Research Articles
Shallow Shotgun Sequencing: A Cost-Effective Path to Species-Level Microbiome Insights for Biomedical Research
This article explores shallow shotgun metagenomic sequencing (SSMS) as a powerful, cost-effective methodology bridging the gap between 16S rRNA gene sequencing and deep shotgun metagenomics.
Optimizing PCR Cycles for 16S rRNA Amplification: A Strategic Guide for Reproducible Microbiome Research
This article provides a comprehensive framework for researchers and drug development professionals to optimize PCR cycle numbers in 16S rRNA gene sequencing protocols.
Navigating the Low Biomass Challenge: A Researcher's Guide to Robust 16S rRNA Gene Sequencing
Accurate 16S rRNA gene sequencing of low biomass samples is critical for exploring microbiomes in environments like the respiratory tract, tissues, and clinical specimens, but it is fraught with challenges...
Optimizing Host DNA Depletion in Shotgun Metagenomics: A 2025 Guide for Biomedical Researchers
Shotgun metagenomics has revolutionized microbiome research, but its application in host-derived samples is severely limited by the overwhelming abundance of host DNA.
Functional Profiling from Metagenomic Data: Methods, Applications, and AI-Driven Insights for Biomedical Research
This article provides a comprehensive overview of functional profiling from shotgun metagenomic data, a powerful approach for decoding the functional potential of microbial communities.
Optimizing Library Preparation for Metagenomic Sequencing: A Comprehensive Guide for Robust Microbial Profiling
Metagenomic next-generation sequencing (mNGS) is revolutionizing microbial community analysis and infectious disease diagnostics by enabling unbiased detection of pathogens.
Host DNA Depletion Methods for Shotgun Metagenomics: A 2025 Guide for Clinical Researchers
Shotgun metagenomic sequencing is revolutionizing pathogen detection and microbiome research but is critically limited by the overwhelming presence of host DNA in clinical samples.
A Strategic Guide to Choosing 16S rRNA Variable Regions for Robust Microbiome Research
Selecting the optimal hypervariable region for 16S rRNA sequencing is a critical, yet complex, decision that directly impacts the taxonomic resolution, accuracy, and reproducibility of microbiome studies.
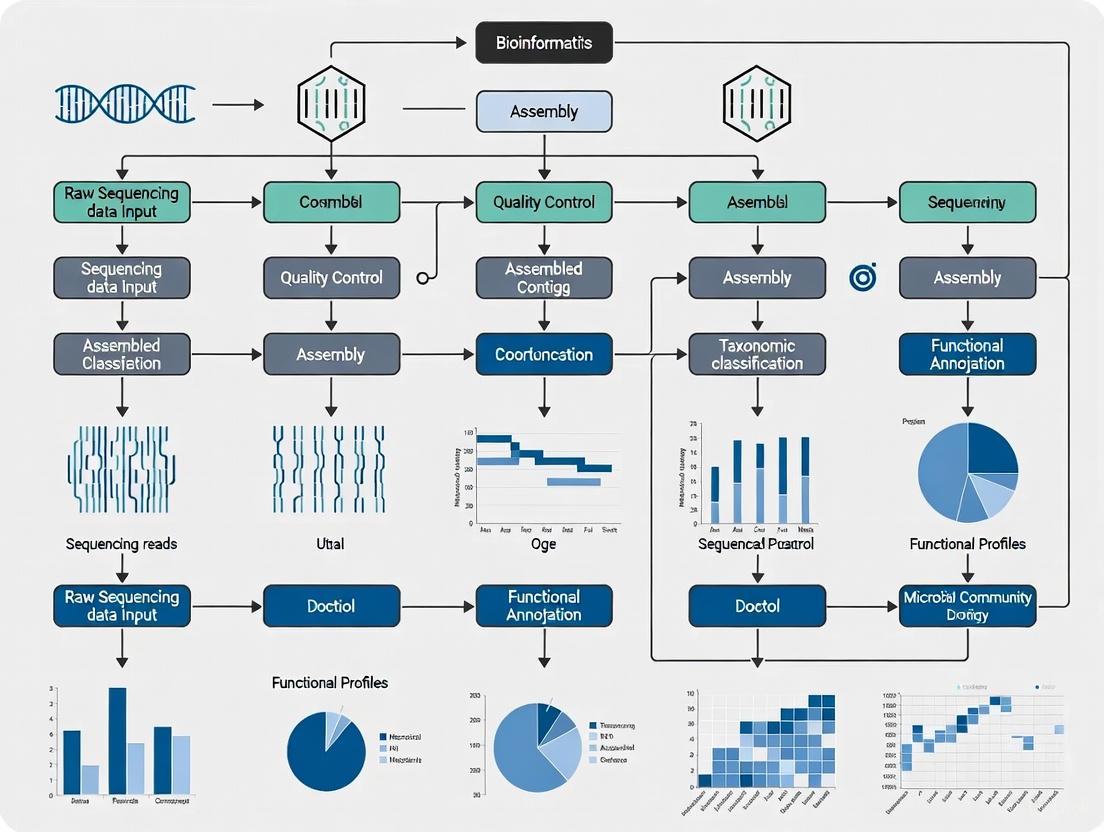
A Comprehensive Guide to Shotgun Metagenomics Bioinformatics Pipelines: From Foundational Concepts to Clinical Validation
This article provides a comprehensive guide to shotgun metagenomics bioinformatics pipelines, tailored for researchers, scientists, and drug development professionals.
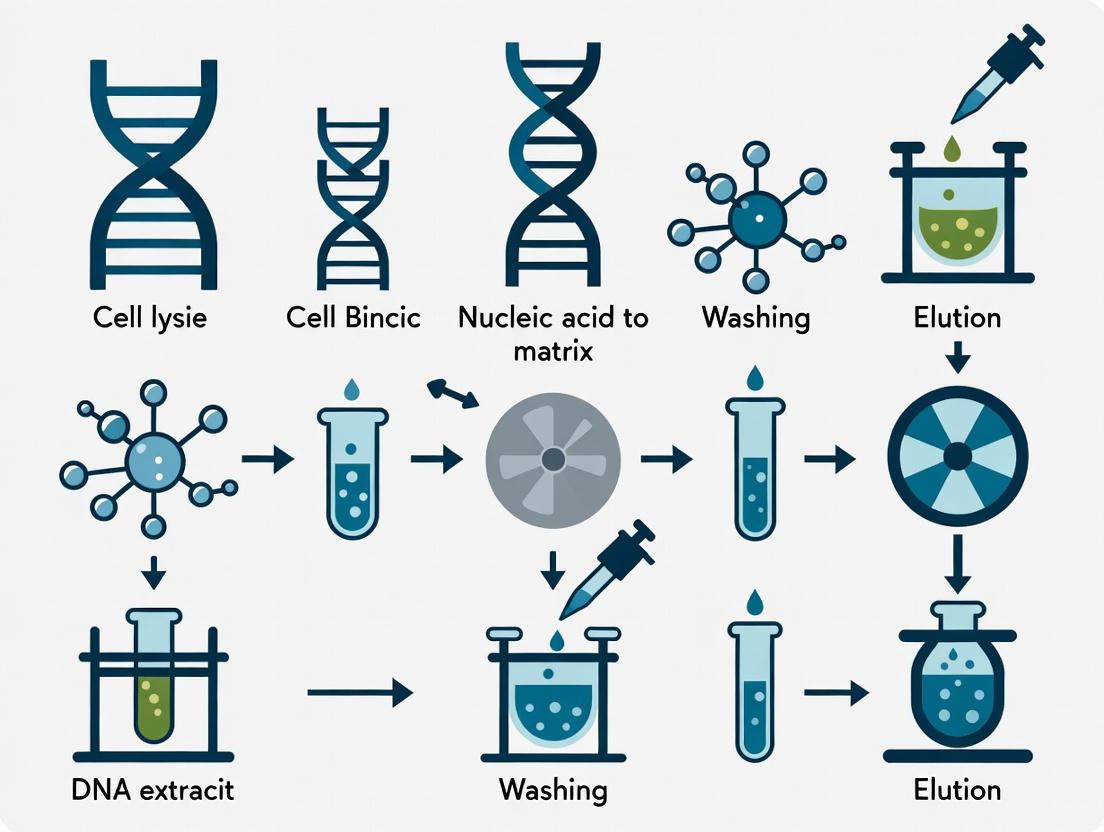
Optimizing DNA Extraction for Metagenomic Sequencing: A Guide for Robust Pathogen Detection and Microbiome Analysis
Metagenomic sequencing has revolutionized pathogen detection and microbiome analysis, but its success is critically dependent on the initial DNA extraction step.