimize RNP complex ratios (typically 1:2 to 1:5 Cas13:crRNA)
- Consider alternative Cas13 orthologs with different PFS requirements
Variable Effects Across Biofilm Models:
- Account for differential penetration in dense biofilm structures
- Optimize delivery timing relative to biofilm developmental stage
- Consider adjuvant compounds to enhance RNP penetration
Off-target Transcript Effects:
- Validate target specificity through RNA-seq in pilot studies
- Design multiple crRNAs against different regions of target transcript
- Implement appropriate controls including non-targeting crRNAs
The integration of chemically modified guide RNAs with advanced delivery platforms, including nanoparticle-based systems, represents the next frontier for CRISPR-Cas13 applications in biofilm research [11]. These hybrid approaches may further enhance the stability and efficacy of RNA-targeting interventions in complex microbial communities.
The application of the CRISPR-Cas13 system for targeting RNA within bacterial biofilm metabolic pathways represents a transformative approach in microbiological research and therapeutic development. Unlike DNA-targeting CRISPR systems, Cas13 specifically interrogates and manipulates the transcriptome, offering unique advantages for dissecting dynamic metabolic processes in biofilm communities without altering the underlying genome [55] [17]. The effectiveness of Cas13-based interventions is profoundly influenced by the precision of its expression, which is governed by promoter selection and regulatory control systems. Proper expression optimization ensures sufficient Cas13 activity for target engagement while minimizing off-target effects and cellular toxicity, which is particularly crucial when investigating essential metabolic pathways in structured biofilm environments [8].
This application note provides a structured framework for selecting promoter systems and implementing regulatory controls to achieve precise Cas13 expression for biofilm metabolic pathway research. We integrate quantitative performance data, detailed methodological protocols, and strategic considerations to guide researchers in designing robust experimental systems for transcriptome-level investigation of biofilm physiology.
Promoter Systems for Cas13 Expression
Selection of an appropriate promoter is fundamental to achieving desired Cas13 expression levels and kinetics. The choice depends on the specific application, whether for high-throughput screening, transient knockdown studies, or long-term metabolic pathway analysis. The table below summarizes the key characteristics of commonly used promoter systems for Cas13 expression.
Table 1: Comparison of Promoter Systems for Cas13 Expression
| Promoter Type | Induction Mechanism | Expression Kinetics | Leakiness | Best Applications |
|---|---|---|---|---|
| T7 Polymerase | IPTG-inducible | Very strong, rapid onset (hours) | Low with proper engineering | High-level expression for diagnostics and screening [56] |
| J23100 (Constitutive) | N/A | Medium, constant | High | Reference standardization, metabolic flux analysis [8] |
| pLuxI (QS-regulated) | AHL autoinducer | Population-density dependent | Variable | Biofilm development studies, pathway timing [8] |
| pTet | Anhydrotetracycline | Tightly regulated, dose-dependent | Very low | Precision knockdowns in essential pathways [8] [55] |
| pBAD | L-Arabinose | Titratable, moderate strength | Low to moderate | Metabolic engineering, fine-tuning expression [8] |
For biofilm metabolic pathway research, inducible systems offer significant advantages. The pTet system provides particularly tight control with minimal basal expression, enabling researchers to initiate Cas13 expression at specific biofilm developmental stages. This temporal precision is valuable when investigating how metabolic shifts influence transitions from planktonic to biofilm states or during biofilm dispersal events [8].
Regulatory Control Elements
Beyond promoter selection, incorporating additional regulatory elements enhances expression precision and system safety, especially for extended biofilm studies.
Table 2: Key Regulatory Control Elements for Precision Expression
| Control Element | Function | Considerations for Biofilm Research |
|---|---|---|
| Kill Switches | Terminate expression after set duration | Prevents prolonged metabolic disturbance; essential for in vivo models |
| T7 RNA Polymerase | Enables T7 promoter usage in prokaryotes | Increases portability across biofilm model systems |
| sgRNA Scaffold Optimization | Modulates Cas13 binding and cleavage efficiency | Affects mismatch discrimination; critical for homologous metabolic genes [17] |
| Terminator Sequences | Preforms transcriptional read-through | Insulates Cas13 expression from polar effects in metabolic operons |
Research Reagent Solutions
The following table outlines essential reagents and their specific functions for establishing Cas13 expression systems for biofilm metabolic pathway research.
Table 3: Essential Research Reagents for Cas13-Biofilm Studies
| Reagent/Category | Specific Function | Application Notes |
|---|---|---|
| Cas13 Orthologs | RNA targeting and cleavage | LwaCas13a, LbuCas13a, RfxCas13d show varying efficacy against structured RNA targets [17] |
| Delivery Vectors | Cas13 and guide RNA transport to biofilm cells | Phagemids, conjugative plasmids, or nanocarriers selected based on target species [8] [12] |
| Fluorescent Reporters | Visualize Cas13 activity and localization | RNA-based reporters with stem-loop structures for monitoring activation kinetics [17] |
| Occluded crRNAs | Enhanced specificity for single-nucleotide discrimination | Engineered crRNAs that improve mismatch detection in conserved metabolic genes [17] |
| Nanoparticle Delivery Systems | Enhance penetration through biofilm matrix | Liposomal or gold nanoparticles improve delivery efficiency up to 3.5-fold [12] |
Experimental Protocol: Promoter Efficiency Testing
This protocol provides a standardized method for evaluating and comparing promoter efficiency for Cas13 expression in biofilm-relevant conditions.
Materials
- Plasmids: Cas13 expression vectors with candidate promoters (e.g., pTet, pBAD, T7)
- Bacterial Strains: Target biofilm-forming strain (e.g., Pseudomonas aeruginosa, Escherichia coli)
- Inducers: IPTG (0.1-1mM), anhydrotetracycline (10-200 ng/mL), L-Arabinose (0.001-0.2%)
- Growth Media: Appropriate broth with selection antibiotics
- Detection Reagents: RNA extraction kits, qRT-PCR reagents, fluorescence plate reader
Procedure
Day 1: Strain Preparation
- Transform candidate promoter-Cas13 plasmids into target bacterial strain using standard transformation protocols.
- Plate transformed cells on selective media and incubate overnight at appropriate temperature (e.g., 37°C for most pathogens).
Day 2: Inoculation and Induction
- Pick 3-5 colonies from each transformation and inoculate 5 mL of selective broth.
- Grow cultures to mid-log phase (OD600 ≈ 0.4-0.6).
- Split each culture into two aliquots:
- Uninduced control (no inducer)
- Induced test (with appropriate inducer at mid-range concentration)
- Incubate with shaking for precisely 6 hours post-induction.
Day 2: Sample Collection and Analysis
- Collect 1 mL from each culture for RNA extraction.
- Extract total RNA using commercial kit, including DNase treatment step.
- Perform qRT-PCR using Cas13-specific primers and normalize to housekeeping gene (e.g., rpoB or gyrA).
- For fluorescence-based reporters, measure fluorescence intensity (ex/em appropriate for reporter).
- Calculate fold-induction for each promoter (induced/uninduced).
Day 3: Biofilm Assay
- For selected top performers, repeat induction in biofilm culture using peg lid or flow cell system.
- After 6 hours induction, harvest biofilm cells and quantify Cas13 expression as above.
- Compare planktonic versus biofilm expression ratios.
Data Interpretation
- Optimal Promoter Selection: Choose promoters with high induction ratio (>50-fold), low leakiness (<1% of induced), and consistent performance in biofilm conditions.
- Troubleshooting: High basal expression may require additional genetic insulation (strong terminators) or modified ribosome binding sites. Poor induction may necessitate inducer concentration optimization.
Experimental Protocol: crRNA Design for Structured RNA Targets
Metabolic pathway mRNAs often contain complex secondary structures that can impede Cas13 access. This protocol outlines a bioinformatic and experimental approach for designing effective crRNAs against structured RNA targets in biofilm metabolic genes.
Materials
- Software: NUPACK, RNAfold, or similar RNA structure prediction tools
- Target Sequences: mRNA sequences of biofilm metabolic genes (e.g., pelA, pslG, algC for Pseudomonas; icaA, icaD for Staphylococcus)
- crRNA Synthesis: Commercial synthesis or in vitro transcription kit
- Validation System: Purified Cas13 protein, fluorescent RNA reporter
Procedure
Step 1: Target Sequence Analysis
- Obtain full-length mRNA sequence of target metabolic gene from databases (NCBI, UniProt).
- Identify conserved regions across bacterial strains using multiple sequence alignment (Clustal Omega, MUSCLE).
- Input target sequence into structure prediction software (NUPACK recommended) to identify unstructured regions.
Step 2: crRNA Design
- Design 3-5 crRNA candidates targeting regions with:
- Minimal secondary structure (ΔG > -5 kcal/mol)
- High sequence conservation (for broad application)
- Avoidance of known single-nucleotide polymorphisms
- Incorporate strategic mismatches in seed region (positions 5-8) if enhanced specificity required [17].
- For highly structured targets, design "occluded crRNAs" with 5' extensions that compete with native structure.
Step 3: In Vitro Validation
- Synthesize candidate crRNAs (commercial synthesis recommended for modified bases).
- Set up Cas13 cleavage reactions with:
- 50 nM purified Cas13 protein
- 50 nM crRNA
- 100 nM target RNA transcript
- 500 nM fluorescent RNA reporter
- Reaction buffer (20 mM HEPES, 60 mM NaCl, 6 mM MgCl₂, pH 6.8)
- Measure fluorescence every 30 seconds for 2 hours at 37°C.
- Calculate cleavage rates and compare to positive and negative controls.
Step 4: Cellular Validation
- Clone validated crRNAs into appropriate expression vector.
- Transform into target strain with Cas13 expression system.
- Measure target mRNA knockdown using qRT-PCR after induction.
- Assess metabolic consequences through targeted metabolomics or phenotype assays.
Workflow Visualization
Optimization Workflow for Cas13 Expression Systems
RNA Structure Effects on Cas13 Function
Precise control of Cas13 expression through strategic promoter selection and regulatory engineering is fundamental to successful investigation of biofilm metabolic pathways. The frameworks, protocols, and reagents outlined in this application note provide a systematic approach for researchers to develop optimized Cas13 systems tailored to their specific biofilm models and metabolic targets. As Cas13 technology continues to evolve, incorporating emerging innovations in RNA targeting and delivery will further enhance our ability to precisely manipulate and understand the metabolic networks that underpin biofilm formation and persistence.
The escalating crisis of antimicrobial resistance (AMR) represents one of the most pressing global health challenges of our time, with projections indicating that AMR may cause up to 10 million fatalities annually by 2050 if new antimicrobial strategies are not developed [57] [58]. Biofilm-associated infections are particularly problematic in clinical settings, as the extracellular polymeric substance (EPS) matrix and heterogeneous metabolic states within biofilms can increase bacterial tolerance to antibiotics by up to 1000-fold compared to planktonic cells [11]. This recalcitrance necessitates innovative approaches that extend beyond conventional antibiotic monotherapies.
CRISPR-Cas13 systems have emerged as powerful programmable tools that target RNA rather than DNA, employing guide RNAs to direct the Cas13 nuclease to complementary RNA sequences [2] [58]. Upon target recognition, Cas13 exhibits collateral RNA cleavage activity that can lead to bacterial growth inhibition or death [58]. When deployed in combination with conventional antibiotics, CRISPR-Cas13 systems enable a synergistic, dual-mechanism attack: Cas13 precisely disrupts vital bacterial functions at the RNA level, while antibiotics concurrently target structural or metabolic pathways. This approach is particularly effective against biofilms, as Cas13 can be programmed to interfere with quorum sensing, stress response systems, and EPS production—key pathways supporting biofilm maintenance and antibiotic tolerance [8] [11].
Key Mechanistic Insights and Quantitative Synergy Data
Molecular Mechanisms of Synergy
The synergistic relationship between Cas13 and conventional antibiotics operates through several interconnected mechanisms. Cas13a achieves sequence-specific killing of target bacteria by recognizing and cleaving mRNA of antimicrobial resistance genes, regardless of whether these genes are located chromosomally or on plasmids [58]. This targeting capability fundamentally differs from CRISPR-Cas9 systems, which lack effective bacterial killing capacity when target genes are plasmid-borne [58]. By selectively disrupting resistance gene expression, Cas13 effectively resensitizes resistant bacteria to conventional antibiotics, restoring the efficacy of previously ineffective treatments.
At the molecular level, Cas13's collateral RNA degradation activity after target recognition creates widespread disruption of bacterial metabolic processes [2] [58]. This disruption particularly affects biofilm maintenance pathways, including quorum sensing systems that coordinate community behavior and stress response networks that enhance bacterial survival under antibiotic pressure [8]. When administered concurrently, antibiotics can more effectively penetrate and act upon bacterial cells whose defensive capabilities have been compromised by Cas13-mediated RNA interference.
Quantitative Assessment of Synergistic Effects
Table 1: Quantitative Efficacy of Cas13-Antibiotic Combinations Against Biofilms
| Pathogen | Resistance Profile | Cas13 Target | Combination Partner | Efficacy Outcome | Reference |
|---|---|---|---|---|---|
| E. coli | Carbapenem-resistant (blaIMP-1+) | blaIMP-1 mRNA | Carbapenems | 2-3 log reduction in bacterial counts; effective against chromosomal and plasmid-borne genes | [58] |
| S. aureus | Methicillin-resistant (MRSA) | mecA mRNA | β-lactam antibiotics | Significant restoration of β-lactam susceptibility; enhanced biofilm disruption | [57] [58] |
| P. aeruginosa | Multi-drug resistant | Quorum sensing networks (lasR, rhlR) | Tobramycin | >90% reduction in biofilm biomass with liposomal Cas13 delivery | [11] |
| Multiple Gram-negative pathogens | Carbapenem-resistant | blaNDM-1, blaKPC-2, blaOXA-48 | Multiple antibiotic classes | Sequence-specific killing demonstrated across major carbapenemase genes | [58] |
Research has demonstrated that liposomal Cas13a formulations can reduce Pseudomonas aeruginosa biofilm biomass by over 90% in vitro [11]. When combined with nanoparticle delivery systems, CRISPR-Cas13 constructs have shown editing efficiency enhancements of up to 3.5-fold compared to non-carrier systems, significantly improving the penetration of these therapeutic agents through biofilm matrices [11]. The combination approach has proven particularly effective against carbapenem-resistant Enterobacteriaceae, with Cas13a-based antimicrobials achieving reduction of bacterial counts by two to three orders of magnitude against bacteria carrying blaIMP-1, both on chromosomes and plasmids [58].
Experimental Protocols
Protocol 1: Cas13a-Antibiotic Synergy Screening in Biofilm Models
This protocol describes a standardized method for evaluating the synergistic effects of Cas13a and conventional antibiotics against bacterial biofilms, utilizing a microtiter plate format suitable for high-throughput screening.
Materials and Reagents:
- Cas13a protein (LshCas13a or equivalent)
- In vitro-transcribed guide RNAs targeting resistance or essential genes
- Cationic liposomal transfection reagent or phage capsid delivery system
- Mueller Hinton broth or appropriate bacterial growth medium
- 96-well polystyrene plates for biofilm formation
- SYTO 9/propidium iodide live/dead staining solution
- Crystal violet solution (0.1% for biofilm biomass quantification)
- Microplate reader with fluorescence and absorbance capabilities
Procedure:
- Biofilm Formation: Grow overnight cultures of target bacteria and dilute to 1×10^6 CFU/mL in fresh medium. Add 200 μL aliquots to 96-well plates and incubate for 24-48 hours at appropriate temperature to establish mature biofilms.
- Cas13a Delivery Preparation: Complex Cas13a ribonucleoproteins (100 nM) with guide RNAs (50 nM) targeting specific resistance genes (e.g., blaNDM-1, mecA) using liposomal transfection reagents according to manufacturer protocols. Alternatively, prepare phage capsid-packaged Cas13a constructs.
- Treatment Application: Carefully aspirate planktonic cells and medium from established biofilms. Add treatment groups:
- Group 1: Cas13a-gRNA complexes alone
- Group 2: Antibiotic alone (at sub-MIC concentrations)
- Group 3: Cas13a-gRNA + antibiotic combination
- Group 4: Non-targeting gRNA control + antibiotic
- Group 5: Untreated biofilm control
- Incubation and Assessment: Incubate plates for 6-24 hours at appropriate temperature. Assess biofilm viability using:
- Metabolic activity: MTT assay
- Biofilm biomass: Crystal violet staining (absorbance at 570 nm)
- Bacterial viability: Live/dead staining with fluorescence quantification
- Synergy Calculation: Determine synergistic effects using the Fractional Inhibitory Concentration Index (FICI) where FICI ≤0.5 indicates synergy.
Troubleshooting Notes:
- Optimize Cas13a:gRNA ratio for each bacterial strain (typically 2:1 to 1:1 molar ratio)
- For thick biofilms, extend treatment incubation times to 24-48 hours
- Include appropriate controls for collateral RNA cleavage effects
Protocol 2: Advanced Nanoparticle-Mediated Co-delivery System
This protocol details the formulation and application of hybrid nanoparticle systems for coordinated delivery of Cas13 components and antibiotics directly to biofilm environments.
Materials and Reagents:
- Cationic lipid nanoparticles (LNPs) or gold nanoparticles (AuNPs)
- Cas13a mRNA or recombinant protein
- Guide RNA constructs
- Antibiotics (tobramycin, ciprofloxacin, or imipenem)
- Diethylaminoethyl-dextran (DEAE-dextran) for enhanced biofilm penetration
- Dynamic light scattering (DLS) apparatus for nanoparticle characterization
Formulation Procedure:
- Nanoparticle Preparation: Prepare lipid nanoparticles using microfluidic mixing technology with ionizable cationic lipids, phospholipids, cholesterol, and PEG-lipid at molar ratios of 50:10:38.5:1.5.
- Complexation with CRISPR Components: Encapsulate Cas13a mRNA (0.5 mg/mL) and guide RNA (0.25 mg/mL) using a 5:1 nitrogen-to-phosphate ratio for efficient complexation.
- Antibiotic Loading: Adsorb antibiotics onto nanoparticle surfaces through electrostatic interactions or incorporate into outer lipid layers at 1-5% w/w ratio.
- Characterization: Determine nanoparticle size (target 100-200 nm), zeta potential (>+20 mV for biofilm penetration), and encapsulation efficiency using HPLC for antibiotic quantification.
- Quality Control: Validate Cas13a activity after encapsulation using in vitro RNA cleavage assays with synthetic target RNA sequences.
Application to Biofilms:
- Treat established biofilms in 96-well plates or flow cell systems with nanoparticle formulations containing Cas13a (50-100 nM) and antibiotic (1/4 to 1/2 MIC).
- Incubate for 4-24 hours with gentle agitation to maintain nanoparticle suspension.
- Assess biofilm disruption using confocal laser scanning microscopy with constitutive GFP-expressing bacterial strains.
- Quantify remaining viable bacteria through colony counting after biofilm dissociation via sonication.
Diagram 1: Mechanism of CRISPR-Cas13 and Antibiotic Synergy in Biofilm Elimination. This schematic illustrates the coordinated action of nanoparticle-delivered Cas13 and antibiotics against bacterial biofilms, highlighting the dual-mechanism approach that underlies their synergistic effect.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents for Cas13-Antibiotic Combination Studies
| Reagent/Category | Specific Examples | Function & Application Notes | Commercial Sources |
|---|---|---|---|
| Cas13 Variants | LshCas13a, RfxCas13d, Cas13X.1 | RNA-targeting effectors with varying PFS requirements; Cas13X.1 offers compact size for improved delivery | Addgene, Thermo Fisher, Sigma-Aldrich |
| Delivery Systems | Cationic LNPs, AuNPs, phage capsids, conjugative plasmids | Enhance biofilm penetration and cellular uptake; phage capsids provide species-specific targeting | Formulate in-house or custom-order from nanomedicine specialists |
| Guide RNA Design | Target-specific crRNAs, multiplexed arrays | Sequence-specific targeting of AMR genes (e.g., blaNDM-1, mecA) or essential pathways | Integrated DNA Technologies, Twist Bioscience |
| Biofilm Reactors | Calgary biofilm device, flow cell systems, 96-well peg lids | Generate standardized, reproducible biofilms for therapeutic testing | MBEC Biofilm, Ibidi GmbH, Nunc |
| Viability Assays | ATP bioluminescence, resazurin reduction, LIVE/DEAD BacLight | Quantify metabolic activity and membrane integrity in biofilm populations | Promega, Thermo Fisher, Molecular Probes |
| Antibiotic Panel | Tobramycin, ciprofloxacin, imipenem, colistin | Representative agents from major classes for combination screening | Sigma-Aldrich, TOKU-E |
Pathway Engineering and Target Selection
Effective implementation of Cas13-antibiotic combinations requires strategic selection of target pathways within bacterial biofilms. The most promising targets include:
Resistance Gene Targeting: Program Cas13 to cleave mRNA of clinically relevant resistance genes, including:
- Carbapenemase genes (blaKPC, blaNDM, blaOXA-48)
- Methicillin resistance gene (mecA)
- Colistin resistance genes (mcr-1, mcr-2)
- Extended-spectrum β-lactamase genes (CTX-M-15, TEM-52)
Biofilm Regulatory Networks: Target quorum sensing systems (lasR, rhlR in Pseudomonas; agr in Staphylococcus) to disrupt cell-to-cell communication essential for biofilm maturation and maintenance [8]. Additionally, target secondary messenger systems (c-di-GMP synthesis pathways) that control the transition between planktonic and biofilm lifestyles [37].
Stress Response Pathways: Program Cas13 to interfere with bacterial stress response systems, including:
- Stringent response pathways regulated by (p)ppGpp alarmones
- Oxidative stress defense mechanisms
- SOS response and DNA repair systems
Diagram 2: Strategic Targeting Approach for Cas13-Antibiotic Synergy. This workflow illustrates the rational selection of Cas13 targets to potentiate conventional antibiotic activity through multiple complementary mechanisms.
Technical Considerations and Optimization Strategies
Successful implementation of Cas13-antibiotic combination therapies requires careful attention to several technical aspects:
Delivery Optimization: Efficient delivery remains the primary challenge for CRISPR-based antimicrobials. Strategies to enhance delivery include:
- Conjugative plasmids that enable transfer between bacterial cells
- Phage capsids engineered for specific bacterial receptor targeting
- Cationic nanoparticles optimized for biofilm penetration
- Chemical modifications to guide RNAs to enhance stability in biofilm environments
Dosing and Timing: The temporal relationship between Cas13 and antibiotic administration significantly impacts efficacy. Conduct time-kill studies to establish optimal sequencing—generally, pre-treatment with Cas13 for 2-4 hours before antibiotic administration maximizes resensitization effects.
Resistance Mitigation: While Cas13-antibiotic combinations reduce resistance development compared to monotherapies, potential escape mechanisms include:
- Mutations in Cas13 target sequences
- Upregulation of efflux pumps
- Enhanced DNA repair systems Counter these mechanisms by using multiplexed guide RNAs targeting multiple essential genes simultaneously and incorporating efflux pump inhibitors into combination regimens.
The strategic combination of CRISPR-Cas13 systems with conventional antibiotics represents a paradigm shift in our approach to combating biofilm-associated antimicrobial resistance. By enabling precise targeting of resistance mechanisms and biofilm maintenance pathways, Cas13 resensitizes resistant bacteria to traditional antibiotics, creating powerful synergistic effects that overcome the limitations of monotherapies. The protocols and frameworks presented herein provide researchers with validated methodologies for developing and optimizing these innovative combination strategies.
Future advancements in this field will likely focus on enhancing delivery efficiency through novel nanocarriers, expanding the repertoire of targetable pathways through improved bioinformatic selection, and developing personalized approaches based on rapid diagnostics of resistance patterns. As these technologies mature, CRISPR-Cas13 and antibiotic combinations hold exceptional promise for addressing some of the most challenging infections in clinical practice, potentially extending the useful lifespan of our existing antibiotic arsenal while overcoming the defensive advantages conferred by biofilm growth.
Benchmarking Success: Validating Cas13 Performance and Comparing RNA-Targeting Platforms
The advent of CRISPR-Cas13 technology has revolutionized RNA targeting, offering a powerful approach for functional genomics research. Within biofilm metabolic pathways research, precise knockdown of key transcripts enables researchers to decipher complex regulatory networks and identify potential therapeutic targets. This Application Note provides detailed protocols and methodologies for quantitatively assessing both transcript reduction and subsequent phenotypic consequences in biofilm studies utilizing CRISPR-Cas13 systems. The procedures outlined specifically address the unique challenges of working with biofilm models, including delivery efficiency in extracellular polymeric substance (EPS)-rich matrices and differentiation between transient and stable knockdown effects.
Quantitative Assessment of Knockdown Efficiency
Transcript-Level Quantification Methods
Reverse Transcription Quantitative PCR (RT-qPCR) remains the gold standard for directly quantifying transcript knockdown efficiency following CRISPR-Cas13 treatment. This method provides sensitive, quantitative data on target RNA reduction when properly optimized with appropriate controls.
Table 1: Key Reagents for RT-qPCR-based Transcript Quantification
| Research Reagent | Function/Application | Example/Notes |
|---|---|---|
| CRISPR-RfxCas13d Plasmid | RNA-targeting effector | Addgene-sourced backbone [59] [60] |
| Target-Specific crRNAs | Guides Cas13 to RNA target | Designed to span back-splicing junctions or exon-exon junctions [59] [61] |
| RNA Extraction Kit | Isolate intact RNA from biofilms | Ensure compatibility with EPS-rich matrices [8] |
| Reverse Transcriptase | cDNA synthesis | Use consistent amounts across samples [59] |
| SYBR Green Master Mix | qPCR detection | Enables quantification of target vs. reference genes [59] |
| Junction-Specific Primers | Amplify unique splice variants | Critical for isoform-specific quantification [61] |
The experimental workflow for transcript quantification must account for several crucial factors to ensure accurate results. Junction-targeting approaches are particularly valuable in biofilm research, where alternative splicing may generate multiple transcript isoforms with distinct functions in metabolic pathway regulation. Guide RNAs targeting exon-exon junctions (EEJs) provide isoform-specific knockdown capability, enabling precise functional studies of individual splice variants within complex biofilm regulatory networks [61].
Diagram 1: Transcript quantification workflow for assessing CRISPR-Cas13 knockdown efficiency.
Optimization of crRNA design significantly impacts knockdown efficiency. In comparative studies targeting circRNAs, CRISPR-Cas13d with 24-nucleotide crRNAs demonstrated approximately 50% knockdown efficiency initially, with optimization of transfection conditions increasing efficiency to 70% - significantly higher than the 40% achieved with siRNA methods [59] [60]. When designing crRNAs for biofilm metabolic pathway targets, consider:
- Length optimization: Systematically test crRNAs of different nucleotide lengths (e.g., 24, 26, 30, 34 nt)
- Junction spanning: Design crRNAs that span back-splicing junctions for circular RNAs or exon-exon junctions for linear transcript isoforms
- Accessibility validation: Perform essentiality screens to confirm target accessibility
Phenotypic Assessment Methodologies
Following transcript knockdown, comprehensive phenotypic assessment is essential to establish functional consequences in biofilm metabolic pathways. Multi-dimensional approaches provide the most complete understanding of knockdown effects.
Table 2: Phenotypic Assessment Methods for Biofilm Metabolic Pathways
| Phenotypic Category | Assessment Method | Key Metrics | Application in Biofilm Research |
|---|---|---|---|
| Matrix Production | Western Blot | Collagen I, α-SMA expression | Quantify extracellular matrix components [59] [60] |
| Biofilm Architecture | Confocal Microscopy | Biomass, thickness, porosity | Visualize structural changes post-knockdown [8] |
| Metabolic Activity | Resazurin Assay | Metabolic flux, viability | Assess pathway-specific functional consequences [8] |
| Antimicrobial Susceptibility | MIC/MBC assays | Biofilm eradication efficiency | Evaluate potential synergy with antimicrobials [8] |
| Virulence Factor Production | ELISA, Enzymatic Assays | Toxin, enzyme secretion | Monitor pathogenicity alterations [8] |
Diagram 2: Signaling pathway from transcript knockdown to phenotypic consequences in biofilms.
In biofilm metabolic pathway research, multiplexed phenotypic assessment provides comprehensive functional data. For example, knockdown of circAdpgk-0001 using Cas13d reduced expression of activation-related factors collagen I and α-SMA by approximately 40%, outperforming siRNA-mediated knockdown [59] [60]. This demonstrates the importance of correlating transcript reduction with downstream functional effects in biofilm systems.
Experimental Protocols
Protocol 1: CRISPR-Cas13d Mediated Transcript Knockdown in Biofilm Models
Materials:
- CRISPR-RfxCas13d plasmid (Addgene)
- BsmBI-v2 enzyme (NEB)
- T4 DNA Ligase (Takara)
- Lipofectamine3000 or jetPRIME Transfection Reagents
- Target-specific crRNAs
Procedure:
- crRNA Design: Design crRNAs targeting specific exon-exon junctions of your target transcript. For a 24-nt crRNA, take 11 nucleotides on each side of the junction (excluding the junction itself) [59].
- Plasmid Construction:
- Digest 2 µg CRISPR-RfxCas13d plasmid with 2 µL BsmBI-v2 enzyme in 1× NEBuffer 3.1 at 55°C for 1 hour [59] [60].
- Purify linearized vector using PCR purification kit.
- Anneal crRNA oligonucleotides by mixing forward and reverse strands in NEBuffer 2.1 and gradually cooling from 95°C to 22°C at 1°C per minute [59].
- Ligate annealed product into linearized vector using T4 DNA Ligase at 16°C overnight [59].
- Transform into E. coli DH5α, then culture and purify plasmid using endotoxin-free maxiprep kit.
- Biofilm Transfection:
- Culture biofilm-forming cells to 70-90% confluency.
- Transfect using Lipofectamine3000 or jetPRIME according to manufacturer protocols.
- For stimulation experiments, add relevant inducters (e.g., TGF-β at 10 ng/mL) concurrently or post-transfection based on optimization [59].
- Harvest cells at 24-48 hours post-transfection for analysis.
Protocol 2: Quantitative Assessment of Knockdown Efficiency
Materials:
- RNA extraction kit (compatible with biofilm material)
- cDNA synthesis kit
- SYBR Green qPCR master mix
- Junction-specific primers
Procedure:
- RNA Extraction:
- Harvest biofilm cells using appropriate dissociation methods.
- Extract RNA using validated methods, ensuring minimal degradation.
- Quantify RNA concentration and assess quality (A260/A280 ratio >1.8).
- cDNA Synthesis:
- Use consistent amounts of RNA (500-1000 ng) across all samples.
- Perform reverse transcription using random hexamers or oligo-dT primers.
- qPCR Analysis:
- Design primers spanning the same junction targeted by crRNAs.
- Include housekeeping genes (GAPDH, β-actin) for normalization.
- Set up reactions in triplicate using SYBR Green chemistry.
- Use the following cycling conditions: 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min.
- Data Analysis:
- Calculate ΔΔCt values comparing treated vs. control samples.
- Determine knockdown efficiency using the formula: % Knockdown = (1 - 2^(-ΔΔCt)) × 100
Protocol 3: Functional Phenotypic Assessment in Biofilms
Materials:
- Lysis buffer for protein extraction
- Primary antibodies for target proteins
- Resazurin solution
- Microtiter plates for biofilm cultivation
Procedure:
- Western Blot Analysis:
- Extract proteins from transfected biofilms using RIPA buffer with protease inhibitors.
- Separate proteins by SDS-PAGE and transfer to PVDF membranes.
- Probe with primary antibodies against target proteins (e.g., collagen I, α-SMA).
- Use appropriate secondary antibodies and develop using ECL.
- Quantify band intensity normalized to loading controls.
- Metabolic Activity Assessment:
- Culture biofilms in 96-well plates.
- Add resazurin solution (10% v/v) and incubate 2-4 hours.
- Measure fluorescence at 560ex/590em.
- Normalize to biomass or cell number controls.
- Biofilm Architecture Analysis:
- Stain biofilms with SYTO9/propidium iodide for viability assessment.
- Image using confocal microscopy at multiple random fields.
- Analyze biomass, thickness, and viability using image analysis software.
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for CRISPR-Cas13 Biofilm Studies
| Tool Category | Specific Product/Type | Function | Considerations for Biofilm Research |
|---|---|---|---|
| CRISPR-Cas13 System | RfxCas13d plasmid | RNA-targeting effector | Higher efficiency than siRNA for certain targets [59] [60] |
| Delivery Vehicles | Lipofectamine3000, jetPRIME, nanocarriers | Cas13 plasmid/crRNA delivery | Must penetrate EPS matrix; efficiency varies [8] |
| crRNA Design Tools | TIGER model, Isoviz R package | Predict gRNA efficacy | ~30% of randomly selected EEJ gRNAs are effective [61] |
| Biofilm Growth Substrata | Calgary biofilm device, flow cells | Reproducible biofilm cultivation | Enable high-throughput screening [8] |
| Detection Assays | SHERLOCK (Cas13-based) | Pathogen/target detection | Attomolar sensitivity in complex samples [8] |
Troubleshooting and Optimization
Common challenges in quantifying CRISPR-Cas13 knockdown in biofilm systems include delivery efficiency barriers due to the extracellular polymeric substance matrix, transcriptional redundancy in metabolic pathways, and temporal stability of knockdown effects. To address these:
- Optimize delivery: Test multiple transfection reagents and consider nanocarrier systems designed for biofilm penetration [8]
- Validate specificity: Include multiple crRNAs targeting the same transcript and utilize non-targeting controls
- Monitor kinetics: Assess knockdown at multiple timepoints (24, 48, 72 hours) to establish effect duration
- Employ multiplexing: Target multiple nodes in parallel pathways to overcome redundancy
Recent advances in Cas13d-based systems have demonstrated superior performance compared to traditional siRNA, with optimized protocols achieving 70% knockdown efficiency versus 40% with siRNA methods [59] [60]. Furthermore, the application of deep learning models like TIGER for gRNA efficacy prediction has improved the success rate of target engagement in complex transcriptomes [61].
For researchers investigating biofilm metabolic pathways, these quantification protocols provide a standardized framework for validating CRISPR-Cas13 mediated knockdown and establishing robust correlations between transcript reduction and functional phenotypic outcomes, accelerating the identification of novel therapeutic targets for biofilm-associated infections.
This application note provides a systematic evaluation of four CRISPR-Cas13 orthologs—Cas13a, Cas13d, Cas13x, and Cas13y—for targeted RNA knockdown in the context of biofilm metabolic pathways research. We present quantitative performance data, detailed experimental protocols for assessing knockdown efficiency, and visualization of the experimental workflow. This resource is designed to assist researchers in selecting appropriate Cas13 variants and implementing robust methods for interrogating RNA-level regulation in biofilm-forming pathogens.
CRISPR-Cas13 systems have emerged as powerful tools for programmable RNA targeting, enabling transcript knockdown without permanent genomic alteration. Unlike DNA-targeting Cas nucleases, Cas13 effectors possess two distinct ribonuclease activities: one for processing its own precursor CRISPR RNA (pre-crRNA) and another for cleaving target RNA sequences guided by mature crRNAs [6] [13]. This functionality is particularly valuable for studying biofilm formation, a complex process involving dynamic gene expression changes where transient transcript modulation is often preferable to permanent genetic disruption.
For researchers investigating metabolic pathways in biofilms, Cas13 technology offers several advantages: (1) ability to target both chromosomal and plasmid-borne transcripts, including antibiotic resistance genes; (2) compatibility with multiplexed targeting through customized crRNA arrays; and (3) potential for reversible gene suppression to study essential metabolic genes [8] [62]. This application note focuses on the comparative analysis of four Cas13 orthologs to guide selection for biofilm pathway research.
Ortholog Performance Comparison
A recent systematic evaluation of Cas13 orthologs provides critical quantitative data for variant selection. The study assessed knockdown efficiency against endogenous transcripts and RNA viruses in plant systems, offering comparable performance metrics relevant to microbial biofilm research [63].
Table 1: Comparative Characteristics of Cas13 Orthologs
| Ortholog | Subtype | Size (amino acids) | Knockdown Efficiency | PFS Preference | Key Features |
|---|---|---|---|---|---|
| LwaCas13a | VI-A | ~1,200 | ~40-60% | 3' U, A, or C | First characterized, robust activity |
| RfxCas13d | VI-D | ~930 | 70-80% | None | High efficiency, minimal constraints |
| Cas13x.1 | VI-X | ~775 | 65-75% | None | Ultra-compact, efficient packaging |
| Cas13y.1 | VI-Y | ~790 | 58-68% | None | Compact size, good activity |
Table 2: Quantitative Knockdown Efficiency of Cas13 Orthologs
| Ortholog | Target Transcript 1 Efficiency | Target Transcript 2 Efficiency | Multiplexing Efficiency | Viral RNA Reduction |
|---|---|---|---|---|
| LwaCas13a | 42.5% ± 3.2% | 51.7% ± 4.1% | 38.2% ± 5.1% | ~60% |
| RfxCas13d | 78.3% ± 2.7% | 75.6% ± 3.3% | 72.4% ± 4.2% | ~85% |
| Cas13x.1 | 73.8% ± 3.1% | 69.5% ± 2.9% | 68.9% ± 3.8% | ~80% |
| Cas13y.1 | 65.2% ± 3.5% | 61.8% ± 3.7% | 58.3% ± 4.1% | ~75% |
The data reveal that RfxCas13d, Cas13x.1, and Cas13y.1 exhibit enhanced stability with editing efficiencies ranging from 58% to 80%, with RfxCas13d demonstrating the highest consistent knockdown across diverse targets [63]. Notably, Cas13x.1 and Cas13y.1 achieve these efficiencies with significantly smaller protein sizes, facilitating delivery via size-constrained vectors like adeno-associated viruses (AAVs) - an important consideration for potential therapeutic applications against biofilm-related infections [16].
Experimental Protocol for Cas13-Mediated Transcript Knockdown in Biofilm Models
Reagent Preparation
Cas13 Expression Constructs:
- Clone codon-optimized Cas13 orthologs (LwaCas13a, RfxCas13d, Cas13x.1, Cas13y.1) into appropriate expression vectors under inducible promoters (e.g., pTet, pBAD).
- For bacterial biofilm studies, ensure compatibility with origin of replication and selection markers relevant to target species (e.g., p15A ori for Gram-negative bacteria; rolling circle replication for Gram-positive species).
crRNA Design and Array Cloning:
- Design spacer sequences (20-28 nt) complementary to target mRNA regions within biofilm-related genes (e.g., quorum sensing, EPS production, metabolic pathway genes).
- Avoid target regions with strong secondary structure that may impair Cas13 binding [2].
- For multiplexed targeting, clone up to 5 crRNAs as a single array with direct repeats separating individual spacers, leveraging Cas13's native pre-crRNA processing capability [13].
- Clone crRNA arrays into expression vectors with strong, constitutive promoters.
Bacterial Transformation and Biofilm Cultivation
Day 1: Transformation
- Prepare electrocompetent cells of target biofilm-forming strain (e.g., Pseudomonas aeruginosa, Staphylococcus aureus).
- Co-transform with Cas13 expression plasmid and crRNA array plasmid (or use single plasmid system).
- Plate on selective media and incubate overnight at appropriate temperature.
Day 2: Biofilm Induction
- Inoculate 3-5 transformed colonies into 5 mL selective media with inducer (if using inducible Cas13).
- Incubate with shaking to mid-exponential phase (OD600 ≈ 0.5-0.6).
- Dilute culture to OD600 ≈ 0.1 in fresh media with appropriate antibiotics and inducer.
- Aliquot 1-2 mL into biofilm cultivation vessels (e.g., 24-well plates with or without inserts).
- Incubate statically for biofilm formation (24-48 hours, species-dependent).
Knockdown Efficiency Assessment
Day 4: RNA Extraction and Analysis
- Carefully aspirate planktonic culture from established biofilms.
- Gently wash biofilm twice with PBS to remove loosely attached cells.
- Add RNA stabilization reagent directly to biofilm and incubate 5 minutes.
- Disrupt biofilm by pipetting or gentle scraping and transfer suspension to RNAse-free tubes.
- Extract total RNA using commercial kit with DNase treatment step.
- Quantify RNA concentration and purity (A260/280 ≈ 2.0).
- Convert 1 μg total RNA to cDNA using reverse transcription kit.
- Perform qPCR with target gene primers and reference gene primers (e.g., rpoB, gyrA).
- Calculate knockdown efficiency using ΔΔCt method relative to non-targeting crRNA control.
Visualizing the Experimental Workflow
Cas13 Biofilm Analysis Workflow
This workflow outlines the key stages for evaluating Cas13-mediated transcript knockdown in biofilm models, from initial construct design to quantitative assessment of targeting efficiency.
Cas13 Molecular Mechanism in Biofilm Pathway Targeting
Cas13 RNA Targeting Mechanism
This diagram illustrates the molecular mechanism of Cas13 orthologs in targeting biofilm pathway transcripts. Upon forming a complex with guide crRNA, Cas13 recognizes and binds complementary target mRNA through base pairing, leading to HEPN domain-mediated cleavage and transcript degradation. The resulting knockdown of key biofilm genes (e.g., quorum sensing regulators, metabolic enzymes) leads to measurable changes in biofilm formation. Note the potential for collateral RNA cleavage, an important consideration for experimental design [13] [2].
Research Reagent Solutions
Table 3: Essential Research Reagents for Cas13 Biofilm Studies
| Reagent Category | Specific Examples | Function & Application Notes |
|---|---|---|
| Cas13 Expression Plasmids | pC013-LwaCas13a, pRfxCas13d, pC013X.1, pC013Y.1 | Codon-optimized variants with inducible promoters for controlled expression in target organisms. |
| crRNA Cloning Systems | pCRRNA-Array (tRNA-based), Direct Golden Gate assembly | Enable multiplexed guide expression; tRNA-flanked arrays enhance processing efficiency [63]. |
| Biofilm Cultivation Materials | 96-well polystyrene plates, Calgary biofilm device, flow cell systems | Provide standardized surfaces for reproducible biofilm growth and analysis. |
| RNA Analysis Tools | Biofilm RNA extraction kits, DNase I treatment, reverse transcription kits | Optimized for difficult-to-lyse biofilm samples; essential for accurate transcript quantification. |
| Delivery Systems | Electroporation apparatus, liposomal nanoparticles, conjugated polymers | Facilitate Cas13 component delivery; nanoparticles enhance biofilm penetration [12]. |
Troubleshooting and Optimization Guidelines
Low Knockdown Efficiency:
- Verify crRNA target accessibility by testing multiple spacer sequences along the transcript.
- Optimize Cas13 expression levels through inducer concentration titration.
- Ensure crRNA processing by checking array design with appropriate direct repeats.
- For RfxCas13d, utilize the minimal 5'-DR-ggRNA-3' architecture for optimal performance [63].
Cellular Toxicity:
- Reduce Cas13 expression levels using weaker promoters or lower inducer concentrations.
- Consider engineered Cas13 variants with reduced collateral activity [2].
- Limit exposure time through transient induction protocols.
Biofilm-Specific Challenges:
- Implement enhanced delivery strategies for mature biofilms, such as nanoparticle conjugation [12].
- Include appropriate controls for biofilm heterogeneity (multiple biological replicates).
- Combine with biofilm disruption methods (e.g., mild sonication) for more uniform sampling.
Application in Biofilm Metabolic Pathway Research
The Cas13 orthologs characterized here enable precise dissection of metabolic pathways essential for biofilm formation and maintenance. Specific applications include:
- Functional Analysis of Essential Genes: Knockdown of essential metabolic genes without permanent lethality allows study of their roles in biofilm-specific metabolic adaptations [62].
- Quorum Sensing Circuits: Targeted degradation of autoinducer synthase transcripts or regulator mRNAs to interrogate signaling dynamics.
- Metabolic Specialization: Investigation of metabolic division of labor in heterogeneous biofilm communities through zone-specific transcript knockdown.
- Antibiotic Persistence: Role of metabolic dormancy pathways in biofilm-associated antibiotic tolerance through transient transcript suppression.
The compact size of Cas13x.1 and Cas13y.1 orthologs is particularly advantageous for delivery challenges in dense biofilm matrices, enabling more efficient penetration and distribution [16].
This systematic evaluation demonstrates that Cas13d, Cas13x, and Cas13y orthologs offer superior knockdown efficiency compared to Cas13a, with RfxCas13d showing the highest consistent performance against diverse transcript targets. The provided protocols and troubleshooting guidelines enable robust implementation of Cas13 technology for dissecting metabolic pathways in biofilm systems. As CRISPR-Cas13 tools continue to evolve, with ongoing development of engineered variants with reduced collateral activity and enhanced specificity, their application in biofilm research will provide increasingly precise insights into the RNA-level regulation of these complex microbial communities.
In the field of RNA-targeting for biofilm research, two primary technologies enable gene silencing: RNA interference (RNAi) and CRISPR-Cas13. Understanding their distinct specificity and off-target profiles is crucial for designing reliable experiments investigating biofilm metabolic pathways. While RNAi has been the traditional knockdown method, the more recently developed CRISPR-Cas13 system offers a fundamentally different mechanism with potential advantages for precision. This Application Note provides a direct, evidence-based comparison of these platforms, focusing on their application in studying biofilm-associated genes, with structured data and actionable protocols to guide researchers and drug development professionals.
The core distinction between these technologies lies in their targeting mechanism: RNAi functions as a catalytic knockdown system, while Cas13 can operate as a multiple-turnover cleaver or, in its inactive form (dCas13), as a steric blocker.
RNAi relies on the cell's endogenous RNA-induced silencing complex (RISC). Introduced small interfering RNAs (siRNAs) or short hairpin RNAs (shRNAs) are loaded into RISC, guiding it to complementary mRNA targets primarily through base-pairing in the "seed region" (nucleotides 2-8 of the guide). This leads to Argonaute-mediated cleavage of the mRNA or translational repression if complementarity is imperfect [64].
CRISPR-Cas13 is a prokaryotic-derived system where a single Cas13 nuclease complexed with a guide RNA (gRNA) directly binds and cleaves single-stranded RNA targets. Cas13's activation requires near-perfect complementarity between the gRNA spacer and the target RNA. Upon activation, Cas13 exhibits collateral RNase activity, non-specifically cleaving nearby bystander RNAs, a significant concern for off-target effects in some contexts [14] [65]. The catalytically dead Cas13 (dCas13) variant binds target RNA without cleavage, functioning as a steric block to processes like translation initiation [65].
The table below summarizes the fundamental differences.
Table 1: Fundamental Mechanisms of RNAi and CRISPR-Cas13
| Feature | RNAi (siRNA/shRNA) | CRISPR-Cas13 (Active) | CRISPR-dCas13 (Inactive) |
|---|---|---|---|
| Mechanism of Action | Endogenous RISC-mediated cleavage or translational repression | RNA-guided, HEPN domain-mediated cleavage of target RNA | Steric blockade of ribosomal scanning or protein binding |
| Guide Component | siRNA (21-23 nt) or shRNA | gRNA (~28-30 nt spacer for Cas13a/b, ~23 nt for Cas13d) | Same as active Cas13 |
| Complementarity Requirement | Imperfect pairing tolerated, especially outside seed region | Near-perfect complementarity required for activation | Near-perfect complementarity required for tight binding |
| Catalytic Nature | Single-turnover (guide is degraded) | Multiple-turnover (guide is not degraded) | Not applicable (catalytically inactive) |
| Primary Off-Target Sources | Seed-based off-targeting; interferon response | Collateral RNase activity; gRNA-dependent off-target binding | gRNA-dependent off-target binding (no collateral activity) |
Figure 1: Comparative Signaling Pathways for RNAi, Cas13, and dCas13. The Cas13 pathway highlights the risk of collateral RNA cleavage, a major source of off-target effects, which is absent in the dCas13 and RNAi pathways.
Quantitative Comparison of Specificity and Off-Target Effects
A critical consideration for biofilm metabolic pathway research is the minimization of off-target effects that can confound phenotypic results. Comparative studies and high-throughput screens consistently show CRISPR-Cas13 has superior specificity compared to RNAi.
RNAi is notoriously prone to sequence-dependent off-targets. The RISC complex requires only limited complementarity, particularly in the 2-8 nucleotide "seed" region of the guide, to suppress transcripts. One consequence is that a single siRNA can inadvertently target hundreds of genes with partial seed matches [64]. Furthermore, sequence-independent off-targets can occur via activation of the innate immune response and interferon pathways.
CRISPR-Cas13 requires extensive complementarity for activation, theoretically granting higher specificity. However, its primary confounder is collateral RNase activity. Upon target recognition, the HEPN domains of activated Cas13 non-specifically cleave nearby RNAs, potentially causing widespread transcriptome damage and cytotoxicity [14] [65]. A key advancement is the use of catalytically dead Cas13 (dCas13). dCas13 binds RNA without cleavage, repressing translation through steric hindrance. Genome-wide ribosome profiling has confirmed that dCas13-mediated silencing (CRISPRδ) achieves ultra-high gene silencing specificity without measurable collateral effects [65].
Deep learning models like DeepCas13 have been developed to predict gRNA on-target efficiency, helping to optimize guide design and minimize off-target viability effects. Models show that features determining off-target viability effects are closely related to on-target efficiency [66].
Table 2: Direct Comparison of Specificity and Off-Target Profiles
| Parameter | RNAi | CRISPR-Cas13 (Active) | CRISPR-dCas13 (Inactive) |
|---|---|---|---|
| Guide-Dependent Off-Target Rate | High (up to hundreds of genes per siRNA) [64] | Moderate (dependent on gRNA design) [66] | Low (high specificity binding) [65] |
| Primary Off-Target Mechanism | Seed-based hybridization; imperfect complementarity | Collateral RNase activity upon activation; gRNA mismatch tolerance | gRNA mismatch tolerance (no cleavage) |
| Immunogenicity / Cytotoxicity | Can trigger interferon response [64] | Cytotoxicity from collateral RNA cleavage [14] [65] | Generally low |
| Key Mitigation Strategies | Chemical modifications; pooled controls; seed region analysis [64] | Optimized gRNA design (e.g., DeepCas13); using dCas13 variants [66] [65] | Optimal gRNA placement (start codon) [65] |
| Typical Knockdown Efficiency | Variable; often 70-90% (highly sequence-dependent) | High (>90% for efficient guides) [61] | Moderate to High (50-90%, position-dependent) [65] |
Experimental Protocols for Biofilm Metabolic Pathway Studies
Protocol: dCas13-Mediated Translational Repression for High-Specificity Knockdown
This protocol uses dCas13 for precise gene silencing without collateral activity, ideal for probing essential biofilm metabolic genes where off-target effects could be lethal or produce misleading phenotypes [65].
Research Reagent Solutions Table 3: Essential Reagents for dCas13 Translational Repression
| Item | Function / Description | Example |
|---|---|---|
| dCas13 Effector Plasmid | Expresses catalytically inactive dCas13 (e.g., dPspCas13b-NES). | Addgene # # (e.g., for dPspCas13b) |
| gRNA Expression Vector | Cloning vector for target-specific gRNA sequence. | Custom synthesized or cloned into backbone (e.g., U6 promoter). |
| Delivery Vehicle | For transfection in bacterial biofilms. | Liposomes, polymeric nanoparticles, or conjugative systems. |
| Negative Control gRNAs | Non-targeting gRNAs to control for dCas13 binding effects. | Designed to have no perfect match in the host transcriptome. |
| Validation Primers | For qRT-PCR to confirm mRNA level stability. | Target the mRNA region outside the gRNA binding site. |
Step-by-Step Workflow:
gRNA Design and Cloning:
- Design: Design gRNAs with a 23-30 nt spacer sequence targeting the start codon region of the target mRNA. Tiling several gRNAs across this region is recommended for optimal efficacy [65].
- Specificity Check: Use tools like DeepCas13 [66] to predict on-target efficiency and BLAST against the host genome to minimize off-target binding.
- Cloning: Synthesize and clone the gRNA sequence into the expression vector under a U6 or T7 promoter.
Complex Formation and Delivery:
- For in vitro assays, the dCas13 protein can be pre-complexed with in vitro transcribed gRNA to form a ribonucleoprotein (RNP). This reduces off-targets and increases editing efficiency [64].
- For bacterial biofilm delivery, use engineered nanoparticles (e.g., gold or lipid nanoparticles) or conjugative systems to deliver the dCas13 expression plasmid and gRNA expression vector [8] [11]. The choice of delivery vector is critical for penetrating the biofilm matrix.
Transfection and Induction:
- Apply the delivery vehicle to pre-formed biofilms. For planktonic controls, standard transfection methods can be used.
- If using an inducible promoter for dCas13/gRNA expression, induce with the appropriate agent (e.g., anhydrous tetracycline).
Validation and Phenotyping:
- Specificity Validation (48-72 hours post-delivery):
- Extract total RNA.
- Perform qRT-PCR for the target gene and a panel of potential off-target genes (e.g., genes with partial gRNA homology). The key differentiator from active Cas13 is that target mRNA levels should remain stable with dCas13, while protein output is reduced [65].
- For genome-wide specificity assessment, ribosome profiling (Ribo-seq) can be used to confirm the absence of widespread translational disruption [65].
- Phenotypic Assay: Assess the impact on biofilm metabolism (e.g., using ATP assays, resazurin metabolism assays, or specific nutrient consumption/production assays) and biofilm integrity (e.g., crystal violet staining, confocal microscopy).
- Specificity Validation (48-72 hours post-delivery):
Figure 2: Experimental workflow for dCas13-mediated gene silencing in biofilm studies, highlighting key steps from gRNA design to phenotypic validation.
Protocol: Isoform-Specific Knockdown Using Cas13d for Biofilm Pathway Analysis
Biofilm bacteria often express distinct gene isoforms in response to environmental stress. This protocol details the use of Cas13d (CasRx) with junction-spanning gRNAs for isoform-specific knockdown, enabling precise functional dissection of metabolic pathway components [61].
Step-by-Step Workflow:
Identify Target Isoform and Unique Junction:
- Use RNA-seq data from biofilm conditions to identify differentially expressed isoforms of the target metabolic gene.
- Identify a fully unique exon-exon junction (EEJ) present only in the target isoform using annotation databases (e.g., GENCODE) and custom scripts. The Isoviz R package can assist in experimental design [61].
Design and Select EEJ-Targeting gRNAs:
- Design gRNAs (typically 23 nt for Cas13d) where the spacer sequence spans the unique EEJ. Tiling several gRNAs across the junction (e.g., from -15 bp to +15 bp relative to the junction) is recommended to ensure high activity [61].
- Use predictive models like TIGER [61] or DeepCas13 [66] to score and select gRNAs with high predicted efficiency. For Cas13d, approximately 30% of randomly selected EEJ gRNAs are active, a rate that can be significantly enriched with predictive modeling.
Delivery and Screening in Biofilm Models:
- Clone selected gRNAs into a Cas13d (RfxCas13d) expression system.
- Deliver the constructs to your bacterial biofilm model using appropriate vectors (e.g., phagemids, conjugative plasmids) [8].
- Screen for successful knockdown using isoform-specific RT-qPCR (using primers spanning the unique junction) or long-read RNA-sequencing.
Functional Validation:
- Assess the impact of isoform-specific knockdown on the biofilm's metabolic phenotype, such as resistance to specific antibiotics, nutrient utilization efficiency, or production of key metabolites.
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Research Reagent Solutions for RNA-Targeting Experiments
| Category | Item | Function / Application |
|---|---|---|
| CRISPR-Cas13 Systems | dPspCas13b-NES/dPspCas13b-NLS [65] | Effector proteins for translational repression; NES for cytoplasmic, NLS for nuclear roles. |
| RfxCas13d (CasRx) [66] [61] | Compact Cas13 variant ideal for screening and isoform-specific targeting. | |
| gRNA Design & Analysis | DeepCas13 Web Server [66] | Deep learning model to predict Cas13d gRNA on-target activity. |
| TIGER Model & Isoviz R Package [61] | Predicts gRNA efficiency and aids in designing isoform-specific EEJ-targeting experiments. | |
| Delivery Platforms | Lipid Nanoparticles (LNPs) [11] | Efficient delivery of CRISPR components in vitro and in vivo. |
| Gold Nanoparticles (AuNPs) [11] | Co-delivery of Cas9/gRNA for biofilm disruption; can be adapted for Cas13. | |
| Conjugative Plasmids / Phagemids [8] | For efficient delivery of CRISPR constructs into bacterial biofilms. | |
| Validation Tools | Ribosome Profiling (Ribo-seq) [65] | Gold-standard for genome-wide assessment of translational repression specificity. |
| Long-Read RNA-Sequencing (LRS) [61] | For unambiguous validation of isoform-specific knockdown. |
Validation in Complex Multi-Species Biofilm Models
The study of biofilm-mediated antibiotic resistance and pathogenesis requires models that accurately recapitulate the complexity of natural microbial communities. Complex multi-species biofilm models are essential tools for validating the efficacy and specificity of novel RNA-targeting interventions, such as those utilizing the CRISPR-Cas13 system. This protocol outlines the methodology for establishing, treating, and analyzing multi-species biofilms to validate CRISPR-Cas13a-based targeting of metabolic pathways, providing researchers with a standardized framework for assessing therapeutic efficacy in conditions that mimic clinical and environmental scenarios. The application of CRISPR-Cas13 in this context represents a paradigm shift in biofilm research, enabling precise RNA-level modulation of key metabolic and virulence pathways without permanent genomic alteration [8] [67].
Theoretical Framework: CRISPR-Cas13a for RNA Targeting in Biofilms
The CRISPR-Cas13a system (Type VI-A) represents a powerful RNA-targeting platform derived from bacterial adaptive immune systems. Unlike DNA-targeting Cas nucleases, Cas13a possesses RNA-guided RNase activity, making it ideally suited for transcript-specific knockdown in biofilm communities. Upon recognition and cleavage of its target RNA through complementary crRNA, the Cas13a protein exhibits collateral RNase activity, enabling amplified signal detection in diagnostic applications and potent degradation of transcript targets [67].
In biofilm research, this system enables precise dissection of metabolic pathway contributions to extracellular polymeric substance (EPS) production, quorum sensing, and stress response without selecting for resistant mutants through permanent genetic changes. The programmable nature of Cas13a allows researchers to simultaneously target multiple transcripts within critical biofilm pathways, offering unprecedented capability for analyzing functional redundancy in multi-species communities [8] [62].
Table 1: CRISPR-Cas13 System Components and Functions in Biofilm Research
| Component | Function in Biofilm Studies | Considerations |
|---|---|---|
| Cas13a Nuclease | RNA-guided RNase for transcript knockdown | HEPN domain catalytic sites require R/H active motifs |
| crRNA Guide | 20-27nt spacer for target specificity; 5' guanine recommended | Avoid functional RNA motifs; design 3-5 guides per target |
| Delivery Vector | Plasmid, phage, or nanoparticle-based delivery | Must penetrate EPS barrier; inducible promoters preferred |
| HEP Reporter | Fluorescent RNA molecule for collateral activity detection | Validation of targeting efficiency in real-time |
Experimental Models and Validation Workflow
Multi-Species Biofilm Model Establishment
The selection of appropriate microbial species is critical for developing clinically relevant multi-species biofilm models. A recommended model system for gastrointestinal and medical device-related biofilm studies includes:
- Primary Species: Escherichia coli (gram-negative), Staphylococcus aureus (gram-positive), and Candida albicans (fungal component)
- Culture Conditions: YESCA media at 25-28°C for 3-4 days under static conditions to promote pellicle biofilm formation with characteristic curli, cellulose, and LPS matrix components [68]
- Surface Substrates: Cholesterol-coated surfaces (gallstone-mimetic), medical-grade silicone, or stainless steel coupons to simulate clinical and industrial environments
Biofilm maturity should be validated through scanning electron microscopy (SEM) to confirm the presence of characteristic heterogeneous architecture with microcolonies, water channels, and dense extracellular matrix [11]. The model should demonstrate at least 1000-fold increased antimicrobial tolerance compared to planktonic cultures before experimental interventions [11].
CRISPR-Cas13a Delivery Systems for Biofilm Penetration
Efficient delivery of CRISPR-Cas13 components through the protective biofilm matrix represents a significant technical challenge. The following delivery systems have demonstrated efficacy in complex biofilm models:
- Lipid-Based Nanoparticles: Formulated with cationic lipids to complex with CRISPR-Cas13a RNPs; shown to reduce P. aeruginosa biofilm biomass by >90% in vitro [11]
- Engineered Phage Systems: Bacteriophage modified to deliver Cas13a expression constructs; provides species-specific targeting within multi-species communities
- Conjugative Plasmids: Enable interbacterial transfer of Cas13a machinery; particularly effective for gram-negative species
- Gold Nanoparticle Carriers: Enhance editing efficiency up to 3.5-fold compared to non-carrier systems; enable co-delivery with antibiotics [11]
Table 2: Quantitative Efficacy of CRISPR-Cas13 Against Biofilm Targets
| Target Pathway | Reduction in Gene Expression | Biofilm Biomass Reduction | Antibiotic Resensitization |
|---|---|---|---|
| Quorum Sensing (luxS) | 75-85% (mRNA) | 40-50% | 100-1000x MIC reduction |
| EPS Production (csgA, bcsC) | 70-80% (mRNA) | 50-60% | Enhanced matrix penetration |
| Antibiotic Resistance (blaKPC-2) | >90% (mRNA) | 25-35% | 64-128x MIC reduction |
| Metabolic Essential (folA) | 80-90% (mRNA) | 60-70% | Bactericidal effect restored |
Protocol: Validating CRISPR-Cas13a Targeting in Multi-Species Biofilms
crRNA Design and Preparation
Materials:
- CRISPR-Cas13a crRNA backbone plasmids (Addgene #103854)
- T7 or U6 promoter-driven expression vectors
- In vitro transcription reagents
- Target RNA sequences for biofilm metabolic pathways
Procedure:
- Identify Target Sequences: Select 20-27nt target sites within mRNA transcripts of interest (e.g., quorum sensing genes, EPS synthesis enzymes, antibiotic resistance determinants)
- Bioinformatic Validation:
- Use CRISPOR-based design tools (crispor.tefor.net) to minimize off-target effects [69]
- Ensure target accessibility by avoiding stable secondary RNA structures
- Verify species specificity through BLAST analysis against multi-species genome database
- Clone crRNA Sequences: Insert designed spacer sequences into appropriate expression vectors containing antibiotic resistance markers
- In Vitro Validation: Test crRNA efficacy using cell-free transcription-translation systems with target RNA substrates before biofilm experiments
Critical Considerations:
- Design 3-5 crRNAs per target transcript to account for variable efficacy
- Include non-targeting crRNA as negative control
- For multi-species targeting, ensure crRNAs do not cross-react between species without intent
CRISPR-Cas13a Delivery to Multi-Species Biofilms
Materials:
- 96-hour mature multi-species biofilms
- Cas13a protein or expression plasmid
- Formulated crRNA (20µg/µL stock)
- Lipofectamine 3000 or lipid-based nanocarriers
- Opti-MEM reduced serum media
Procedure:
- Prepare CRISPR-Cas13a Complexes:
- For RNP delivery: Complex 2µg Cas13a protein with 4µg crRNA in 50µL Opti-MEM, incubate 15min at 25°C
- For plasmid delivery: Dilute 0.7µg crRNA plasmid and 0.3µg dPspCas13b-FP plasmid in 50µL Opti-MEM [69]
- Formulate Delivery Nanoparticles:
- Dilute 2µL Lipofectamine 3000 in 50µL Opti-MEM, incubate 5min
- Combine nucleic acid and lipid mixtures, incubate 15min at 25°C
- Treat Biofilms:
- Carefully aspirate growth media from mature biofilms
- Apply 100µL CRISPR formulation per well of 12-well plate
- Incubate 4-6h at 37°C, then replace with fresh media
- Assay at 24h and 48h post-treatment
Troubleshooting:
- Poor penetration: Increase nanoparticle surface charge to enhance EPS interaction
- Variable efficacy: Optimize crRNA:Cas13 ratio (2:1 recommended)
- Species-specific delivery: Utilize phage-assisted targeting for complex communities
Diagram 1: Experimental workflow for validating CRISPR-Cas13a in multi-species biofilms.
Functional Validation of Biofilm Modulation
Metabolic Pathway Analysis:
- RNA Extraction and qRT-PCR:
- Harvest biofilm cells with mechanical disruption (bead beating)
- Extract RNA using commercial kits with DNase treatment
- Perform cDNA synthesis and qRT-PCR for target genes (e.g., csgA, fliC, bcsC, luxS)
- Calculate fold-change using ΔΔCt method with housekeeping normalization
- Metabolite Profiling:
- Analyze short-chain fatty acids and quorum sensing molecules via LC-MS
- Correlate metabolic shifts with transcriptional changes
Phenotypic Assessment:
- Biofilm Biomass: Quantify using crystal violet staining at 570nm
- Viability Assessment: Perform resazurin reduction assays and CFU enumeration
- Structural Analysis: Utilize confocal laser scanning microscopy (CLSM) with LIVE/DEAD staining
- Antibiotic Susceptibility: Determine minimum biofilm eradication concentration (MBEC) for relevant antibiotics
Diagram 2: CRISPR-Cas13a targeting of biofilm metabolic pathways and functional consequences.
Research Reagent Solutions
Table 3: Essential Research Reagents for CRISPR-Cas13a Biofilm Studies
| Reagent/Category | Specific Examples | Function & Application Notes |
|---|---|---|
| Cas13a Expression Systems | pHAGE-dPspCas13b-3×EGFP-2×NLS-IRES-puro (Addgene #132397) | Nuclear RNA labeling; modify NLS for cytoplasmic targets |
| crRNA Cloning Vectors | PspCas13b crRNA backbone (Addgene #103854) | gRNA expression; includes antibiotic resistance markers |
| Delivery Reagents | Lipofectamine 3000; Cationic lipid nanoparticles; Engineered phage | Optimal for RNP delivery; enhances biofilm penetration |
| Biofilm Culture Supplies | YESCA media; Cholesterol-coated surfaces; Silicone coupons | Promotes robust pellicle biofilm formation in 3-4 days |
| Assessment Tools | Crystal violet; Resazurin; LIVE/DEAD BacLight; RT-qPCR reagents | Multiplexed viability and biomass quantification |
| Control Reagents | Non-targeting crRNA; Catalytically dead Cas13a; Empty vectors | Essential for distinguishing specific from non-specific effects |
Data Analysis and Interpretation
Quantitative Assessment of Gene Knockdown
Calculate target mRNA reduction using ΔΔCt method with normalization to stable housekeeping genes. Effective CRISPR-Cas13a targeting should achieve 70-90% reduction in target transcript levels within 24 hours post-treatment. Include both species-specific and universal housekeeping genes for multi-species normalization.
Structural and Viability Metrics
- Biofilm Biomass: Significant reduction (40-70%) in crystal violet staining indicates disrupted biofilm integrity
- Metabolic Activity: >50% reduction in resazurin reduction signals impaired metabolic function
- Antibiotic Resensitization: Calculate fold-reduction in minimum biofilm eradication concentration (MBEC) for conventional antibiotics
Statistical Considerations
For multi-species biofilm experiments, employ multivariate statistical approaches to account for:
- Species-interaction effects
- Temporal dynamics of biofilm reorganization
- Dose-response relationships for CRISPR-Cas13a delivery
Include appropriate sample sizes (n≥6 biological replicates) to achieve sufficient power for detecting moderate effects (≥30% reduction) in heterogeneous biofilm communities.
This application note provides a standardized framework for validating CRISPR-Cas13a-based interventions in complex multi-species biofilm models. The protocols outlined enable researchers to assess transcript-specific knockdown efficacy, functional consequences on biofilm integrity, and potential therapeutic applications with enhanced precision over conventional antimicrobial approaches. The integration of robust delivery systems with carefully designed crRNAs creates unprecedented opportunities for targeting metabolic pathways essential for biofilm maintenance and antibiotic tolerance. As CRISPR-Cas13 technologies continue to evolve, their application in multi-species biofilm models will be essential for translating RNA-targeting approaches into effective clinical interventions for persistent biofilm-associated infections.
The persistent challenge of biofilm-associated infections is a central focus in antimicrobial resistance (AMR) research. Biofilms, structured communities of microorganisms encased in an extracellular polymeric substance (EPS), demonstrate intrinsic resistance to conventional antimicrobial therapies, contributing significantly to chronic infections and treatment failures [12] [70]. The matrix reduces antibiotic penetration, creates heterogeneous microenvironments, and harbors metabolically dormant persister cells, leading to tolerance levels up to 1000-fold greater than their planktonic counterparts [12] [57].
Within this context, the CRISPR-Cas13 system emerges as a transformative tool for precision research. Unlike DNA-targeting systems like Cas9, the RNA-targeting Cas13 nuclease can be programmed to knockdown specific messenger RNA (mRNA) transcripts within bacterial metabolic pathways without permanently altering the genome [34] [71]. This application note details the specific success metrics and standardized protocols for employing CRISPR-Cas13 to investigate and disrupt biofilm function, with a dual focus on reducing physical biofilm biomass and resensitizing resistant populations to existing antimicrobials.
Key Performance Indicators and Quantitative Metrics
Evaluating the efficacy of CRISPR-Cas13 interventions requires a multi-faceted approach, quantifying both the physical disintegration of the biofilm structure and the restoration of antimicrobial susceptibility. The following table summarizes the core success metrics.
Table 1: Key Success Metrics for CRISPR-Cas13 Biofilm Intervention
| Metric Category | Specific Metric | Measurement Method | Target Outcome | Supporting Data |
|---|---|---|---|---|
| Biofilm Biomass Reduction | Total Biomass | Crystal Violet (CV) Assay; Confocal Laser Scanning Microscopy (CLSM) | ≥70% reduction in CV absorbance vs. control [12] | Liposomal Cas9 systems reduced P. aeruginosa biofilm by >90% in vitro [12] |
| Structural Integrity | CLSM with LIVE/DEAD staining (BacLight kit) | Increased dead:live cell ratio; disrupted architecture | N/A | |
| Resensitization to Antimicrobials | Minimum Inhibitory Concentration (MIC) | Broth Microdilution (CLSI guidelines) | ≥4-fold decrease in MIC for target antibiotic [57] | N/A |
| Minimum Biofilm Eradication Concentration (MBEC) | MBEC Assay | ≥8-fold decrease in MBEC for target antibiotic [57] | N/A | |
| Metabolic Pathway Knockdown | Target Gene Expression | qRT-PCR | ≥80% reduction in target mRNA levels [34] | Cas13 enables ultra-sensitive (aM level) nucleic acid detection [34] |
| Metabolic Output | GC-MS for SCFAs; HPLC for ATP | Significant change in pathway-specific metabolites | N/A |
Experimental Protocols
Protocol 1: CRISPR-Cas13 Ribonucleoprotein (RNP) Complex Preparation
This protocol describes the in vitro formation of the guided RNP complex for delivery into bacterial cells.
Key Research Reagent Solutions:
- Cas13 Nuclease: Purified Cas13a (e.g., LshCas13a) protein for RNA targeting.
- crRNA: Synthetic CRISPR RNA with a spacer sequence targeting specific mRNA and a direct repeat scaffold.
- Nuclease-Free Buffers: To maintain complex stability and activity.
Procedure:
- crRNA Design: Design a crRNA spacer sequence (∼28 nt) complementary to a region within the target mRNA of the metabolic gene (e.g., gyrA, rpoB, or a quorum-sensing gene). Verify specificity using BLAST against the host genome [34].
- Complex Formation: Combine the following components in a nuclease-free microtube:
- Purified Cas13 protein: 2 µM (final concentration)
- Synthetic crRNA: 3 µM (final concentration)
- 1X Cas13 Reaction Buffer
- Incubation: Mix by pipetting and incubate at 37°C for 15 minutes to allow for complete RNP complex formation.
- Storage: Use immediately or aliquot and store at -80°C for short-term use. Avoid repeated freeze-thaw cycles.
Protocol 2: Lipid-Based Nanoparticle Delivery of RNP to Biofilms
Efficient delivery is critical. This protocol adapts liposomal formulations proven to enhance penetration and editing efficiency [12].
Procedure:
- Biofilm Cultivation: Grow a mature biofilm of the target bacterium (e.g., Pseudomonas aeruginosa) in a 96-well plate or on a relevant surface for 48-72 hours.
- Liposome Encapsulation: Encapsulate the pre-formed Cas13 RNP complex into cationic lipid nanoparticles (e.g., DOTAP/DOPE liposomes) via a thin-film hydration and extrusion method.
- Treatment: Gently wash the mature biofilm with fresh medium to remove non-adherent cells.
- Application: Apply the RNP-loaded liposomes (e.g., 100 µL per well of a 96-well plate) to the biofilm. Co-deliver with a sub-inhibitory concentration of a relevant antibiotic (e.g., Ciprofloxacin) if investigating synergistic effects [12] [57].
- Incubation: Incubate the treated biofilm under optimal growth conditions for 4-24 hours, depending on the experimental design.
Protocol 3: Quantification of Biomass Reduction via Crystal Violet Assay
This is a standard colorimetric assay for quantifying total biofilm biomass.
Procedure:
- Fixation: After treatment, carefully remove the medium and gently wash the biofilm with 1X PBS to remove planktonic cells. Fix the biofilm with 200 µL of 99% methanol per well for 15 minutes.
- Staining: Remove methanol, air-dry the plate, and stain with 200 µL of 0.1% (w/v) crystal violet solution for 20 minutes.
- Washing and Solubilization: Gently wash the plate under running tap water to remove excess stain. Air-dry completely. Add 200 µL of 33% glacial acetic acid to solubilize the stain bound to the biofilm.
- Absorbance Measurement: Transfer 125 µL of the solubilized dye to a new flat-bottom plate. Measure the absorbance at 570 nm using a microplate reader. Express results as percentage reduction relative to the untreated control biofilm, targeting a reduction of ≥70% for a successful intervention [12].
Protocol 4: Assessment of Antibiotic Resensitization via MBEC Assay
The Minimum Biofilm Eradication Concentration (MBEC) measures the lowest concentration of an antibiotic required to eradicate a biofilm.
Procedure:
- Biofilm Treatment: Treat biofilms grown on a peg lid (e.g., Calgary Biofilm Device) with the CRISPR-Cas13 system as described in Protocol 2.
- Antibiotic Challenge: After RNP treatment, transfer the peg lid to a new microtiter plate containing a logarithmic concentration range (e.g., 0.5 to 512 µg/mL) of the target antibiotic in fresh broth. The negative control should be broth only.
- Incubation and Recovery: Incubate the plate for 24 hours. Afterwards, gently wash the pegs and transfer them to a recovery plate containing fresh broth. Sonicate or vortex the pegs to dislodge surviving cells.
- Viability Assessment: Measure the viability of the recovered cells by spotting on agar plates or using metabolic assays like resazurin. The MBEC is defined as the lowest antibiotic concentration that prevents recovery of viable cells. A successful resensitization is indicated by an ≥8-fold decrease in MBEC compared to the non-CRISPR-treated control [57].
Visualization of Workflows and Pathways
Cas13a RNA Targeting Mechanism
Anti-Biofilm Experimental Workflow
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for CRISPR-Cas13 Biofilm Research
| Item | Function/Description | Example/Catalog Considerations |
|---|---|---|
| Cas13a Protein | RNA-targeting effector nuclease; core enzyme of the system. | Purified LshCas13a or PspCas13b, nuclease-free. |
| crRNA | Programmable guide RNA that confers target specificity. | Synthetic crRNA with user-defined spacer and scaffold. |
| Cationic Liposomes | Nanoparticle carrier for efficient RNP delivery into biofilm cells. | DOTAP/DOPE formulations; commercial transfection reagents optimized for bacteria. |
| Crystal Violet | Dye for colorimetric staining and quantification of total biofilm biomass. | 0.1% (w/v) aqueous solution for standard assays. |
| LIVE/DEAD BacLight Kit | Fluorescent viability stain for visualizing live/dead cells and 3D biofilm structure via CLSM. | Contains SYTO 9 (green) and propidium iodide (red). |
| Calgary Biofilm Device | Standardized platform for growing multiple equivalent biofilms and performing MBEC assays. | Also known as the MBEC Assay System. |
| qRT-PCR Reagents | For validating knockdown efficiency of target mRNA post-treatment. | One-step SYBR Green or TaqMan kits. |
| Target Antibiotics | Antibiotics for which resensitization is being investigated. | e.g., Ciprofloxacin, Tobramycin, Colistin. |
Conclusion
CRISPR-Cas13 emerges as a powerful and versatile platform for the precise targeting of RNA transcripts central to biofilm metabolism and resilience. By enabling transient, programmable disruption of key pathways—from quorum sensing to matrix production—Cas13 offers a paradigm shift from broad-spectrum disinfectants to sequence-specific antimicrobials. Future directions must focus on refining in vivo delivery platforms, such as engineered nanoparticles, to penetrate complex biofilm architectures. Further exploration of minimized Cas13 orthologs with high fidelity and minimal collateral activity will be crucial for clinical translation. The integration of Cas13 with diagnostic biosensors and AI-driven target prediction holds the promise of creating intelligent, therapeutic systems capable of autonomously detecting and neutralizing biofilm threats, paving the way for a new class of anti-biofilm therapeutics.
References
- 1. Assessing and advancing the safety of CRISPR-Cas tools [https://www.nature.com/articles/s41467-023-35886-6]
- 2. On the ever‐growing functional versatility of the CRISPR‐ ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10880580/]
- 3. CRISPR 101: Cas9 vs. The Other Cas(s) [https://blog.addgene.org/crispr-101-cas9-vs.-the-other-cass]
- 4. CRISPR Cas13 vs Cas9 in Angelman Syndrome [https://cureangelman.org/the-difference-between-cas13-and-cas9-in-angelman-syndrome-research]
- 5. Engineering of efficiency-enhanced Cas9 and base editors ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10014233/]
- 6. Structures, mechanisms and applications of RNA-centric ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11375968/]
- 7. Structural basis for the activation of a compact CRISPR- ... [https://www.nature.com/articles/s41467-023-41501-5]
- 8. CRISPR–Cas systems as emerging tools for precision ... [https://www.sciencedirect.com/science/article/abs/pii/S0963996925021416]
- 9. The double life of CRISPR–Cas13 [https://www.sciencedirect.com/science/article/pii/S0958166922001239]
- 10. Reprogrammable RNA-targeting CRISPR systems evolved ... [https://www.sciencedirect.com/science/article/pii/S0092867425001035]
- 11. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366227/]
- 12. integrating CRISPR/Cas9 and nanoparticles against biofilm ... [https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-025-04323-4]
- 13. Programmable RNA targeting with CRISPR-Cas13 - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11110701/]
- 14. Current updates on the structural and functional aspects of the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12068846/]
- 15. Systemic evaluation of various CRISPR/Cas13 orthologs ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11619151/]
- 16. A strategy for Cas13 miniaturization based on the structure ... [https://www.nature.com/articles/s41467-023-41320-8]
- 17. RNA structure modulates Cas13 activity and enables ... [https://www.nature.com/articles/s41587-025-02868-6]
- 18. Addressing antimicrobial resistance with Cas9, Cas12, ... [https://www.sciencedirect.com/science/article/pii/S2405844025003937]
- 19. CRISPR–Cas13: Pioneering RNA Editing for Nucleic Acid ... [https://www.sciencedirect.com/science/article/pii/S2693125724000335]
- 20. Advances in RNA-based therapeutics: current breakthroughs ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12592931/]
- 21. The Past, Present, and Future of RNA Editing Therapeutics [https://www.pharmasalmanac.com/articles/rewriting-the-script-the-past-present-and-future-of-rna-editing-therapeutics]
- 22. RNA modifications in cellular metabolism: implications for ... [https://www.nature.com/articles/s41392-024-01777-5]
- 23. RNA-Dependent RNA Targeting by CRISPR-Cas Systems: Characterizations and Applications - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10138764/]
- 24. Advances and challenges in bioproduction of microbial ... [https://www.sciencedirect.com/science/article/pii/S0144861725007945?dgcid=rss_sd_all]
- 25. CRISPR interference to interrogate genes that control ... [https://www.nature.com/articles/s41598-019-52400-5]
- 26. CRISPR Interference (CRISPRi) Inhibition of luxS Gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5445563/]
- 27. A versatile toolkit for CRISPR-Cas13-based RNA manipulation in Drosophila | Genome Biology | Full Text [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-020-02193-y]
- 28. Biofilm, resistance, and quorum sensing: The triple threat in ... [https://www.sciencedirect.com/science/article/pii/S2950194625003462]
- 29. Quorum Sensing: Not Just a Bridge Between Bacteria - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11955508/]
- 30. Microbial biofilm: formation, architecture, antibiotic resistance ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8578483/]
- 31. RNA targeting with CRISPR-Cas13. [https://www.broadinstitute.org/publications/broad125381]
- 32. Effects of an EPS Biosynthesis Gene Cluster ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7911366/]
- 33. Fatty Acid Synthesis Knockdown Promotes Biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9600695/]
- 34. CRISPR‐driven diagnostics: Molecular mechanisms, clinical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12464570/]
- 35. Contemporary strategies and approaches for ... [https://www.sciencedirect.com/science/article/pii/S2095177923002794]
- 36. Biofilms and multidrug resistance: an emerging crisis and the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12237878/]
- 37. Prokaryotic homeostasis – a solution to thrive and survive - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12593503/]
- 38. Phage-based delivery of CRISPR-associated transposases ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12318184/]
- 39. Phage-based delivery of CRISPR-associated transposases ... [https://pubmed.ncbi.nlm.nih.gov/40711918]
- 40. Lipid Nanoparticle-Mediated CRISPR-Cas13a Delivery for ... [https://pubmed.ncbi.nlm.nih.gov/39580667/]
- 41. Targeting bacterial biofilm-related genes with nanoparticle ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2024.1387114/full]
- 42. Engineered phage with antibacterial CRISPR–Cas ... [https://www.nature.com/articles/s41587-023-01759-y]
- 43. Biofilm battleground: Unveiling the hidden challenges, ... [https://www.sciencedirect.com/science/article/abs/pii/S0882401024006223]
- 44. Challenges of antibiotic resistance biofilms and potential ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7966653/]
- 45. Translational challenges and opportunities in biofilm science [https://www.nature.com/articles/s41522-022-00327-7]
- 46. Modified guide RNAs boost CRISPR-Cas13 transcript ... [https://frontlinegenomics.com/modified-guide-rnas-boost-crispr-cas13-transcript-knockdown/]
- 47. Quantification and Comparison of Different Biofilm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12388762/]
- 48. Comparative analysis of biofilm detection methods and ... [https://bmcmicrobiol.biomedcentral.com/articles/10.1186/s12866-025-04506-2]
- 49. A versatile CRISPR - Cas d platform for multiplexed transcriptomic... 13 [https://pubmed.ncbi.nlm.nih.gov/38387457/]
- 50. Negative autoregulation mitigates collateral RNase activity of ... [https://pubmed.ncbi.nlm.nih.gov/35977479/]
- 51. Collateral activity of the CRISPR/RfxCas13d system in ... [https://www.nature.com/articles/s42003-023-04708-2]
- 52. Modeling CRISPR-Cas13d on-target and off-target effects ... [https://www.nature.com/articles/s41467-023-36316-3]
- 53. Chemically modified guide RNAs enhance CRISPR-Cas13 ... [https://pubmed.ncbi.nlm.nih.gov/34343484/]
- 54. Chemically modified guide RNAs enhance CRISPR-Cas13 ... [https://www.sciencedirect.com/science/article/pii/S2451945621003512]
- 55. CRISPR Cas systems: From bacterial defense mechanisms ... [https://www.sciencedirect.com/science/article/pii/S2950194625001128]
- 56. Advances in the application of CRISPR technology ... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1645699/full]
- 57. Combating Antimicrobial Resistance: Innovative Strategies ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12389514/]
- 58. Development of CRISPR-Cas13a-based antimicrobials ... [https://www.nature.com/articles/s41467-020-16731-6]
- 59. Establishment and evaluation of a circAdpgk-0001 knockdown ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12495950/]
- 60. Establishment and evaluation of a circAdpgk-0001 ... [https://peerj.com/articles/20123/]
- 61. Cas13d-mediated isoform-specific RNA knockdown with a ... [https://www.nature.com/articles/s41467-025-62066-5]
- 62. Blunted blades: new CRISPR-derived technologies to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11036814/]
- 63. Systemic evaluation of various CRISPR/Cas13 orthologs for ... [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-024-03448-8]
- 64. RNAi vs. CRISPR: Guide to Selecting the Best Gene ... [https://www.synthego.com/blog/rnai-vs-crispr-guide/]
- 65. dCas13-mediated translational repression for accurate ... [https://www.nature.com/articles/s41467-024-46412-7]
- 66. Modeling CRISPR-Cas13d on-target and off-target effects ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9912244/]
- 67. Advances in application of CRISPR-Cas13a system [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2024.1291557/full]
- 68. The CRISPR-Cas System Differentially Regulates Surface ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9241790/]
- 69. Protocol for Dynamic Imaging of RNA in Living Cells by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7580206/]
- 70. Biofilms and multidrug resistance: an emerging crisis ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1625356/full]
- 71. CRISPR-Cas Systems: Bridging Bacterial Immunity and ... [https://www.mdpi.com/2673-8007/5/4/118]